Ninon Burgos
for The Alzheimer's Disease Neuroimaging Initiative, APPRIMAGE Study Group
Multi-Domain Brain Vessel Segmentation Through Feature Disentanglement
Oct 02, 2025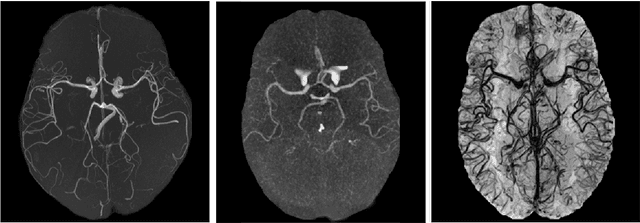


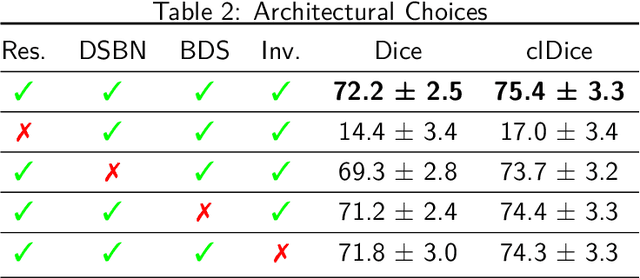
Abstract:The intricate morphology of brain vessels poses significant challenges for automatic segmentation models, which usually focus on a single imaging modality. However, accurately treating brain-related conditions requires a comprehensive understanding of the cerebrovascular tree, regardless of the specific acquisition procedure. Our framework effectively segments brain arteries and veins in various datasets through image-to-image translation while avoiding domain-specific model design and data harmonization between the source and the target domain. This is accomplished by employing disentanglement techniques to independently manipulate different image properties, allowing them to move from one domain to another in a label-preserving manner. Specifically, we focus on manipulating vessel appearances during adaptation while preserving spatial information, such as shapes and locations, which are crucial for correct segmentation. Our evaluation effectively bridges large and varied domain gaps across medical centers, image modalities, and vessel types. Additionally, we conduct ablation studies on the optimal number of required annotations and other architectural choices. The results highlight our framework's robustness and versatility, demonstrating the potential of domain adaptation methodologies to perform cerebrovascular image segmentation in multiple scenarios accurately. Our code is available at https://github.com/i-vesseg/MultiVesSeg.
* 19 pages, 7 figures, 3 tables. Joint first authors: Francesco Galati and Daniele Falcetta. Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2025:021. Code available at https://github.com/i-vesseg/MultiVesSeg
Unsupervised anomaly detection using Bayesian flow networks: application to brain FDG PET in the context of Alzheimer's disease
Jul 23, 2025Abstract:Unsupervised anomaly detection (UAD) plays a crucial role in neuroimaging for identifying deviations from healthy subject data and thus facilitating the diagnosis of neurological disorders. In this work, we focus on Bayesian flow networks (BFNs), a novel class of generative models, which have not yet been applied to medical imaging or anomaly detection. BFNs combine the strength of diffusion frameworks and Bayesian inference. We introduce AnoBFN, an extension of BFNs for UAD, designed to: i) perform conditional image generation under high levels of spatially correlated noise, and ii) preserve subject specificity by incorporating a recursive feedback from the input image throughout the generative process. We evaluate AnoBFN on the challenging task of Alzheimer's disease-related anomaly detection in FDG PET images. Our approach outperforms other state-of-the-art methods based on VAEs (beta-VAE), GANs (f-AnoGAN), and diffusion models (AnoDDPM), demonstrating its effectiveness at detecting anomalies while reducing false positive rates.
False Promises in Medical Imaging AI? Assessing Validity of Outperformance Claims
May 07, 2025Abstract:Performance comparisons are fundamental in medical imaging Artificial Intelligence (AI) research, often driving claims of superiority based on relative improvements in common performance metrics. However, such claims frequently rely solely on empirical mean performance. In this paper, we investigate whether newly proposed methods genuinely outperform the state of the art by analyzing a representative cohort of medical imaging papers. We quantify the probability of false claims based on a Bayesian approach that leverages reported results alongside empirically estimated model congruence to estimate whether the relative ranking of methods is likely to have occurred by chance. According to our results, the majority (>80%) of papers claims outperformance when introducing a new method. Our analysis further revealed a high probability (>5%) of false outperformance claims in 86% of classification papers and 53% of segmentation papers. These findings highlight a critical flaw in current benchmarking practices: claims of outperformance in medical imaging AI are frequently unsubstantiated, posing a risk of misdirecting future research efforts.
Confidence intervals uncovered: Are we ready for real-world medical imaging AI?
Sep 27, 2024



Abstract:Medical imaging is spearheading the AI transformation of healthcare. Performance reporting is key to determine which methods should be translated into clinical practice. Frequently, broad conclusions are simply derived from mean performance values. In this paper, we argue that this common practice is often a misleading simplification as it ignores performance variability. Our contribution is threefold. (1) Analyzing all MICCAI segmentation papers (n = 221) published in 2023, we first observe that more than 50% of papers do not assess performance variability at all. Moreover, only one (0.5%) paper reported confidence intervals (CIs) for model performance. (2) To address the reporting bottleneck, we show that the unreported standard deviation (SD) in segmentation papers can be approximated by a second-order polynomial function of the mean Dice similarity coefficient (DSC). Based on external validation data from 56 previous MICCAI challenges, we demonstrate that this approximation can accurately reconstruct the CI of a method using information provided in publications. (3) Finally, we reconstructed 95% CIs around the mean DSC of MICCAI 2023 segmentation papers. The median CI width was 0.03 which is three times larger than the median performance gap between the first and second ranked method. For more than 60% of papers, the mean performance of the second-ranked method was within the CI of the first-ranked method. We conclude that current publications typically do not provide sufficient evidence to support which models could potentially be translated into clinical practice.
Automated MRI Quality Assessment of Brain T1-weighted MRI in Clinical Data Warehouses: A Transfer Learning Approach Relying on Artefact Simulation
Jun 18, 2024



Abstract:The emergence of clinical data warehouses (CDWs), which contain the medical data of millions of patients, has paved the way for vast data sharing for research. The quality of MRIs gathered in CDWs differs greatly from what is observed in research settings and reflects a certain clinical reality. Consequently, a significant proportion of these images turns out to be unusable due to their poor quality. Given the massive volume of MRIs contained in CDWs, the manual rating of image quality is impossible. Thus, it is necessary to develop an automated solution capable of effectively identifying corrupted images in CDWs. This study presents an innovative transfer learning method for automated quality control of 3D gradient echo T1-weighted brain MRIs within a CDW, leveraging artefact simulation. We first intentionally corrupt images from research datasets by inducing poorer contrast, adding noise and introducing motion artefacts. Subsequently, three artefact-specific models are pre-trained using these corrupted images to detect distinct types of artefacts. Finally, the models are generalised to routine clinical data through a transfer learning technique, utilising 3660 manually annotated images. The overall image quality is inferred from the results of the three models, each designed to detect a specific type of artefact. Our method was validated on an independent test set of 385 3D gradient echo T1-weighted MRIs. Our proposed approach achieved excellent results for the detection of bad quality MRIs, with a balanced accuracy of over 87%, surpassing our previous approach by 3.5 percent points. Additionally, we achieved a satisfactory balanced accuracy of 79% for the detection of moderate quality MRIs, outperforming our previous performance by 5 percent points. Our framework provides a valuable tool for exploiting the potential of MRIs in CDWs.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2024:012
Evaluation of pseudo-healthy image reconstruction for anomaly detection with deep generative models: Application to brain FDG PET
Jan 29, 2024Abstract:Over the past years, pseudo-healthy reconstruction for unsupervised anomaly detection has gained in popularity. This approach has the great advantage of not requiring tedious pixel-wise data annotation and offers possibility to generalize to any kind of anomalies, including that corresponding to rare diseases. By training a deep generative model with only images from healthy subjects, the model will learn to reconstruct pseudo-healthy images. This pseudo-healthy reconstruction is then compared to the input to detect and localize anomalies. The evaluation of such methods often relies on a ground truth lesion mask that is available for test data, which may not exist depending on the application. We propose an evaluation procedure based on the simulation of realistic abnormal images to validate pseudo-healthy reconstruction methods when no ground truth is available. This allows us to extensively test generative models on different kinds of anomalies and measuring their performance using the pair of normal and abnormal images corresponding to the same subject. It can be used as a preliminary automatic step to validate the capacity of a generative model to reconstruct pseudo-healthy images, before a more advanced validation step that would require clinician's expertise. We apply this framework to the reconstruction of 3D brain FDG PET using a convolutional variational autoencoder with the aim to detect as early as possible the neurodegeneration markers that are specific to dementia such as Alzheimer's disease.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2024:003
Leveraging healthy population variability in deep learning unsupervised anomaly detection in brain FDG PET
Nov 20, 2023Abstract:Unsupervised anomaly detection is a popular approach for the analysis of neuroimaging data as it allows to identify a wide variety of anomalies from unlabelled data. It relies on building a subject-specific model of healthy appearance to which a subject's image can be compared to detect anomalies. In the literature, it is common for anomaly detection to rely on analysing the residual image between the subject's image and its pseudo-healthy reconstruction. This approach however has limitations partly due to the pseudo-healthy reconstructions being imperfect and to the lack of natural thresholding mechanism. Our proposed method, inspired by Z-scores, leverages the healthy population variability to overcome these limitations. Our experiments conducted on FDG PET scans from the ADNI database demonstrate the effectiveness of our approach in accurately identifying Alzheimer's disease related anomalies.
A2V: A Semi-Supervised Domain Adaptation Framework for Brain Vessel Segmentation via Two-Phase Training Angiography-to-Venography Translation
Sep 12, 2023



Abstract:We present a semi-supervised domain adaptation framework for brain vessel segmentation from different image modalities. Existing state-of-the-art methods focus on a single modality, despite the wide range of available cerebrovascular imaging techniques. This can lead to significant distribution shifts that negatively impact the generalization across modalities. By relying on annotated angiographies and a limited number of annotated venographies, our framework accomplishes image-to-image translation and semantic segmentation, leveraging a disentangled and semantically rich latent space to represent heterogeneous data and perform image-level adaptation from source to target domains. Moreover, we reduce the typical complexity of cycle-based architectures and minimize the use of adversarial training, which allows us to build an efficient and intuitive model with stable training. We evaluate our method on magnetic resonance angiographies and venographies. While achieving state-of-the-art performance in the source domain, our method attains a Dice score coefficient in the target domain that is only 8.9% lower, highlighting its promising potential for robust cerebrovascular image segmentation across different modalities.
Frequency Disentangled Learning for Segmentation of Midbrain Structures from Quantitative Susceptibility Mapping Data
Feb 25, 2023



Abstract:One often lacks sufficient annotated samples for training deep segmentation models. This is in particular the case for less common imaging modalities such as Quantitative Susceptibility Mapping (QSM). It has been shown that deep models tend to fit the target function from low to high frequencies. One may hypothesize that such property can be leveraged for better training of deep learning models. In this paper, we exploit this property to propose a new training method based on frequency-domain disentanglement. It consists of two main steps: i) disentangling the image into high- and low-frequency parts and feature learning; ii) frequency-domain fusion to complete the task. The approach can be used with any backbone segmentation network. We apply the approach to the segmentation of the red and dentate nuclei from QSM data which is particularly relevant for the study of parkinsonian syndromes. We demonstrate that the proposed method provides considerable performance improvements for these tasks. We further applied it to three public datasets from the Medical Segmentation Decathlon (MSD) challenge. For two MSD tasks, it provided smaller but still substantial improvements (up to 7 points of Dice), especially under small training set situations.
Fourier Disentangled Multimodal Prior Knowledge Fusion for Red Nucleus Segmentation in Brain MRI
Nov 02, 2022



Abstract:Early and accurate diagnosis of parkinsonian syndromes is critical to provide appropriate care to patients and for inclusion in therapeutic trials. The red nucleus is a structure of the midbrain that plays an important role in these disorders. It can be visualized using iron-sensitive magnetic resonance imaging (MRI) sequences. Different iron-sensitive contrasts can be produced with MRI. Combining such multimodal data has the potential to improve segmentation of the red nucleus. Current multimodal segmentation algorithms are computationally consuming, cannot deal with missing modalities and need annotations for all modalities. In this paper, we propose a new model that integrates prior knowledge from different contrasts for red nucleus segmentation. The method consists of three main stages. First, it disentangles the image into high-level information representing the brain structure, and low-frequency information representing the contrast. The high-frequency information is then fed into a network to learn anatomical features, while the list of multimodal low-frequency information is processed by another module. Finally, feature fusion is performed to complete the segmentation task. The proposed method was used with several iron-sensitive contrasts (iMag, QSM, R2*, SWI). Experiments demonstrate that our proposed model substantially outperforms a baseline UNet model when the training set size is very small.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge