Philippe C. Cattin
NOIR: Neural Operator mapping for Implicit Representations
Mar 13, 2026Abstract:This paper presents NOIR, a framework that reframes core medical imaging tasks as operator learning between continuous function spaces, challenging the prevailing paradigm of discrete grid-based deep learning. Instead of operating on fixed pixel or voxel grids, NOIR embeds discrete medical signals into shared Implicit Neural Representations and learns a Neural Operator that maps between their latent modulations, enabling resolution-independent function-to-function transformations. We evaluate NOIR across multiple 2D and 3D downstream tasks, including segmentation, shape completion, image-to-image translation, and image synthesis, on several public datasets such as Shenzhen, OASIS-4, SkullBreak, fastMRI, as well as an in-house clinical dataset. It achieves competitive performance at native resolution while demonstrating strong robustness to unseen discretizations, and empirically satisfies key theoretical properties of neural operators. The project page is available here: https://github.com/Sidaty1/NOIR-io.
AutoFFS: Adversarial Deformations for Facial Feminization Surgery Planning
Mar 02, 2026Abstract:Facial feminization surgery (FFS) is a key component of gender affirmation for transgender and gender diverse patients, aiming to reshape craniofacial structures toward a female morphology. Current surgical planning procedures largely rely on subjective clinical assessment, lacking quantitative and reproducible anatomical guidance. We therefore propose AutoFFS, a novel data-driven framework that generates counterfactual skull morphologies through adversarial free-form deformations. Our method performs a deformation-based targeted adversarial attack on an ensemble of pre-trained binary sex classifiers that learned sexual dimorphism, effectively transforming individual skull shapes toward the target sex. The generated counterfactual skull morphologies provide a quantitative foundation for preoperative planning in FFS, driving advances in this largely overlooked patient group. We validate our approach through classifier-based evaluation and a human perceptual study, confirming that the generated morphologies exhibit target sex characteristics.
End2Reg: Learning Task-Specific Segmentation for Markerless Registration in Spine Surgery
Dec 15, 2025



Abstract:Purpose: Intraoperative navigation in spine surgery demands millimeter-level accuracy. Current systems based on intraoperative radiographic imaging and bone-anchored markers are invasive, radiation-intensive and workflow disruptive. Recent markerless RGB-D registration methods offer a promising alternative, but existing approaches rely on weak segmentation labels to isolate relevant anatomical structures, which can propagate errors throughout registration. Methods: We present End2Reg an end-to-end deep learning framework that jointly optimizes segmentation and registration, eliminating the need for weak segmentation labels and manual steps. The network learns segmentation masks specifically optimized for registration, guided solely by the registration objective without direct segmentation supervision. Results: The proposed framework achieves state-of-the-art performance on ex- and in-vivo benchmarks, reducing median Target Registration Error by 32% to 1.83mm and mean Root Mean Square Error by 45% to 3.95mm, respectively. An ablation study confirms that end-to-end optimization significantly improves registration accuracy. Conclusion: The presented end-to-end RGB-D registration pipeline removes dependency on weak labels and manual steps, advancing towards fully automatic, markerless intraoperative navigation. Code and interactive visualizations are available at: https://lorenzopettinari.github.io/end-2-reg/.
Towards MR-Based Trochleoplasty Planning
Aug 08, 2025Abstract:To treat Trochlear Dysplasia (TD), current approaches rely mainly on low-resolution clinical Magnetic Resonance (MR) scans and surgical intuition. The surgeries are planned based on surgeons experience, have limited adoption of minimally invasive techniques, and lead to inconsistent outcomes. We propose a pipeline that generates super-resolved, patient-specific 3D pseudo-healthy target morphologies from conventional clinical MR scans. First, we compute an isotropic super-resolved MR volume using an Implicit Neural Representation (INR). Next, we segment femur, tibia, patella, and fibula with a multi-label custom-trained network. Finally, we train a Wavelet Diffusion Model (WDM) to generate pseudo-healthy target morphologies of the trochlear region. In contrast to prior work producing pseudo-healthy low-resolution 3D MR images, our approach enables the generation of sub-millimeter resolved 3D shapes compatible for pre- and intraoperative use. These can serve as preoperative blueprints for reshaping the femoral groove while preserving the native patella articulation. Furthermore, and in contrast to other work, we do not require a CT for our pipeline - reducing the amount of radiation. We evaluated our approach on 25 TD patients and could show that our target morphologies significantly improve the sulcus angle (SA) and trochlear groove depth (TGD). The code and interactive visualization are available at https://wehrlimi.github.io/sr-3d-planning/.
fastWDM3D: Fast and Accurate 3D Healthy Tissue Inpainting
Jul 17, 2025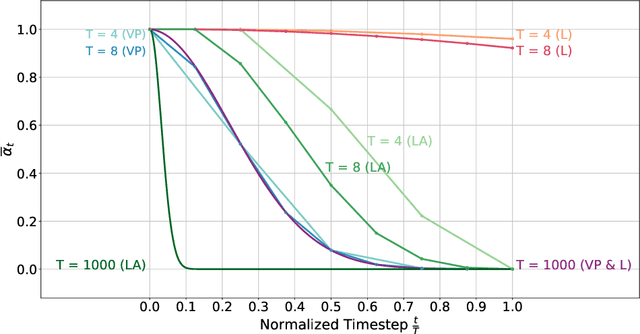
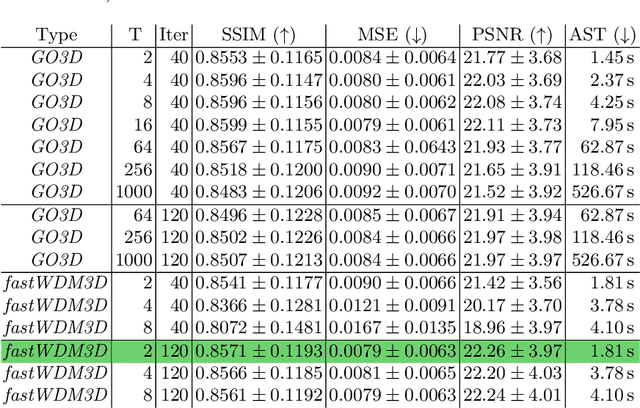
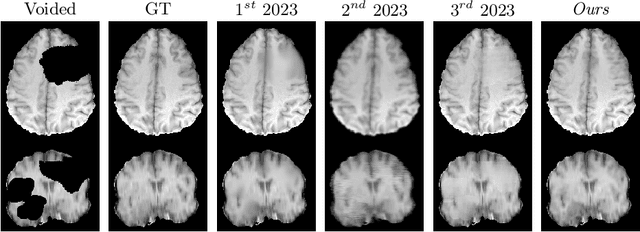
Abstract:Healthy tissue inpainting has significant applications, including the generation of pseudo-healthy baselines for tumor growth models and the facilitation of image registration. In previous editions of the BraTS Local Synthesis of Healthy Brain Tissue via Inpainting Challenge, denoising diffusion probabilistic models (DDPMs) demonstrated qualitatively convincing results but suffered from low sampling speed. To mitigate this limitation, we adapted a 2D image generation approach, combining DDPMs with generative adversarial networks (GANs) and employing a variance-preserving noise schedule, for the task of 3D inpainting. Our experiments showed that the variance-preserving noise schedule and the selected reconstruction losses can be effectively utilized for high-quality 3D inpainting in a few time steps without requiring adversarial training. We applied our findings to a different architecture, a 3D wavelet diffusion model (WDM3D) that does not include a GAN component. The resulting model, denoted as fastWDM3D, obtained a SSIM of 0.8571, a MSE of 0.0079, and a PSNR of 22.26 on the BraTS inpainting test set. Remarkably, it achieved these scores using only two time steps, completing the 3D inpainting process in 1.81 s per image. When compared to other DDPMs used for healthy brain tissue inpainting, our model is up to 800 x faster while still achieving superior performance metrics. Our proposed method, fastWDM3D, represents a promising approach for fast and accurate healthy tissue inpainting. Our code is available at https://github.com/AliciaDurrer/fastWDM3D.
cIDIR: Conditioned Implicit Neural Representation for Regularized Deformable Image Registration
Jul 17, 2025



Abstract:Regularization is essential in deformable image registration (DIR) to ensure that the estimated Deformation Vector Field (DVF) remains smooth, physically plausible, and anatomically consistent. However, fine-tuning regularization parameters in learning-based DIR frameworks is computationally expensive, often requiring multiple training iterations. To address this, we propose cIDI, a novel DIR framework based on Implicit Neural Representations (INRs) that conditions the registration process on regularization hyperparameters. Unlike conventional methods that require retraining for each regularization hyperparameter setting, cIDIR is trained over a prior distribution of these hyperparameters, then optimized over the regularization hyperparameters by using the segmentations masks as an observation. Additionally, cIDIR models a continuous and differentiable DVF, enabling seamless integration of advanced regularization techniques via automatic differentiation. Evaluated on the DIR-LAB dataset, $\operatorname{cIDIR}$ achieves high accuracy and robustness across the dataset.
Generating 3D Pseudo-Healthy Knee MR Images to Support Trochleoplasty Planning
Dec 13, 2024



Abstract:Purpose: Trochlear Dysplasia (TD) is a common malformation in adolescents, leading to anterior knee pain and instability. Surgical interventions such as trochleoplasty require precise planning to correct the trochlear groove. However, no standardized preoperative plan exists to guide surgeons in reshaping the femur. This study aims to generate patient-specific, pseudo-healthy MR images of the trochlear region that should theoretically align with the respective patient's patella, potentially supporting the pre-operative planning of trochleoplasty. Methods: We employ a Wavelet Diffusion Model (WDM) to generate personalized pseudo-healthy, anatomically plausible MR scans of the trochlear region. We train our model using knee MR scans of healthy subjects. During inference, we mask out pathological regions around the patella in scans of patients affected by TD, and replace them with their pseudo-healthy counterpart. An orthopedic surgeon measured the sulcus angle (SA), trochlear groove depth (TGD) and D\'ejour classification in MR scans before and after inpainting. The code is available at https://github.com/wehrlimi/Generate-Pseudo-Healthy-Knee-MRI . Results: The inpainting by our model significantly improves the SA, TGD and D\'ejour classification in a study with 49 knee MR scans. Conclusion: This study demonstrates the potential of WDMs in providing surgeons with patient-specific guidance. By offering anatomically plausible MR scans, the method could potentially enhance the precision and preoperative planning of trochleoplasty, and pave the way to more minimally invasive surgeries.
cWDM: Conditional Wavelet Diffusion Models for Cross-Modality 3D Medical Image Synthesis
Nov 26, 2024



Abstract:This paper contributes to the "BraTS 2024 Brain MR Image Synthesis Challenge" and presents a conditional Wavelet Diffusion Model (cWDM) for directly solving a paired image-to-image translation task on high-resolution volumes. While deep learning-based brain tumor segmentation models have demonstrated clear clinical utility, they typically require MR scans from various modalities (T1, T1ce, T2, FLAIR) as input. However, due to time constraints or imaging artifacts, some of these modalities may be missing, hindering the application of well-performing segmentation algorithms in clinical routine. To address this issue, we propose a method that synthesizes one missing modality image conditioned on three available images, enabling the application of downstream segmentation models. We treat this paired image-to-image translation task as a conditional generation problem and solve it by combining a Wavelet Diffusion Model for high-resolution 3D image synthesis with a simple conditioning strategy. This approach allows us to directly apply our model to full-resolution volumes, avoiding artifacts caused by slice- or patch-wise data processing. While this work focuses on a specific application, the presented method can be applied to all kinds of paired image-to-image translation problems, such as CT $\leftrightarrow$ MR and MR $\leftrightarrow$ PET translation, or mask-conditioned anatomically guided image generation.
Denoising Diffusion Models for Anomaly Localization in Medical Images
Oct 31, 2024



Abstract:This chapter explores anomaly localization in medical images using denoising diffusion models. After providing a brief methodological background of these models, including their application to image reconstruction and their conditioning using guidance mechanisms, we provide an overview of available datasets and evaluation metrics suitable for their application to anomaly localization in medical images. In this context, we discuss supervision schemes ranging from fully supervised segmentation to semi-supervised, weakly supervised, self-supervised, and unsupervised methods, and provide insights into the effectiveness and limitations of these approaches. Furthermore, we highlight open challenges in anomaly localization, including detection bias, domain shift, computational cost, and model interpretability. Our goal is to provide an overview of the current state of the art in the field, outline research gaps, and highlight the potential of diffusion models for robust anomaly localization in medical images.
Deep Generative Models for 3D Medical Image Synthesis
Oct 23, 2024



Abstract:Deep generative modeling has emerged as a powerful tool for synthesizing realistic medical images, driving advances in medical image analysis, disease diagnosis, and treatment planning. This chapter explores various deep generative models for 3D medical image synthesis, with a focus on Variational Autoencoders (VAEs), Generative Adversarial Networks (GANs), and Denoising Diffusion Models (DDMs). We discuss the fundamental principles, recent advances, as well as strengths and weaknesses of these models and examine their applications in clinically relevant problems, including unconditional and conditional generation tasks like image-to-image translation and image reconstruction. We additionally review commonly used evaluation metrics for assessing image fidelity, diversity, utility, and privacy and provide an overview of current challenges in the field.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge