Samuel Kaski
Department of Computer Science, Aalto University, Department of Computer Science, University of Manchester
A High-Performance Implementation of Bayesian Matrix Factorization with Limited Communication
Apr 14, 2020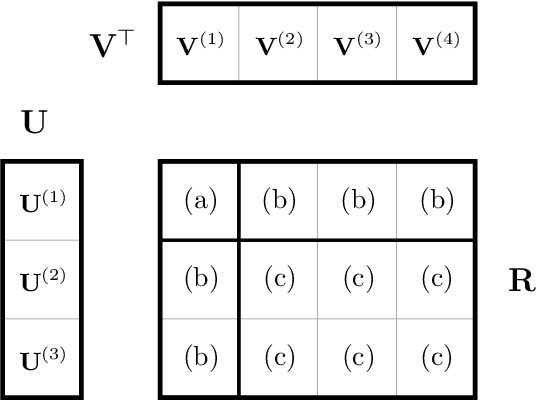
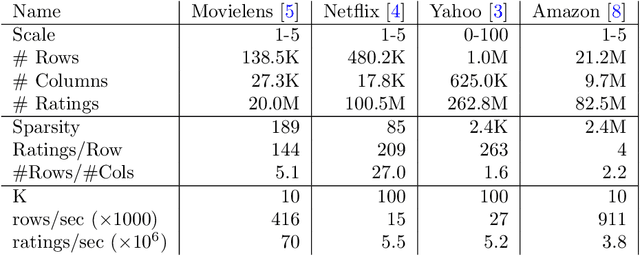
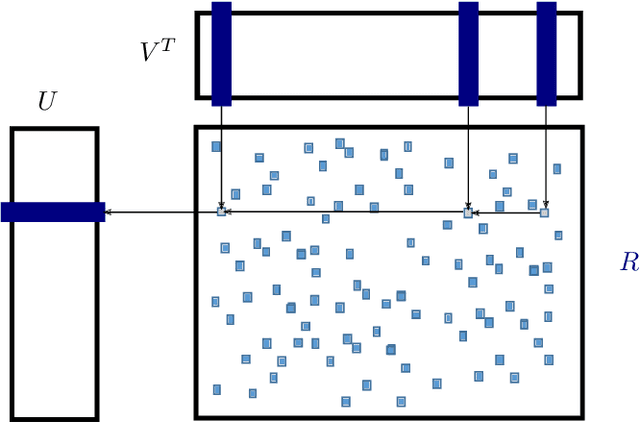
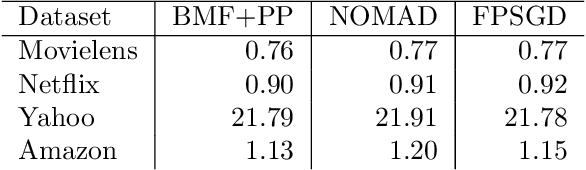
Abstract:Matrix factorization is a very common machine learning technique in recommender systems. Bayesian Matrix Factorization (BMF) algorithms would be attractive because of their ability to quantify uncertainty in their predictions and avoid over-fitting, combined with high prediction accuracy. However, they have not been widely used on large-scale data because of their prohibitive computational cost. In recent work, efforts have been made to reduce the cost, both by improving the scalability of the BMF algorithm as well as its implementation, but so far mainly separately. In this paper we show that the state-of-the-art of both approaches to scalability can be combined. We combine the recent highly-scalable Posterior Propagation algorithm for BMF, which parallelizes computation of blocks of the matrix, with a distributed BMF implementation that users asynchronous communication within each block. We show that the combination of the two methods gives substantial improvements in the scalability of BMF on web-scale datasets, when the goal is to reduce the wall-clock time.
Correlated Feature Selection with Extended Exclusive Group Lasso
Feb 27, 2020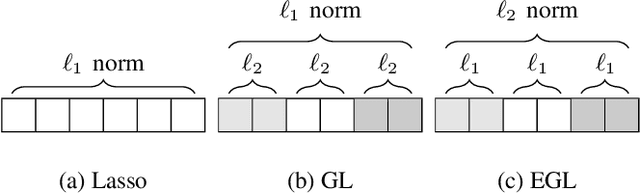
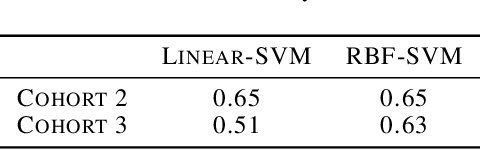
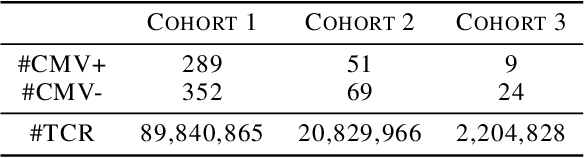
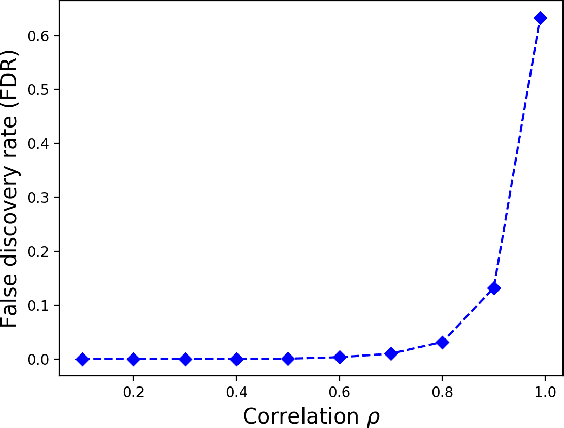

Abstract:In many high dimensional classification or regression problems set in a biological context, the complete identification of the set of informative features is often as important as predictive accuracy, since this can provide mechanistic insight and conceptual understanding. Lasso and related algorithms have been widely used since their sparse solutions naturally identify a set of informative features. However, Lasso performs erratically when features are correlated. This limits the use of such algorithms in biological problems, where features such as genes often work together in pathways, leading to sets of highly correlated features. In this paper, we examine the performance of a Lasso derivative, the exclusive group Lasso, in this setting. We propose fast algorithms to solve the exclusive group Lasso, and introduce a solution to the case when the underlying group structure is unknown. The solution combines stability selection with random group allocation and introduction of artificial features. Experiments with both synthetic and real-world data highlight the advantages of this proposed methodology over Lasso in comprehensive selection of informative features.
Informative Gaussian Scale Mixture Priors for Bayesian Neural Networks
Feb 24, 2020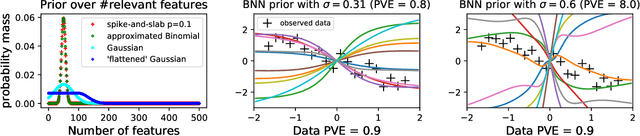
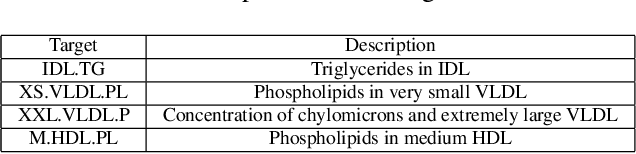
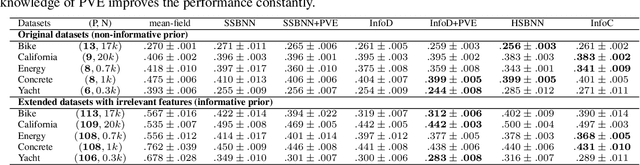
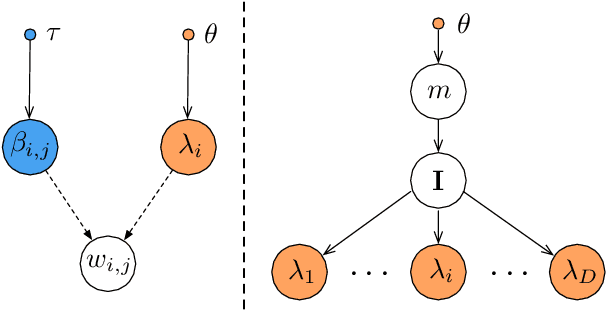

Abstract:Encoding domain knowledge into the prior over the high-dimensional weight space is challenging in Bayesian neural networks. Two types of domain knowledge are commonly available in scientific applications: 1. feature sparsity (number of relevant features); 2. signal-to-noise ratio, quantified, for instance, as the proportion of variance explained (PVE). We show both types of domain knowledge can be encoded into the widely used Gaussian scale mixture priors with Automatic Relevance Determination. Specifically, we propose a new joint prior over the local (i.e., feature-specific) scale parameters to encode the knowledge about feature sparsity, and an algorithm to determine the global scale parameter (shared by all features) according to the PVE. Empirically, we show that the proposed informative prior improves prediction accuracy on publicly available datasets and in a genetics application.
Split-BOLFI for for misspecification-robust likelihood free inference in high dimensions
Feb 21, 2020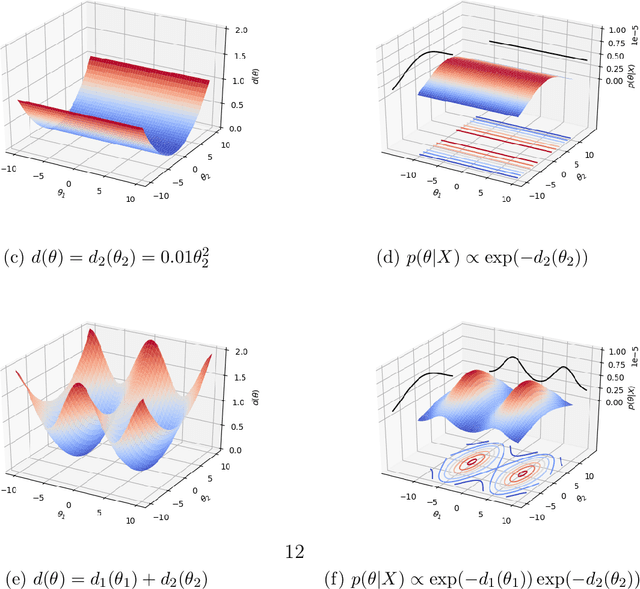
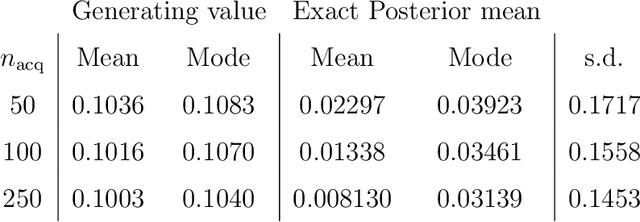
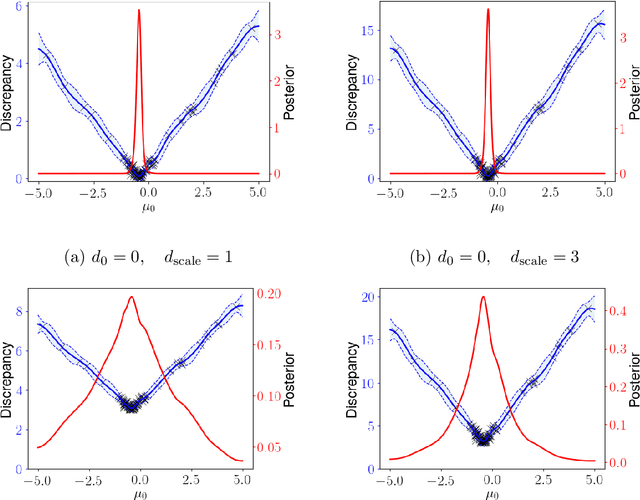

Abstract:Likelihood-free inference for simulator-based statistical models has recently grown rapidly from its infancy to a useful tool for practitioners. However, models with more than a very small number of parameters as the target of inference have remained an enigma, in particular for the approximate Bayesian computation (ABC) community. To advance the possibilities for performing likelihood-free inference in high-dimensional parameter spaces, here we introduce an extension of the popular Bayesian optimisation based approach to approximate discrepancy functions in a probabilistic manner which lends itself to an efficient exploration of the parameter space. Our method achieves computational scalability by using separate acquisition procedures for the discrepancies defined for different parameters. These efficient high-dimensional simulation acquisitions are combined with exponentiated loss-likelihoods to provide a misspecification-robust characterisation of the marginal posterior distribution for all model parameters. The method successfully performs computationally efficient inference in a 100-dimensional space on canonical examples and compares favourably to existing Copula-ABC methods. We further illustrate the potential of this approach by fitting a bacterial transmission dynamics model to daycare centre data, which provides biologically coherent results on the strain competition in a 30-dimensional parameter space.
Projective Preferential Bayesian Optimization
Feb 08, 2020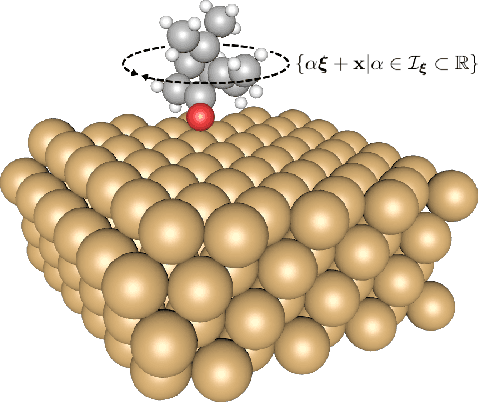
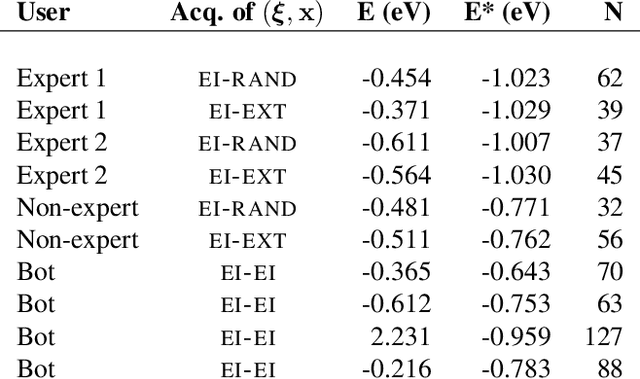
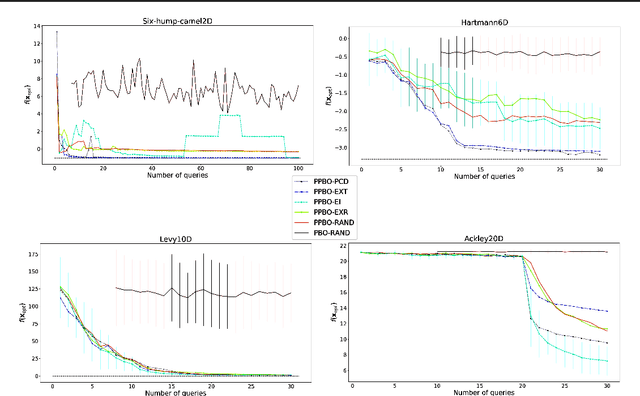
Abstract:Bayesian optimization is an effective method for finding extrema of a black-box function. We propose a new type of Bayesian optimization for learning user preferences in high-dimensional spaces. The central assumption is that the underlying objective function cannot be evaluated directly, but instead a minimizer along a projection can be queried, which we call a projective preferential query. The form of the query allows for feedback that is natural for a human to give, and which enables interaction. This is demonstrated in a user experiment in which the user feedback comes in the form of optimal position and orientation of a molecule adsorbing to a surface. We demonstrate that our framework is able to find a global minimum of a high-dimensional black-box function, which is an infeasible task for existing preferential Bayesian optimization frameworks that are based on pairwise comparisons.
Privacy-preserving data sharing via probabilistic modelling
Jan 29, 2020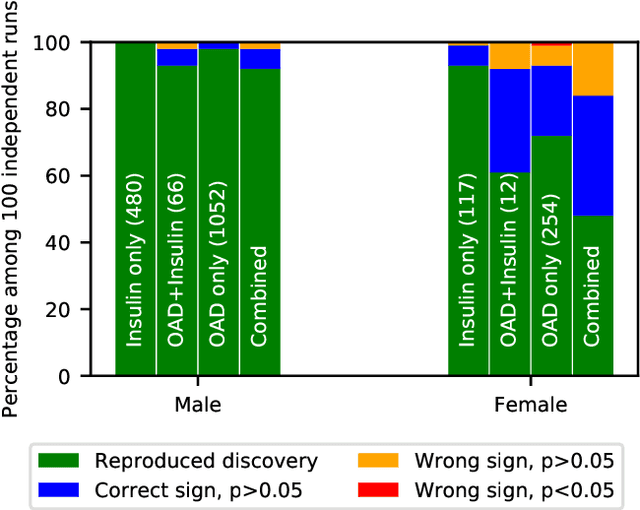
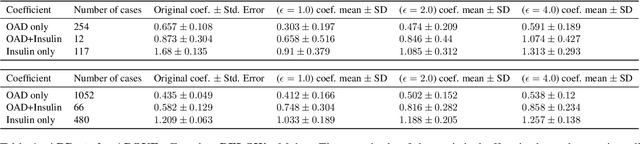
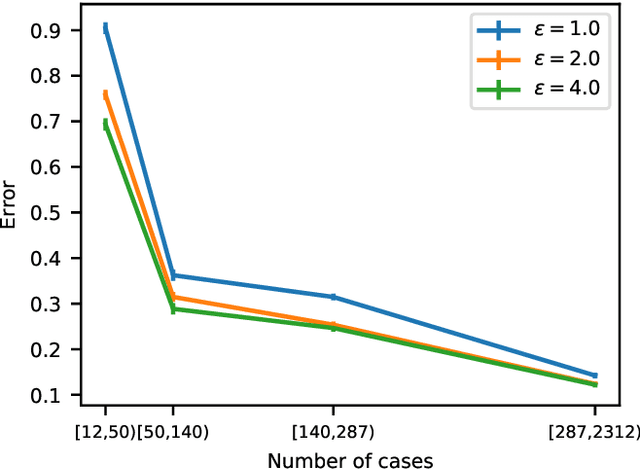
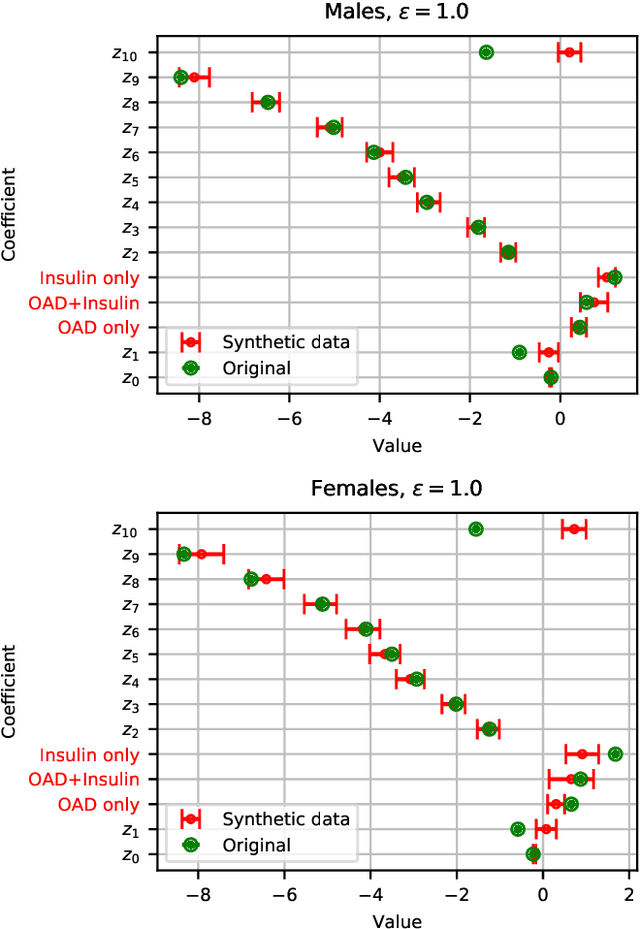
Abstract:Differential privacy allows quantifying privacy loss from computations on sensitive personal data. This loss grows with the number of accesses to the data, making it hard to open the use of such data while respecting privacy. To avoid this limitation, we propose privacy-preserving release of a synthetic version of a data set, which can be used for an unlimited number of analyses with any methods, without affecting the privacy guarantees. The synthetic data generation is based on differentially private learning of a generative probabilistic model which can capture the probability distribution of the original data. We demonstrate empirically that we can reliably reproduce statistical discoveries from the synthetic data. We expect the method to have broad use in sharing anonymized versions of key data sets for research.
Interactive AI with a Theory of Mind
Dec 01, 2019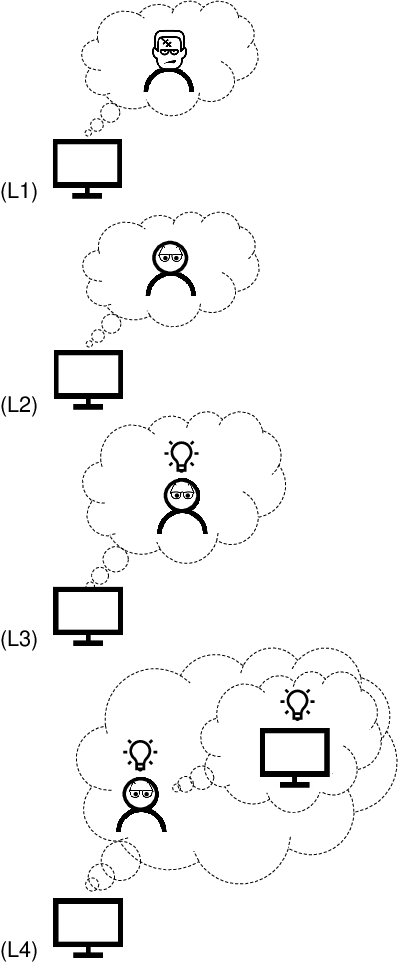
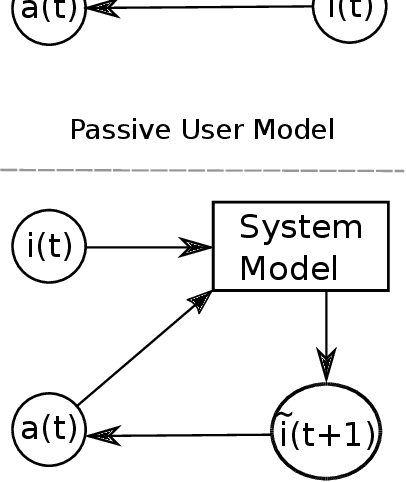
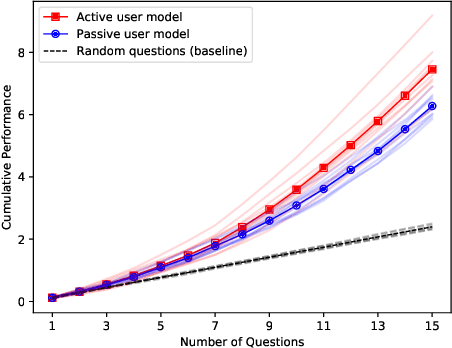
Abstract:Understanding each other is the key to success in collaboration. For humans, attributing mental states to others, the theory of mind, provides the crucial advantage. We argue for formulating human--AI interaction as a multi-agent problem, endowing AI with a computational theory of mind to understand and anticipate the user. To differentiate the approach from previous work, we introduce a categorisation of user modelling approaches based on the level of agency learnt in the interaction. We describe our recent work in using nested multi-agent modelling to formulate user models for multi-armed bandit based interactive AI systems, including a proof-of-concept user study.
Probabilistic Formulation of the Take The Best Heuristic
Nov 01, 2019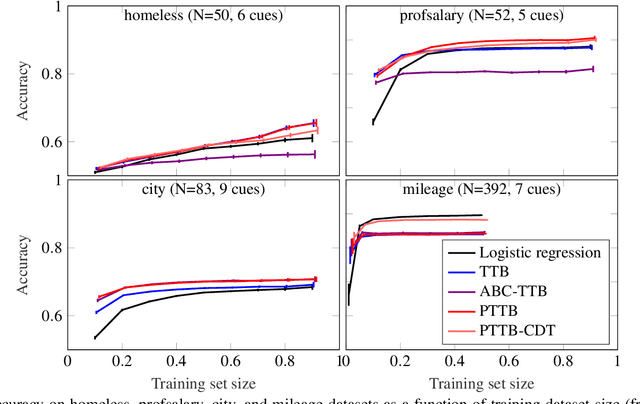
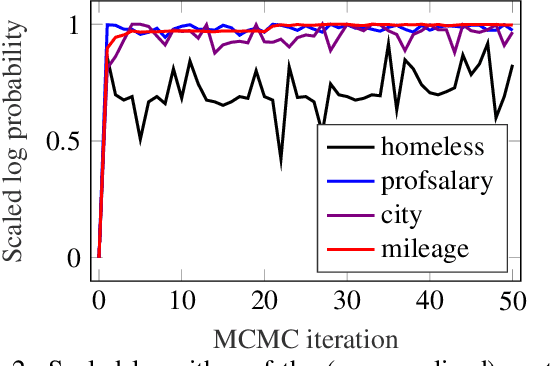
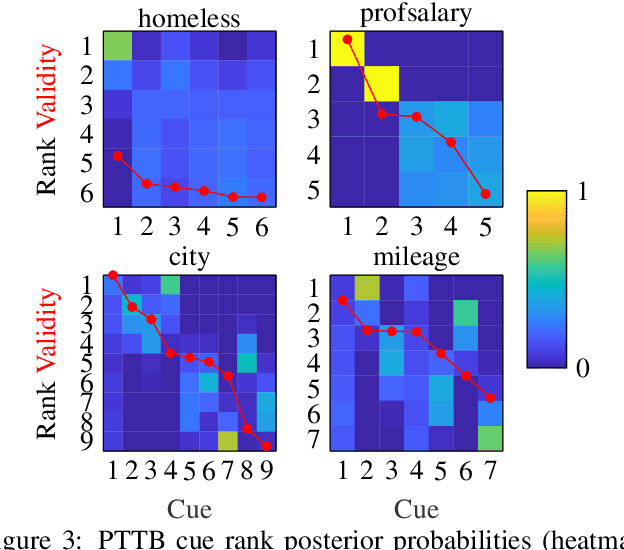
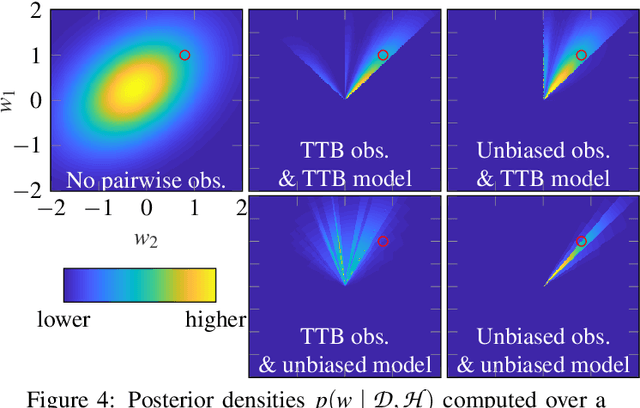
Abstract:The framework of cognitively bounded rationality treats problem solving as fundamentally rational, but emphasises that it is constrained by cognitive architecture and the task environment. This paper investigates a simple decision making heuristic, Take The Best (TTB), within that framework. We formulate TTB as a likelihood-based probabilistic model, where the decision strategy arises by probabilistic inference based on the training data and the model constraints. The strengths of the probabilistic formulation, in addition to providing a bounded rational account of the learning of the heuristic, include natural extensibility with additional cognitively plausible constraints and prior information, and the possibility to embed the heuristic as a subpart of a larger probabilistic model. We extend the model to learn cue discrimination thresholds for continuous-valued cues and experiment with using the model to account for biased preference feedback from a boundedly rational agent in a simulated interactive machine learning task.
Making Bayesian Predictive Models Interpretable: A Decision Theoretic Approach
Oct 21, 2019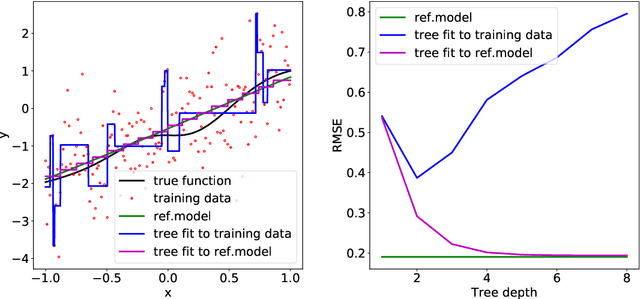
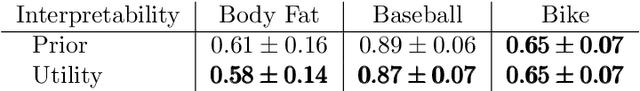
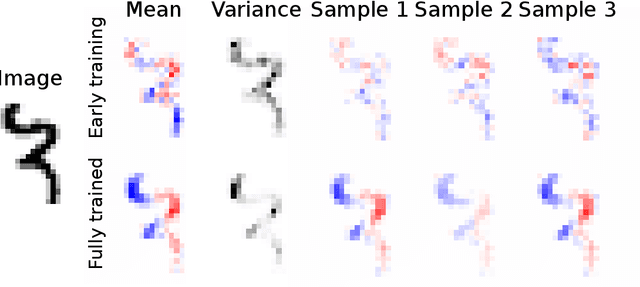
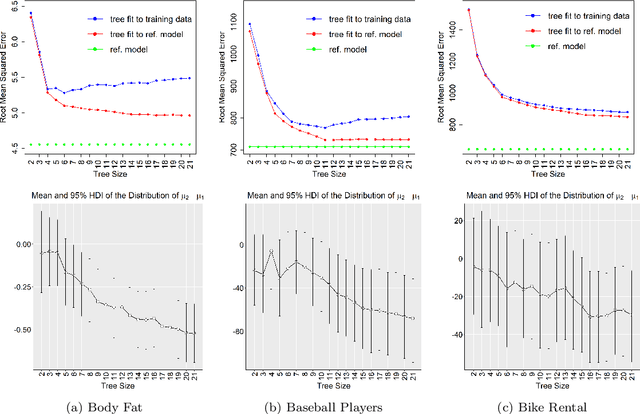
Abstract:A salient approach to interpretable machine learning is to restrict modeling to simple and hence understandable models. In the Bayesian framework, this can be pursued by restricting the model structure and prior to favor interpretable models. Fundamentally, however, interpretability is about users' preferences, not the data generation mechanism: it is more natural to formulate interpretability as a utility function. In this work, we propose an interpretability utility, which explicates the trade-off between explanation fidelity and interpretability in the Bayesian framework. The method consists of two steps. First, a reference model, possibly a black-box Bayesian predictive model compromising no accuracy, is constructed and fitted to the training data. Second, a proxy model from an interpretable model family that best mimics the predictive behaviour of the reference model is found by optimizing the interpretability utility function. The approach is model agnostic - neither the interpretable model nor the reference model are restricted to be from a certain class of models - and the optimization problem can be solved using standard tools in the chosen model family. Through experiments on real-word data sets using decision trees as interpretable models and Bayesian additive regression models as reference models, we show that for the same level of interpretability, our approach generates more accurate models than the earlier alternative of restricting the prior. We also propose a systematic way to measure stabilities of interpretabile models constructed by different interpretability approaches and show that our proposed approach generates more stable models.
Scalable Probabilistic Matrix Factorization with Graph-Based Priors
Sep 11, 2019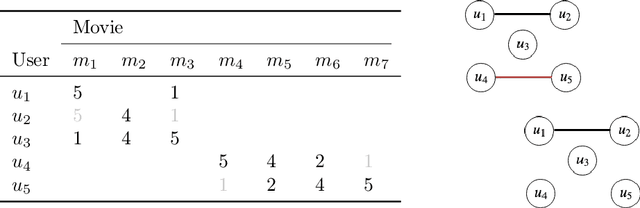
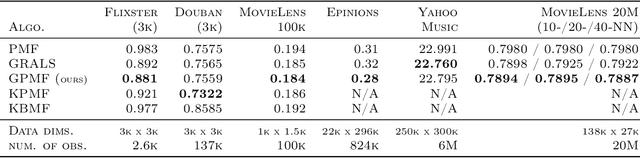
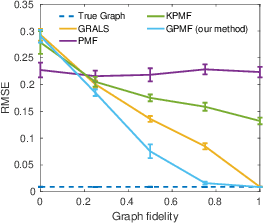
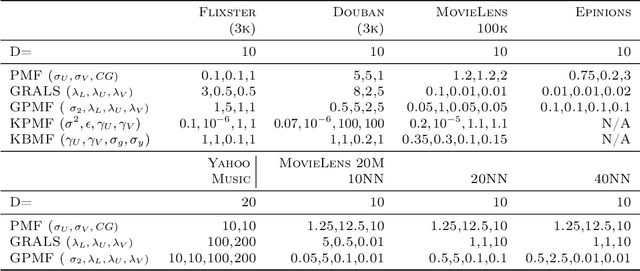
Abstract:In matrix factorization, available graph side-information may not be well suited for the matrix completion problem, having edges that disagree with the latent-feature relations learnt from the incomplete data matrix. We show that removing these $\textit{contested}$ edges improves prediction accuracy and scalability. We identify the contested edges through a highly-efficient graphical lasso approximation. The identification and removal of contested edges adds no computational complexity to state-of-the-art graph-regularized matrix factorization, remaining linear with respect to the number of non-zeros. Computational load even decreases proportional to the number of edges removed. Formulating a probabilistic generative model and using expectation maximization to extend graph-regularised alternating least squares (GRALS) guarantees convergence. Rich simulated experiments illustrate the desired properties of the resulting algorithm. On real data experiments we demonstrate improved prediction accuracy with fewer graph edges (empirical evidence that graph side-information is often inaccurate). A 300 thousand dimensional graph with three million edges (Yahoo music side-information) can be analyzed in under ten minutes on a standard laptop computer demonstrating the efficiency of our graph update.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge