Paul Duckworth
Force-Aware Neural Tangent Kernels for Scalable and Robust Active Learning of MLIPs
May 13, 2026Abstract:Active learning for machine-learning interatomic potentials (MLIPs) must address several challenges to be practical: scaling to large candidate pools, leveraging energy-force supervision, and maintaining robustness when candidate pools are biased relative to the target distribution. In this work, we jointly address these challenges. We first introduce a linearly scaling acquisition framework based on chunked feature-space posterior-variance shortlisting. By avoiding materialisation of the candidate and train set kernels, this approach enables screening of ~200k structures within hours and applies broadly to acquisition strategies that score candidates based on molecular similarity metrics. We then extend the Neural Tangent Kernel (NTK) to a force-aware setting via mixed parameter-coordinate derivatives, yielding a force NTK and a joint energy-force NTK that provide natural similarity metrics for vector-field prediction. We demonstrate the effectiveness of the joint energy-force NTK on the OC20 dataset, where force-aware acquisition is crucial: it achieves the lowest energy and force MAE and RMSE across all metrics and distribution splits. Across T1x, PMechDB, and RGD benchmarks, our force NTK methods remain competitive with established baselines while being significantly more efficient than committee-based approaches. Under a controlled candidate-pool shift case study on T1x, acquisition based on pretrained MLIP embeddings and NTKs remains robust, whereas committee-based methods exhibit higher variance. Overall, these results show that a single pretrained MLIP can enable scalable, force-aware, and distribution-robust active learning for foundation-model fine-tuning.
Pretrained Model Representations as Acquisition Signals for Active Learning of MLIPs
May 05, 2026Abstract:Training machine learning interatomic potentials (MLIPs) for reactive chemistry is often bottlenecked by the high cost of quantum chemical labels and the scarcity of transition state configurations in candidate pools. Active learning (AL) can mitigate these costs, but its effectiveness hinges on the acquisition rule. We investigate whether the latent space of a pretrained MLIP already contains the information necessary for effective acquisition, eliminating the need for auxiliary uncertainty heads, Bayesian training and fine-tuning, or committee ensembles. We introduce two acquisition signals derived directly from a pretrained MACE potential: a finite-width neural tangent kernel (NTK) and an activation kernel built from hidden latent space features. On reactive-chemistry benchmarks, both kernels consistently outperform fixed-descriptor baselines, committee disagreement, and random acquisition, reducing the data required to reach performance targets by an average of 38% for energy error and 28% for force error. We further show that the pretrained model induces similarity spaces that preserve chemically meaningful structure and provide more reliable residual uncertainty estimates than randomly initialised or fixed-descriptor-based kernels. Our results suggest that pretraining aligns latent-space geometry with model error, yielding a practical and sufficient acquisition signal for reactive MLIP fine-tuning.
Overconfident Oracles: Limitations of In Silico Sequence Design Benchmarking
Feb 24, 2025



Abstract:Machine learning methods can automate the in silico design of biological sequences, aiming to reduce costs and accelerate medical research. Given the limited access to wet labs, in silico design methods commonly use an oracle model to evaluate de novo generated sequences. However, the use of different oracle models across methods makes it challenging to compare them reliably, motivating the question: are in silico sequence design benchmarks reliable? In this work, we examine 12 sequence design methods that utilise ML oracles common in the literature and find that there are significant challenges with their cross-consistency and reproducibility. Indeed, oracles differing by architecture, or even just training seed, are shown to yield conflicting relative performance with our analysis suggesting poor out-of-distribution generalisation as a key issue. To address these challenges, we propose supplementing the evaluation with a suite of biophysical measures to assess the viability of generated sequences and limit out-of-distribution sequences the oracle is required to score, thereby improving the robustness of the design procedure. Our work aims to highlight potential pitfalls in the current evaluation process and contribute to the development of robust benchmarks, ultimately driving the improvement of in silico design methods.
Metalic: Meta-Learning In-Context with Protein Language Models
Oct 10, 2024



Abstract:Predicting the biophysical and functional properties of proteins is essential for in silico protein design. Machine learning has emerged as a promising technique for such prediction tasks. However, the relative scarcity of in vitro annotations means that these models often have little, or no, specific data on the desired fitness prediction task. As a result of limited data, protein language models (PLMs) are typically trained on general protein sequence modeling tasks, and then fine-tuned, or applied zero-shot, to protein fitness prediction. When no task data is available, the models make strong assumptions about the correlation between the protein sequence likelihood and fitness scores. In contrast, we propose meta-learning over a distribution of standard fitness prediction tasks, and demonstrate positive transfer to unseen fitness prediction tasks. Our method, called Metalic (Meta-Learning In-Context), uses in-context learning and fine-tuning, when data is available, to adapt to new tasks. Crucially, fine-tuning enables considerable generalization, even though it is not accounted for during meta-training. Our fine-tuned models achieve strong results with 18 times fewer parameters than state-of-the-art models. Moreover, our method sets a new state-of-the-art in low-data settings on ProteinGym, an established fitness-prediction benchmark. Due to data scarcity, we believe meta-learning will play a pivotal role in advancing protein engineering.
SMX: Sequential Monte Carlo Planning for Expert Iteration
Feb 12, 2024Abstract:Developing agents that can leverage planning abilities during their decision and learning processes is critical to the advancement of Artificial Intelligence. Recent works have demonstrated the effectiveness of combining tree-based search methods and self-play learning mechanisms. Yet, these methods typically face scaling challenges due to the sequential nature of their search. While practical engineering solutions can partly overcome this, they still demand extensive computational resources, which hinders their applicability. In this paper, we introduce SMX, a model-based planning algorithm that utilises scalable Sequential Monte Carlo methods to create an effective self-learning mechanism. Grounded in the theoretical framework of control as inference, SMX benefits from robust theoretical underpinnings. Its sampling-based search approach makes it adaptable to environments with both discrete and continuous action spaces. Furthermore, SMX allows for high parallelisation and can run on hardware accelerators to optimise computing efficiency. SMX demonstrates a statistically significant improvement in performance compared to AlphaZero, as well as demonstrating its performance as an improvement operator for a model-free policy, matching or exceeding top model-free methods across both continuous and discrete environments.
Are we going MAD? Benchmarking Multi-Agent Debate between Language Models for Medical Q&A
Nov 29, 2023



Abstract:Recent advancements in large language models (LLMs) underscore their potential for responding to medical inquiries. However, ensuring that generative agents provide accurate and reliable answers remains an ongoing challenge. In this context, multi-agent debate (MAD) has emerged as a prominent strategy for enhancing the truthfulness of LLMs. In this work, we provide a comprehensive benchmark of MAD strategies for medical Q&A, along with open-source implementations. This explores the effective utilization of various strategies including the trade-offs between cost, time, and accuracy. We build upon these insights to provide a novel debate-prompting strategy based on agent agreement that outperforms previously published strategies on medical Q&A tasks.
Jumanji: a Diverse Suite of Scalable Reinforcement Learning Environments in JAX
Jun 16, 2023Abstract:Open-source reinforcement learning (RL) environments have played a crucial role in driving progress in the development of AI algorithms. In modern RL research, there is a need for simulated environments that are performant, scalable, and modular to enable their utilization in a wider range of potential real-world applications. Therefore, we present Jumanji, a suite of diverse RL environments specifically designed to be fast, flexible, and scalable. Jumanji provides a suite of environments focusing on combinatorial problems frequently encountered in industry, as well as challenging general decision-making tasks. By leveraging the efficiency of JAX and hardware accelerators like GPUs and TPUs, Jumanji enables rapid iteration of research ideas and large-scale experimentation, ultimately empowering more capable agents. Unlike existing RL environment suites, Jumanji is highly customizable, allowing users to tailor the initial state distribution and problem complexity to their needs. Furthermore, we provide actor-critic baselines for each environment, accompanied by preliminary findings on scaling and generalization scenarios. Jumanji aims to set a new standard for speed, adaptability, and scalability of RL environments.
DITTO: Offline Imitation Learning with World Models
Feb 06, 2023


Abstract:We propose DITTO, an offline imitation learning algorithm which uses world models and on-policy reinforcement learning to addresses the problem of covariate shift, without access to an oracle or any additional online interactions. We discuss how world models enable offline, on-policy imitation learning, and propose a simple intrinsic reward defined in the world model latent space that induces imitation learning by reinforcement learning. Theoretically, we show that our formulation induces a divergence bound between expert and learner, in turn bounding the difference in reward. We test our method on difficult Atari environments from pixels alone, and achieve state-of-the-art performance in the offline setting.
Invariant Risk Minimisation for Cross-Organism Inference: Substituting Mouse Data for Human Data in Human Risk Factor Discovery
Nov 14, 2021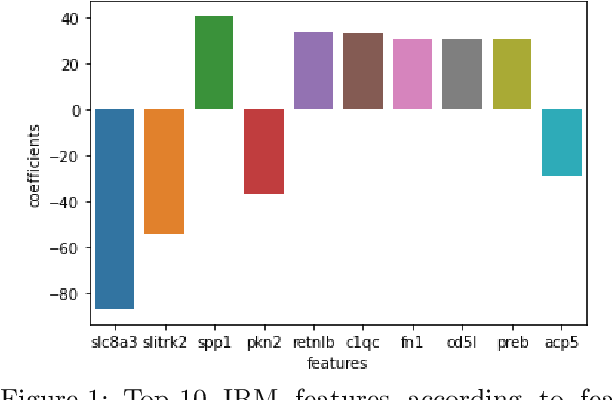
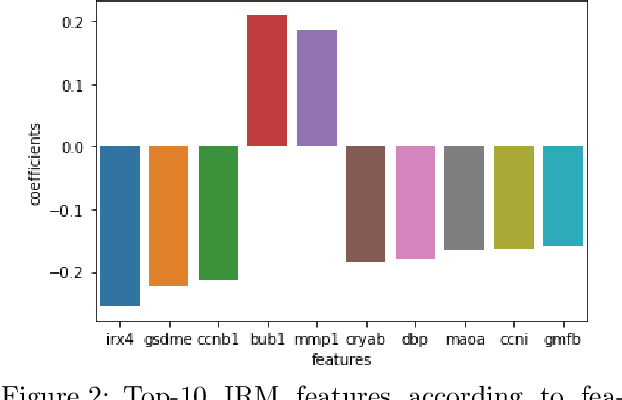
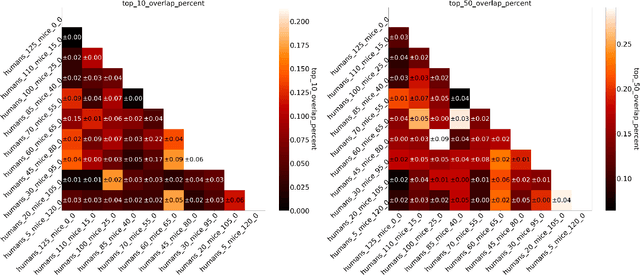
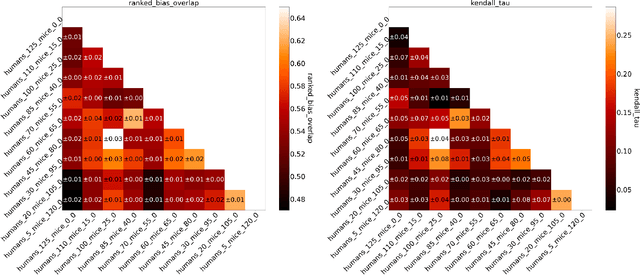
Abstract:Human medical data can be challenging to obtain due to data privacy concerns, difficulties conducting certain types of experiments, or prohibitive associated costs. In many settings, data from animal models or in-vitro cell lines are available to help augment our understanding of human data. However, this data is known for having low etiological validity in comparison to human data. In this work, we augment small human medical datasets with in-vitro data and animal models. We use Invariant Risk Minimisation (IRM) to elucidate invariant features by considering cross-organism data as belonging to different data-generating environments. Our models identify genes of relevance to human cancer development. We observe a degree of consistency between varying the amounts of human and mouse data used, however, further work is required to obtain conclusive insights. As a secondary contribution, we enhance existing open source datasets and provide two uniformly processed, cross-organism, homologue gene-matched datasets to the community.
Lexicographic Optimisation of Conditional Value at Risk and Expected Value for Risk-Averse Planning in MDPs
Oct 25, 2021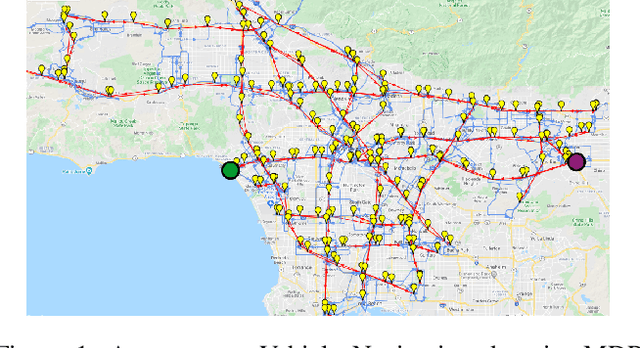
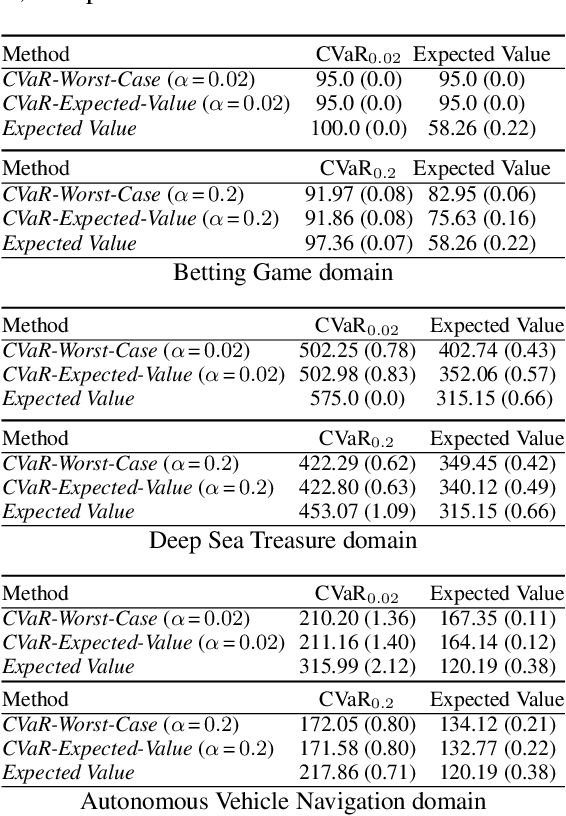
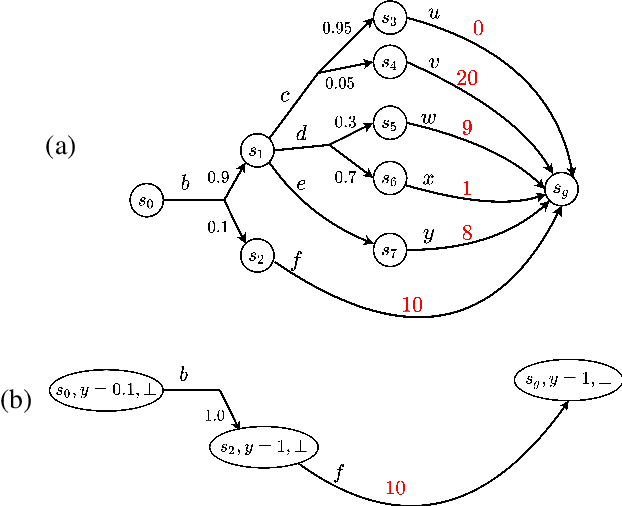
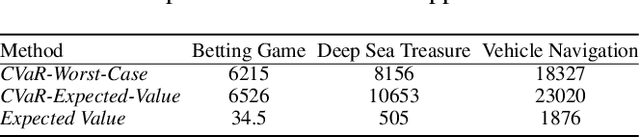
Abstract:Planning in Markov decision processes (MDPs) typically optimises the expected cost. However, optimising the expectation does not consider the risk that for any given run of the MDP, the total cost received may be unacceptably high. An alternative approach is to find a policy which optimises a risk-averse objective such as conditional value at risk (CVaR). In this work, we begin by showing that there can be multiple policies which obtain the optimal CVaR. We formulate the lexicographic optimisation problem of minimising the expected cost subject to the constraint that the CVaR of the total cost is optimal. We present an algorithm for this problem and evaluate our approach on three domains, including a road navigation domain based on real traffic data. Our experimental results demonstrate that our lexicographic approach attains improved expected cost while maintaining the optimal CVaR.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge