Peter Boor
Explainable histomorphology-based survival prediction of glioblastoma, IDH-wildtype
Jan 16, 2026Abstract:Glioblastoma, IDH-wildtype (GBM-IDHwt) is the most common malignant brain tumor. Histomorphology is a crucial component of the integrated diagnosis of GBM-IDHwt. Artificial intelligence (AI) methods have shown promise to extract additional prognostic information from histological whole-slide images (WSI) of hematoxylin and eosin-stained glioblastoma tissue. Here, we present an explainable AI-based method to support systematic interpretation of histomorphological features associated with survival. It combines an explainable multiple instance learning (MIL) architecture with a sparse autoencoder (SAE) to relate human-interpretable visual patterns of tissue to survival. The MIL architecture directly identifies prognosis-relevant image tiles and the SAE maps these tiles post-hoc to visual patterns. The MIL method was trained and evaluated using a new real-world dataset that comprised 720 GBM-IDHwt cases from three hospitals and four cancer registries in Germany. The SAE was trained using 1878 WSIs of glioblastoma from five independent public data collections. Despite the many factors influencing survival time, our method showed some ability to discriminate between patients living less than 180 days or more than 360 days solely based on histomorphology (AUC: 0.67; 95% CI: 0.63-0.72). Cox proportional hazards regression confirmed a significant difference in survival time between the predicted groups after adjustment for established prognostic factors (hazard ratio: 1.47; 95% CI: 1.26-1.72). Our method identified multiple interpretable visual patterns associated with survival. Three neuropathologists separately found that 21 of the 24 most strongly associated patterns could be clearly attributed to seven histomorphological categories. Necrosis and hemorrhage appeared to be associated with shorter survival while highly cellular tumor areas were associated with longer survival.
Recommendations on test datasets for evaluating AI solutions in pathology
Apr 21, 2022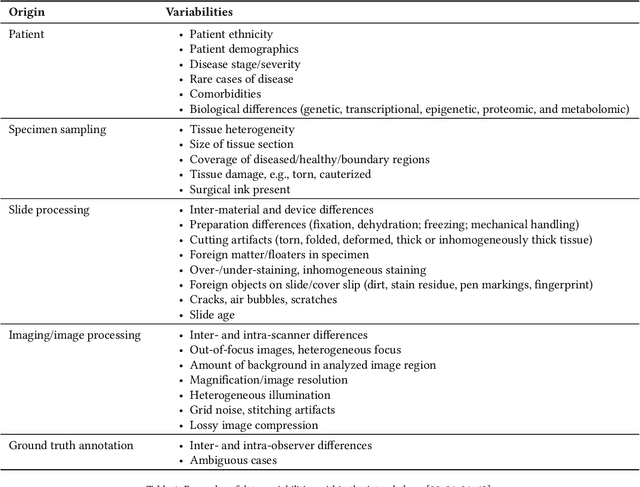
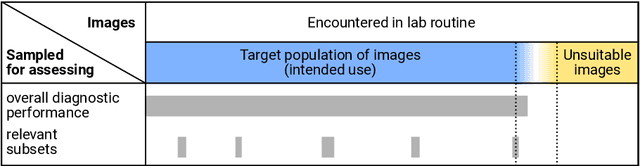
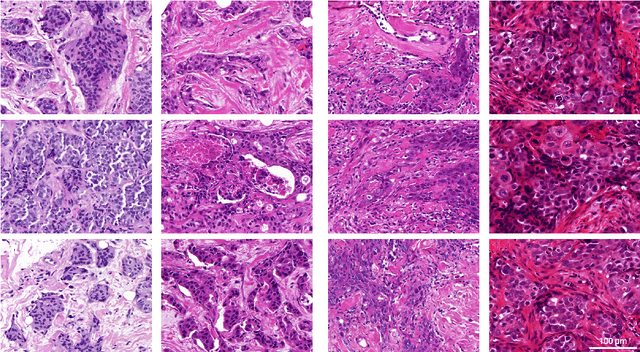

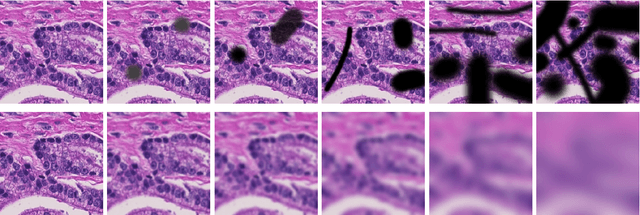
Abstract:Artificial intelligence (AI) solutions that automatically extract information from digital histology images have shown great promise for improving pathological diagnosis. Prior to routine use, it is important to evaluate their predictive performance and obtain regulatory approval. This assessment requires appropriate test datasets. However, compiling such datasets is challenging and specific recommendations are missing. A committee of various stakeholders, including commercial AI developers, pathologists, and researchers, discussed key aspects and conducted extensive literature reviews on test datasets in pathology. Here, we summarize the results and derive general recommendations for the collection of test datasets. We address several questions: Which and how many images are needed? How to deal with low-prevalence subsets? How can potential bias be detected? How should datasets be reported? What are the regulatory requirements in different countries? The recommendations are intended to help AI developers demonstrate the utility of their products and to help regulatory agencies and end users verify reported performance measures. Further research is needed to formulate criteria for sufficiently representative test datasets so that AI solutions can operate with less user intervention and better support diagnostic workflows in the future.
Improving Unsupervised Stain-To-Stain Translation using Self-Supervision and Meta-Learning
Dec 16, 2021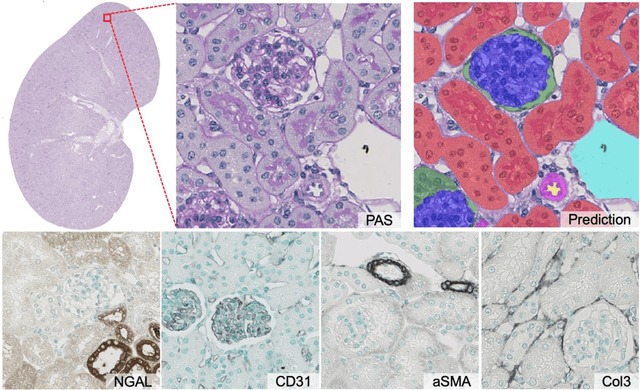

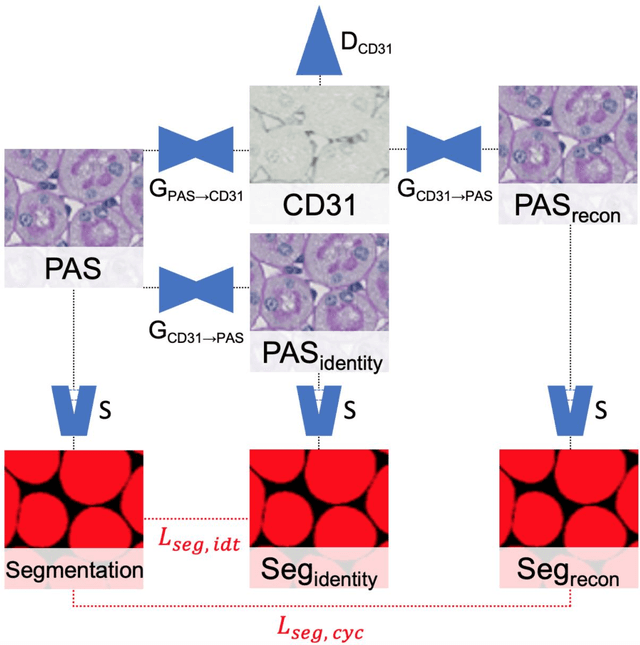
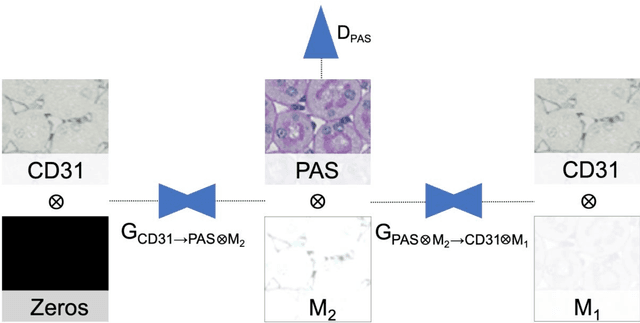
Abstract:In digital pathology, many image analysis tasks are challenged by the need for large and time-consuming manual data annotations to cope with various sources of variability in the image domain. Unsupervised domain adaptation based on image-to-image translation is gaining importance in this field by addressing variabilities without the manual overhead. Here, we tackle the variation of different histological stains by unsupervised stain-to-stain translation to enable a stain-independent applicability of a deep learning segmentation model. We use CycleGANs for stain-to-stain translation in kidney histopathology, and propose two novel approaches to improve translational effectivity. First, we integrate a prior segmentation network into the CycleGAN for a self-supervised, application-oriented optimization of translation through semantic guidance, and second, we incorporate extra channels to the translation output to implicitly separate artificial meta-information otherwise encoded for tackling underdetermined reconstructions. The latter showed partially superior performances to the unmodified CycleGAN, but the former performed best in all stains providing instance-level Dice scores ranging between 78% and 92% for most kidney structures, such as glomeruli, tubules, and veins. However, CycleGANs showed only limited performance in the translation of other structures, e.g. arteries. Our study also found somewhat lower performance for all structures in all stains when compared to segmentation in the original stain. Our study suggests that with current unsupervised technologies, it seems unlikely to produce generally applicable fake stains.
Unsupervisedly Training GANs for Segmenting Digital Pathology with Automatically Generated Annotations
Aug 01, 2018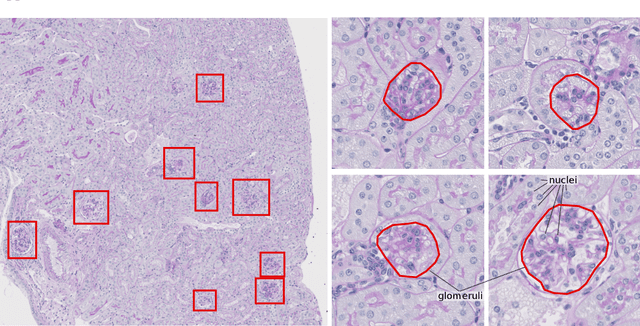
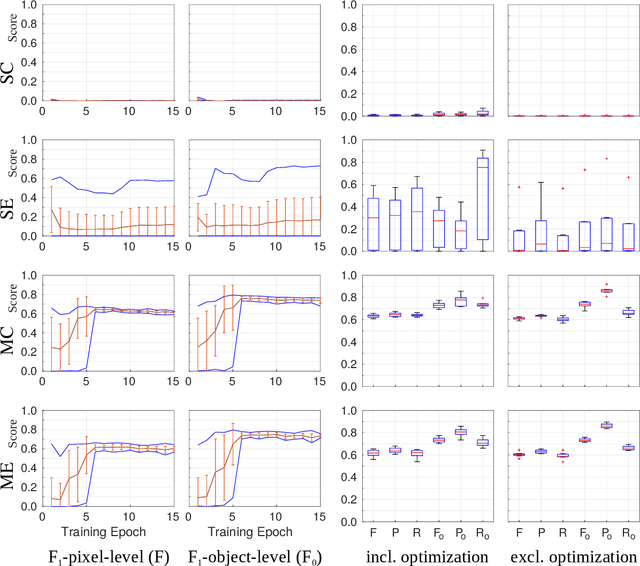
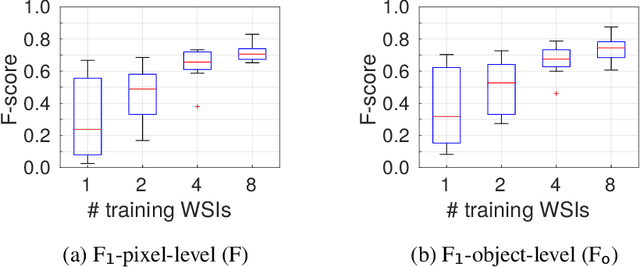
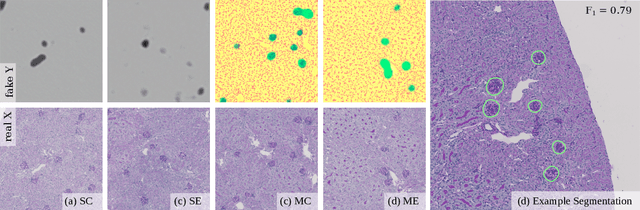
Abstract:Recently, generative adversarial networks exhibited excellent performances in semi-supervised image analysis scenarios. In this paper, we go even further by proposing a fully unsupervised approach for segmentation applications with prior knowledge of the objects' shapes. We propose and investigate different strategies to generate simulated label data and perform image-to-image translation between the image and the label domain using an adversarial model. Specifically, we assess the impact of the annotation model's accuracy as well as the effect of simulating additional low-level image features. For experimental evaluation, we consider the segmentation of the glomeruli, an application scenario from renal pathology. Experiments provide proof of concept and also confirm that the strategy for creating the simulated label data is of particular relevance considering the stability of GAN trainings.
CNN Cascades for Segmenting Whole Slide Images of the Kidney
Aug 01, 2017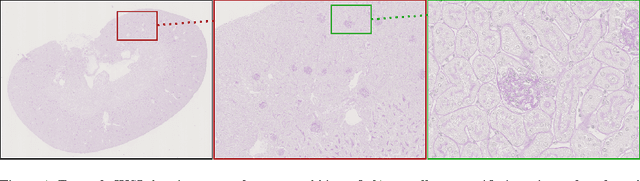
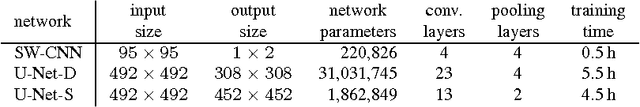

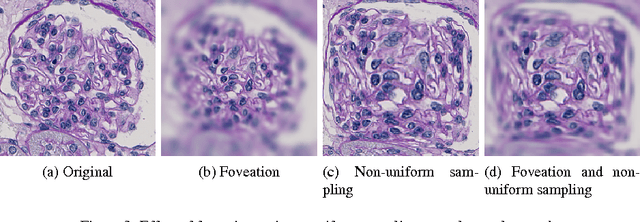
Abstract:Due to the increasing availability of whole slide scanners facilitating digitization of histopathological tissue, there is a strong demand for the development of computer based image analysis systems. In this work, the focus is on the segmentation of the glomeruli constituting a highly relevant structure in renal histopathology, which has not been investigated before in combination with CNNs. We propose two different CNN cascades for segmentation applications with sparse objects. These approaches are applied to the problem of glomerulus segmentation and compared with conventional fully-convolutional networks. Overall, with the best performing cascade approach, single CNNs are outperformed and a pixel-level Dice similarity coefficient of 0.90 is obtained. Combined with qualitative and further object-level analyses the obtained results are assessed as excellent also compared to recent approaches. In conclusion, we can state that especially one of the proposed cascade networks proved to be a highly powerful tool for segmenting the renal glomeruli providing best segmentation accuracies and also keeping the computing time at a low level.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge