Patrick Rinke
Bayesian Optimization for Mixed-Variable Problems in the Natural Sciences
Apr 08, 2026Abstract:Optimizing expensive black-box objectives over mixed search spaces is a common challenge across the natural sciences. Bayesian optimization (BO) offers sample-efficient strategies through probabilistic surrogate models and acquisition functions. However, its effectiveness diminishes in mixed or high-cardinality discrete spaces, where gradients are unavailable and optimizing the acquisition function becomes computationally demanding. In this work, we generalize the probabilistic reparameterization (PR) approach of Daulton et al. to handle non-equidistant discrete variables, enabling gradient-based optimization in fully mixed-variable settings with Gaussian process (GP) surrogates. With real-world scientific optimization tasks in mind, we conduct systematic benchmarks on synthetic and experimental objectives to obtain an optimized kernel formulations and demonstrate the robustness of our generalized PR method. We additionally show that, when combined with a modified BO workflow, our approach can efficiently optimize highly discontinuous and discretized objective landscapes. This work establishes a practical BO framework for addressing fully mixed optimization problems in the natural sciences, and is particularly well suited to autonomous laboratory settings where noise, discretization, and limited data are inherent.
Active Learning of Molecular Data for Task-Specific Objectives
Aug 20, 2024Abstract:Active learning (AL) has shown promise for being a particularly data-efficient machine learning approach. Yet, its performance depends on the application and it is not clear when AL practitioners can expect computational savings. Here, we carry out a systematic AL performance assessment for three diverse molecular datasets and two common scientific tasks: compiling compact, informative datasets and targeted molecular searches. We implemented AL with Gaussian processes (GP) and used the many-body tensor as molecular representation. For the first task, we tested different data acquisition strategies, batch sizes and GP noise settings. AL was insensitive to the acquisition batch size and we observed the best AL performance for the acquisition strategy that combines uncertainty reduction with clustering to promote diversity. However, for optimal GP noise settings, AL did not outperform randomized selection of data points. Conversely, for targeted searches, AL outperformed random sampling and achieved data savings up to 64%. Our analysis provides insight into this task-specific performance difference in terms of target distributions and data collection strategies. We established that the performance of AL depends on the relative distribution of the target molecules in comparison to the total dataset distribution, with the largest computational savings achieved when their overlap is minimal.
Projective Preferential Bayesian Optimization
Feb 08, 2020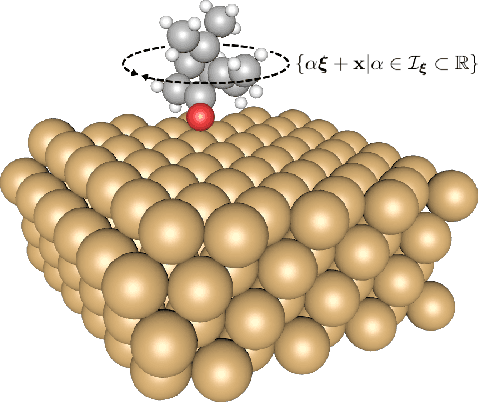
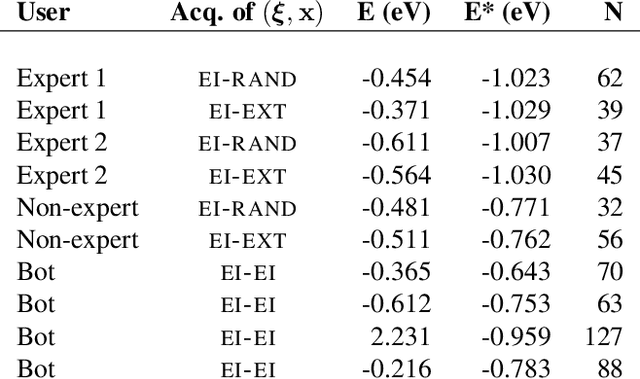
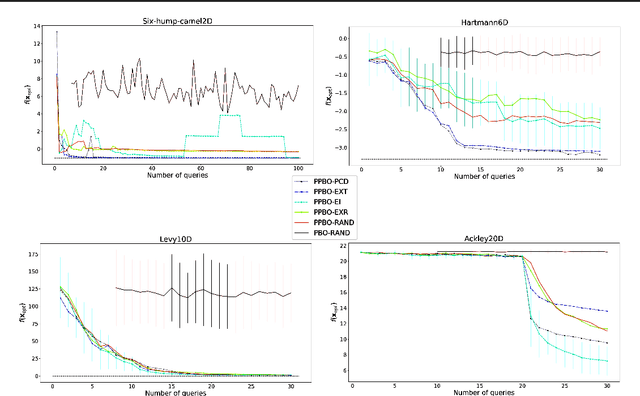
Abstract:Bayesian optimization is an effective method for finding extrema of a black-box function. We propose a new type of Bayesian optimization for learning user preferences in high-dimensional spaces. The central assumption is that the underlying objective function cannot be evaluated directly, but instead a minimizer along a projection can be queried, which we call a projective preferential query. The form of the query allows for feedback that is natural for a human to give, and which enables interaction. This is demonstrated in a user experiment in which the user feedback comes in the form of optimal position and orientation of a molecule adsorbing to a surface. We demonstrate that our framework is able to find a global minimum of a high-dimensional black-box function, which is an infeasible task for existing preferential Bayesian optimization frameworks that are based on pairwise comparisons.
DScribe: Library of Descriptors for Machine Learning in Materials Science
Apr 18, 2019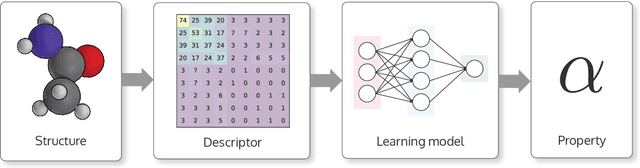
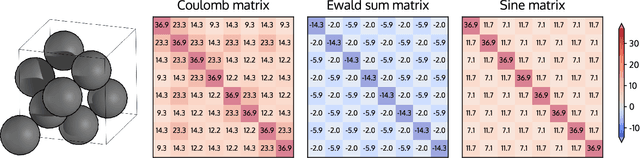

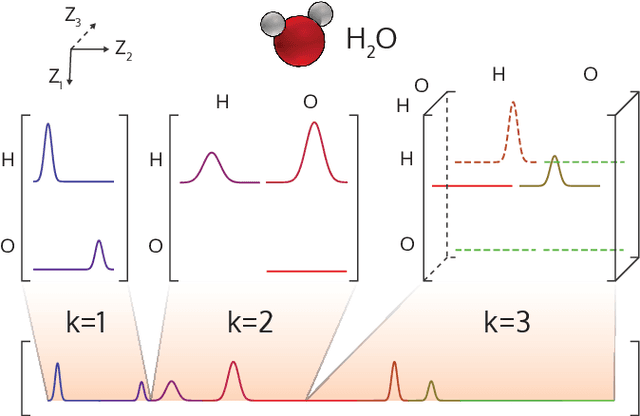
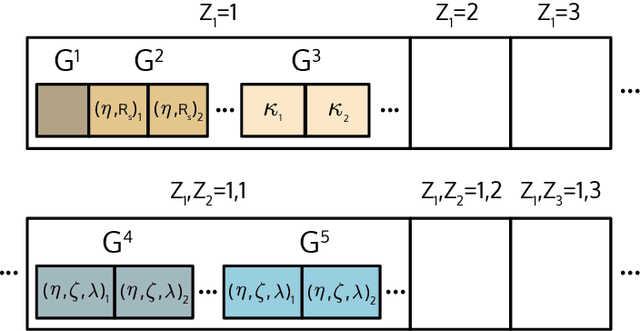
Abstract:DScribe is a software package for machine learning that provides popular feature transformations ("descriptors") for atomistic materials simulations. DScribe accelerates the application of machine learning for atomistic property prediction by providing user-friendly, off-the-shelf descriptor implementations. The package currently contains implementations for Coulomb matrix, Ewald sum matrix, sine matrix, Many-body Tensor Representation (MBTR), Atom-centered Symmetry Function (ACSF) and Smooth Overlap of Atomic Positions (SOAP). Usage of the package is illustrated for two different applications: formation energy prediction for solids and ionic charge prediction for atoms in organic molecules. The package is freely available under the open-source Apache License 2.0.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge