Muriel Medard
MultiTok: Variable-Length Tokenization for Efficient LLMs Adapted from LZW Compression
Oct 28, 2024



Abstract:Large language models have drastically changed the prospects of AI by introducing technologies for more complex natural language processing. However, current methodologies to train such LLMs require extensive resources including but not limited to large amounts of data, expensive machinery, and lengthy training. To solve this problem, this paper proposes a new tokenization method inspired by universal Lempel-Ziv-Welch data compression that compresses repetitive phrases into multi-word tokens. With MultiTok as a new tokenizing tool, we show that language models are able to be trained notably more efficiently while offering a similar accuracy on more succinct and compressed training data. In fact, our results demonstrate that MultiTok achieves a comparable performance to the BERT standard as a tokenizer while also providing close to 2.5x faster training with more than 30% less training data.
Successive Refinement in Large-Scale Computation: Advancing Model Inference Applications
Feb 11, 2024Abstract:Modern computationally-intensive applications often operate under time constraints, necessitating acceleration methods and distribution of computational workloads across multiple entities. However, the outcome is either achieved within the desired timeline or not, and in the latter case, valuable resources are wasted. In this paper, we introduce solutions for layered-resolution computation. These solutions allow lower-resolution results to be obtained at an earlier stage than the final result. This innovation notably enhances the deadline-based systems, as if a computational job is terminated due to time constraints, an approximate version of the final result can still be generated. Moreover, in certain operational regimes, a high-resolution result might be unnecessary, because the low-resolution result may already deviate significantly from the decision threshold, for example in AI-based decision-making systems. Therefore, operators can decide whether higher resolution is needed or not based on intermediate results, enabling computations with adaptive resolution. We present our framework for two critical and computationally demanding jobs: distributed matrix multiplication (linear) and model inference in machine learning (nonlinear). Our theoretical and empirical results demonstrate that the execution delay for the first resolution is significantly shorter than that for the final resolution, while maintaining overall complexity comparable to the conventional one-shot approach. Our experiments further illustrate how the layering feature increases the likelihood of meeting deadlines and enables adaptability and transparency in massive, large-scale computations.
TexShape: Information Theoretic Sentence Embedding for Language Models
Feb 05, 2024



Abstract:With the exponential growth in data volume and the emergence of data-intensive applications, particularly in the field of machine learning, concerns related to resource utilization, privacy, and fairness have become paramount. This paper focuses on the textual domain of data and addresses challenges regarding encoding sentences to their optimized representations through the lens of information-theory. In particular, we use empirical estimates of mutual information, using the Donsker-Varadhan definition of Kullback-Leibler divergence. Our approach leverages this estimation to train an information-theoretic sentence embedding, called TexShape, for (task-based) data compression or for filtering out sensitive information, enhancing privacy and fairness. In this study, we employ a benchmark language model for initial text representation, complemented by neural networks for information-theoretic compression and mutual information estimations. Our experiments demonstrate significant advancements in preserving maximal targeted information and minimal sensitive information over adverse compression ratios, in terms of predictive accuracy of downstream models that are trained using the compressed data.
Analog Compressed Sensing for Sparse Frequency Shift Keying Modulation Schemes
May 31, 2022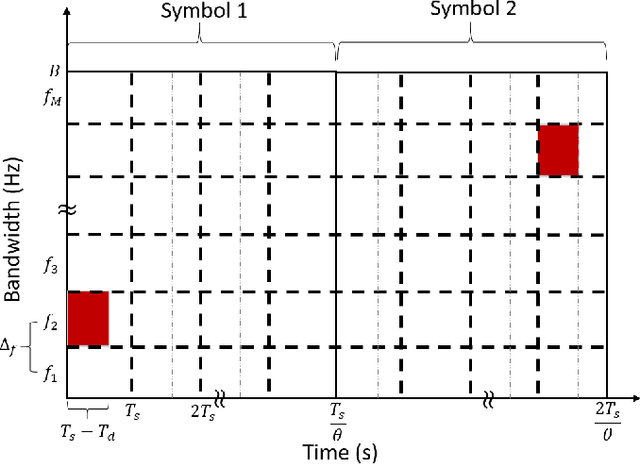
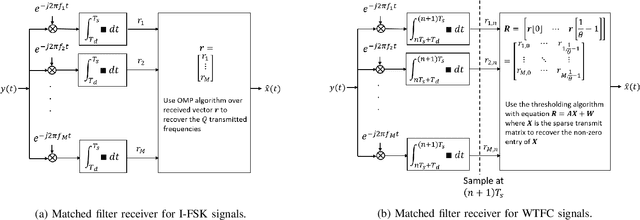
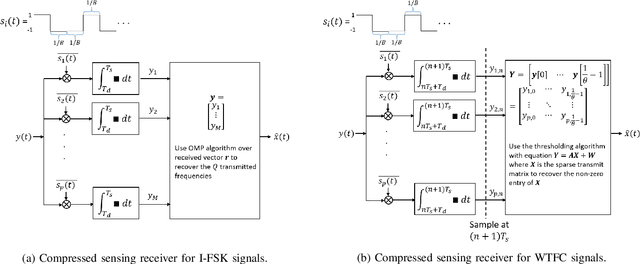
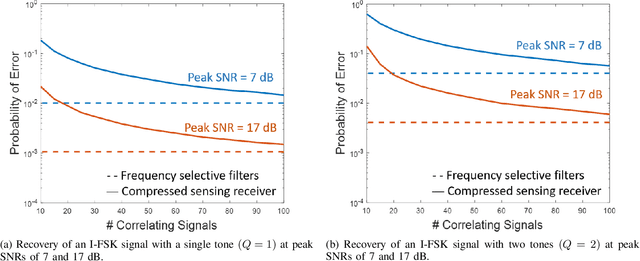
Abstract:There is a growing interest in signaling schemes that operate in the wideband regime due to the crowded frequency spectrum. However, a downside of the wideband regime is that obtaining channel state information is costly, and the capacity of previously used modulation schemes such as code division multiple access and orthogonal frequency division multiplexing begins to diverge from the capacity bound without channel state information. Impulsive frequency shift keying and wideband time frequency coding have been shown to perform well in the wideband regime without channel state information, thus avoiding the costs and challenges associated with obtaining channel state information. However, the maximum likelihood receiver is a bank of frequency-selective filters, which is very costly to implement due to the large number of filters. In this work, we aim to simplify the receiver by using an analog compressed sensing receiver with chipping sequences as correlating signals to detect the sparse signals. Our results show that using a compressed sensing receiver allows for the simplification of the analog receiver with the trade off of a slight degradation in recovery performance. For a fixed frequency separation, symbol time, and peak SNR, the performance loss remains the same for a fixed ratio of number of correlating signals to the number of frequencies.
Wideband Time Frequency Coding
May 31, 2022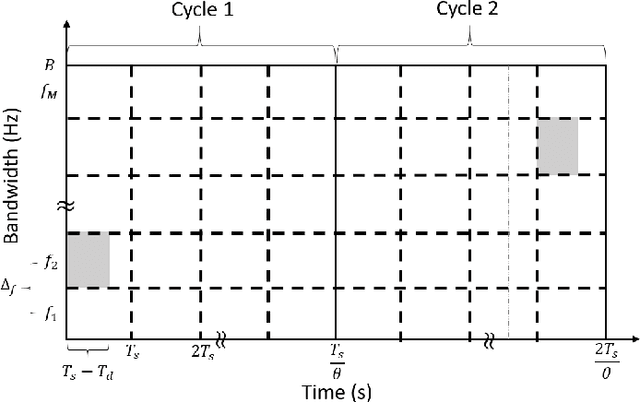
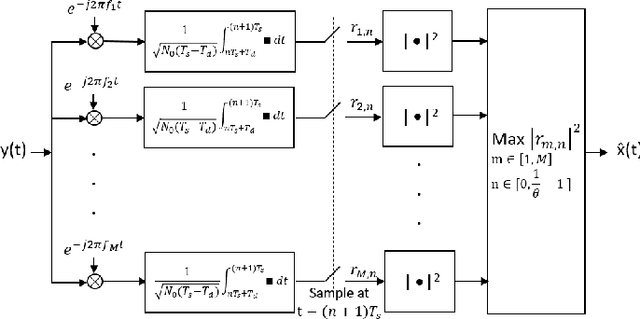
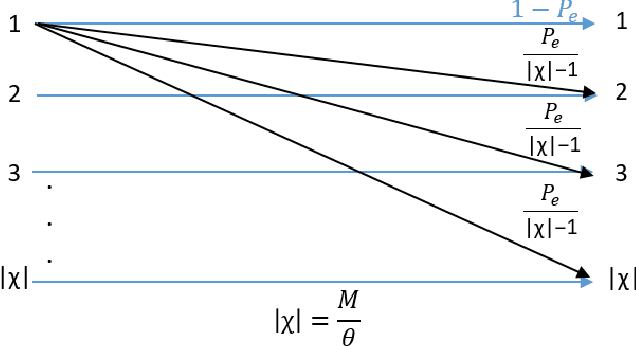
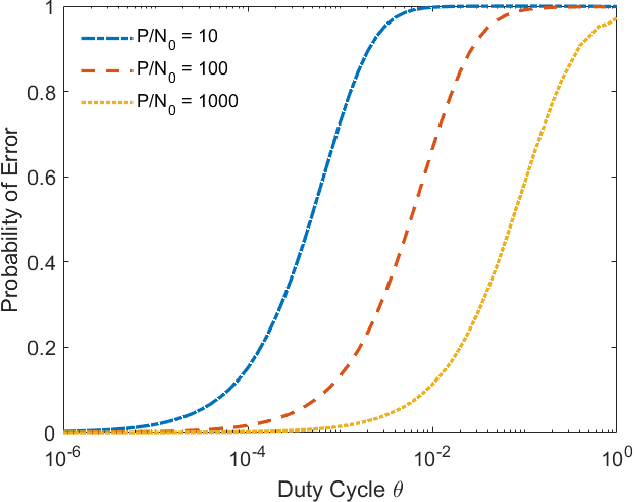
Abstract:In the wideband regime, the performance of many of the popular modulation schemes such as code division multiple access and orthogonal frequency division multiplexing falls quickly without channel state information. Obtaining the amount of channel information required for these techniques to work is costly and difficult, which suggests the need for schemes which can perform well without channel state information. In this work, we present one such scheme, called wideband time frequency coding, which achieves rates on the order of the additive white Gaussian noise capacity without requiring any channel state information. Wideband time frequency coding combines impulsive frequency shift keying with pulse position modulation, which allows for information to be encoded in both the transmitted frequency and the transmission time period. On the detection side, we propose a non-coherent decoder based on a square-law detector, akin to the optimal decoder for frequency shift keying based signals. The impacts of various parameters on the symbol error probability and capacity of wideband time frequency coding are investigated, and the results show that it is robust to shadowing and highly fading channels. When compared to other modulation schemes such as code division multiple access, orthogonal frequency division multiplexing, pulse position modulation, and impulsive frequency shift keying without channel state information, wideband time frequency coding achieves higher rates in the wideband regime, and performs comparably in smaller bandwidths.
NeuraCrypt: Hiding Private Health Data via Random Neural Networks for Public Training
Jun 04, 2021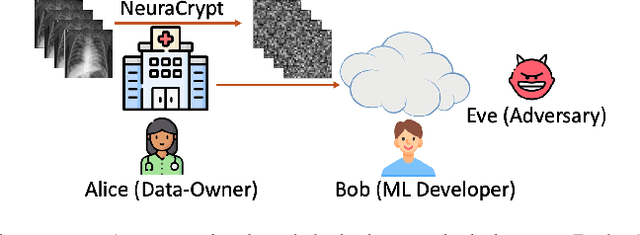
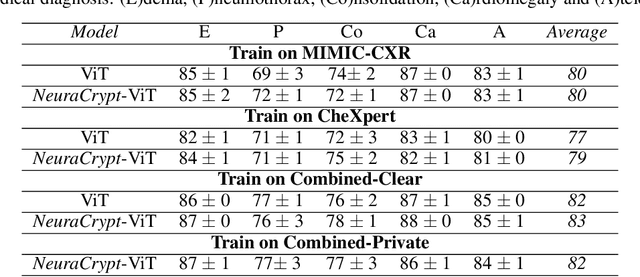
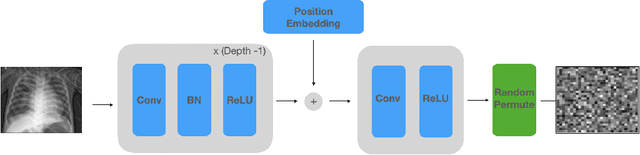

Abstract:Balancing the needs of data privacy and predictive utility is a central challenge for machine learning in healthcare. In particular, privacy concerns have led to a dearth of public datasets, complicated the construction of multi-hospital cohorts and limited the utilization of external machine learning resources. To remedy this, new methods are required to enable data owners, such as hospitals, to share their datasets publicly, while preserving both patient privacy and modeling utility. We propose NeuraCrypt, a private encoding scheme based on random deep neural networks. NeuraCrypt encodes raw patient data using a randomly constructed neural network known only to the data-owner, and publishes both the encoded data and associated labels publicly. From a theoretical perspective, we demonstrate that sampling from a sufficiently rich family of encoding functions offers a well-defined and meaningful notion of privacy against a computationally unbounded adversary with full knowledge of the underlying data-distribution. We propose to approximate this family of encoding functions through random deep neural networks. Empirically, we demonstrate the robustness of our encoding to a suite of adversarial attacks and show that NeuraCrypt achieves competitive accuracy to non-private baselines on a variety of x-ray tasks. Moreover, we demonstrate that multiple hospitals, using independent private encoders, can collaborate to train improved x-ray models. Finally, we release a challenge dataset to encourage the development of new attacks on NeuraCrypt.
A Coding Theory Perspective on Multiplexed Molecular Profiling of Biological Tissues
Feb 02, 2021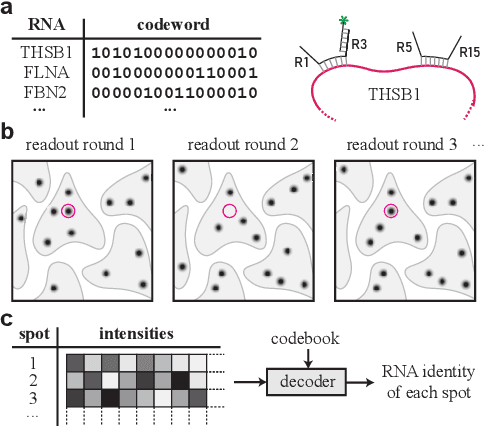

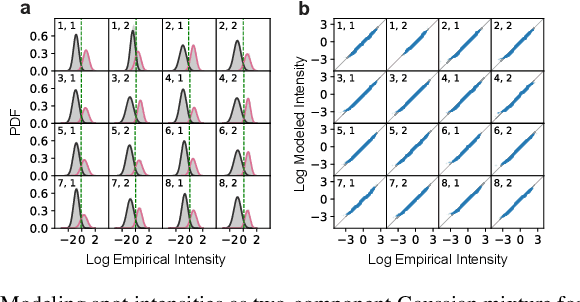
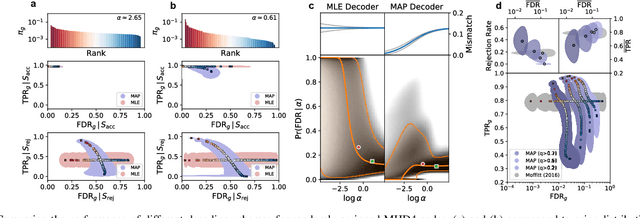
Abstract:High-throughput and quantitative experimental technologies are experiencing rapid advances in the biological sciences. One important recent technique is multiplexed fluorescence in situ hybridization (mFISH), which enables the identification and localization of large numbers of individual strands of RNA within single cells. Core to that technology is a coding problem: with each RNA sequence of interest being a codeword, how to design a codebook of probes, and how to decode the resulting noisy measurements? Published work has relied on assumptions of uniformly distributed codewords and binary symmetric channels for decoding and to a lesser degree for code construction. Here we establish that both of these assumptions are inappropriate in the context of mFISH experiments and substantial decoding performance gains can be obtained by using more appropriate, less classical, assumptions. We propose a more appropriate asymmetric channel model that can be readily parameterized from data and use it to develop a maximum a posteriori (MAP) decoders. We show that false discovery rate for rare RNAs, which is the key experimental metric, is vastly improved with MAP decoders even when employed with the existing sub-optimal codebook. Using an evolutionary optimization methodology, we further show that by permuting the codebook to better align with the prior, which is an experimentally straightforward procedure, significant further improvements are possible.
Network Maximal Correlation
Feb 09, 2017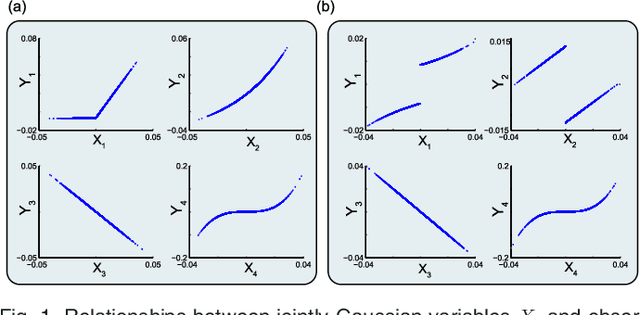
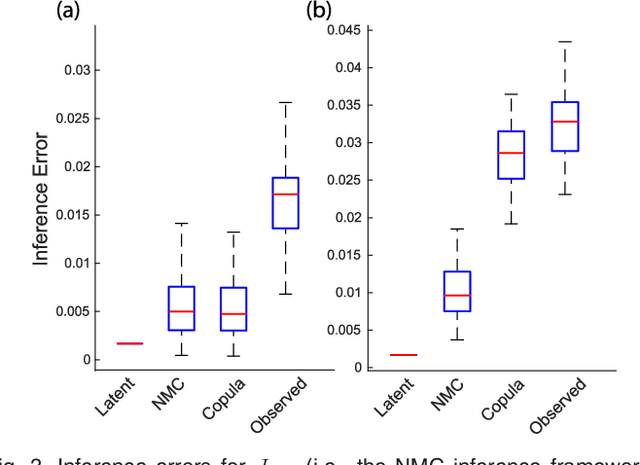
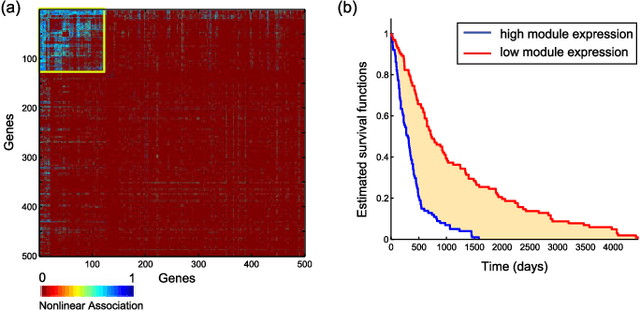
Abstract:We introduce Network Maximal Correlation (NMC) as a multivariate measure of nonlinear association among random variables. NMC is defined via an optimization that infers transformations of variables by maximizing aggregate inner products between transformed variables. For finite discrete and jointly Gaussian random variables, we characterize a solution of the NMC optimization using basis expansion of functions over appropriate basis functions. For finite discrete variables, we propose an algorithm based on alternating conditional expectation to determine NMC. Moreover we propose a distributed algorithm to compute an approximation of NMC for large and dense graphs using graph partitioning. For finite discrete variables, we show that the probability of discrepancy greater than any given level between NMC and NMC computed using empirical distributions decays exponentially fast as the sample size grows. For jointly Gaussian variables, we show that under some conditions the NMC optimization is an instance of the Max-Cut problem. We then illustrate an application of NMC in inference of graphical model for bijective functions of jointly Gaussian variables. Finally, we show NMC's utility in a data application of learning nonlinear dependencies among genes in a cancer dataset.
A Doubly Distributed Genetic Algorithm for Network Coding
Apr 10, 2007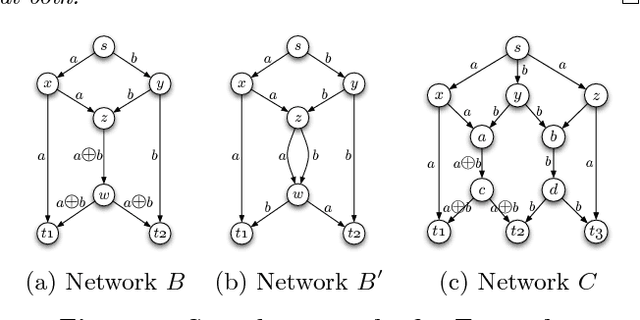
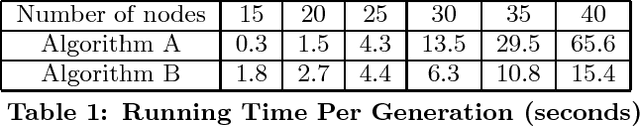
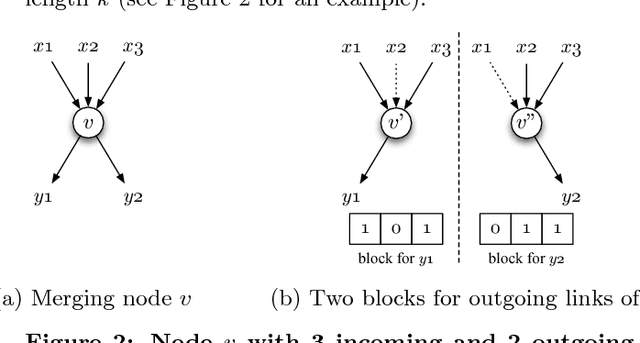
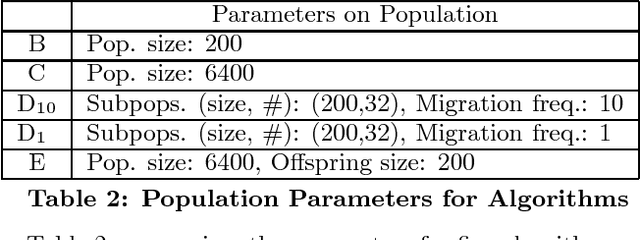
Abstract:We present a genetic algorithm which is distributed in two novel ways: along genotype and temporal axes. Our algorithm first distributes, for every member of the population, a subset of the genotype to each network node, rather than a subset of the population to each. This genotype distribution is shown to offer a significant gain in running time. Then, for efficient use of the computational resources in the network, our algorithm divides the candidate solutions into pipelined sets and thus the distribution is in the temporal domain, rather that in the spatial domain. This temporal distribution may lead to temporal inconsistency in selection and replacement, however our experiments yield better efficiency in terms of the time to convergence without incurring significant penalties.
Genetic Representations for Evolutionary Minimization of Network Coding Resources
Feb 07, 2007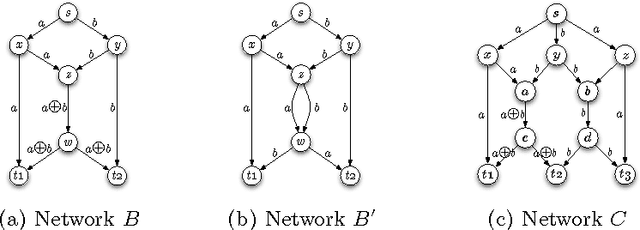
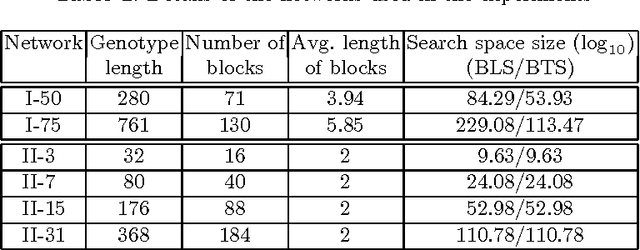
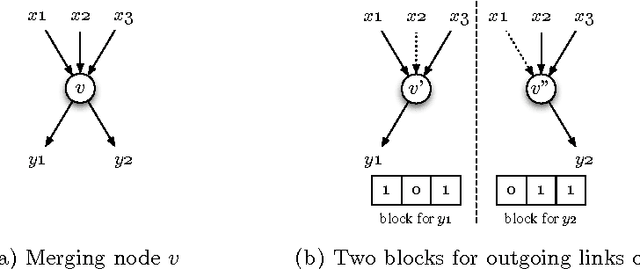

Abstract:We demonstrate how a genetic algorithm solves the problem of minimizing the resources used for network coding, subject to a throughput constraint, in a multicast scenario. A genetic algorithm avoids the computational complexity that makes the problem NP-hard and, for our experiments, greatly improves on sub-optimal solutions of established methods. We compare two different genotype encodings, which tradeoff search space size with fitness landscape, as well as the associated genetic operators. Our finding favors a smaller encoding despite its fewer intermediate solutions and demonstrates the impact of the modularity enforced by genetic operators on the performance of the algorithm.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge