Mingren Shen
Machine learning for interpreting coherent X-ray speckle patterns
Nov 15, 2022Abstract:Speckle patterns produced by coherent X-ray have a close relationship with the internal structure of materials but quantitative inversion of the relationship to determine structure from images is challenging. Here, we investigate the link between coherent X-ray speckle patterns and sample structures using a model 2D disk system and explore the ability of machine learning to learn aspects of the relationship. Specifically, we train a deep neural network to classify the coherent X-ray speckle pattern images according to the disk number density in the corresponding structure. It is demonstrated that the classification system is accurate for both non-disperse and disperse size distributions.
Performance, Successes and Limitations of Deep Learning Semantic Segmentation of Multiple Defects in Transmission Electron Micrographs
Oct 15, 2021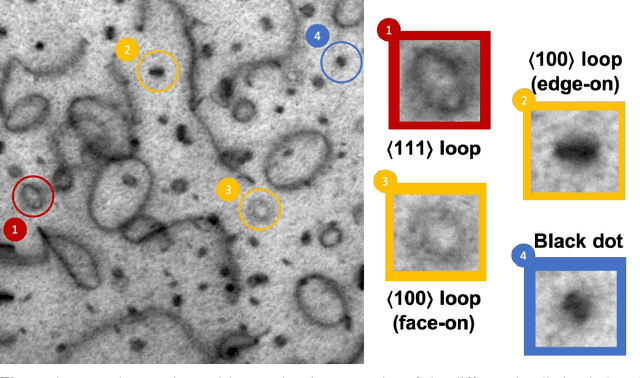
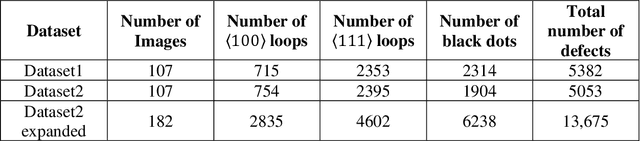
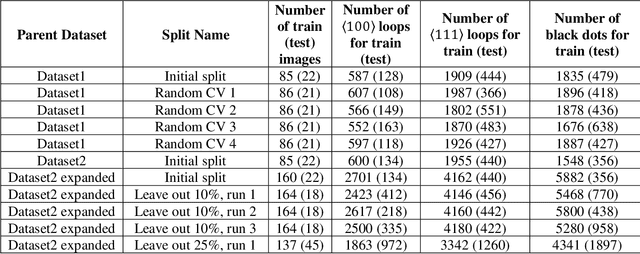
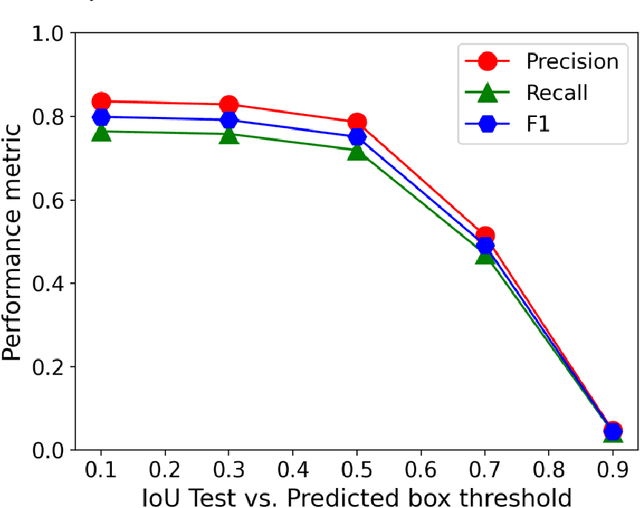
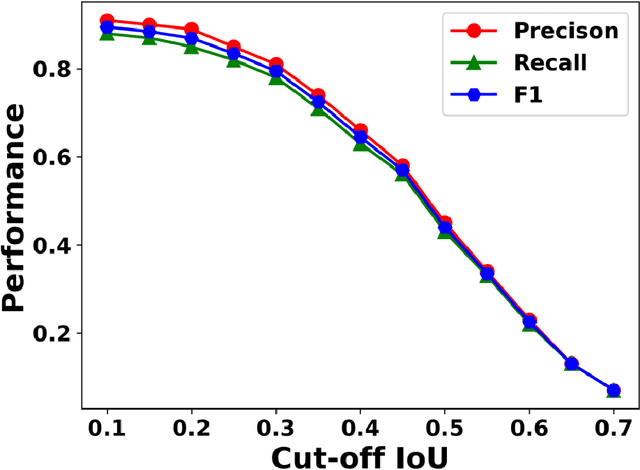
Abstract:In this work, we perform semantic segmentation of multiple defect types in electron microscopy images of irradiated FeCrAl alloys using a deep learning Mask Regional Convolutional Neural Network (Mask R-CNN) model. We conduct an in-depth analysis of key model performance statistics, with a focus on quantities such as predicted distributions of defect shapes, defect sizes, and defect areal densities relevant to informing modeling and understanding of irradiated Fe-based materials properties. To better understand the performance and present limitations of the model, we provide examples of useful evaluation tests which include a suite of random splits, and dataset size-dependent and domain-targeted cross validation tests. Overall, we find that the current model is a fast, effective tool for automatically characterizing and quantifying multiple defect types in microscopy images, with a level of accuracy on par with human domain expert labelers. More specifically, the model can achieve average defect identification F1 scores as high as 0.8, and, based on random cross validation, have low overall average (+/- standard deviation) defect size and density percentage errors of 7.3 (+/- 3.8)% and 12.7 (+/- 5.3)%, respectively. Further, our model predicts the expected material hardening to within 10-20 MPa (about 10% of total hardening), which is about the same error level as experiments. Our targeted evaluation tests also suggest the best path toward improving future models is not expanding existing databases with more labeled images but instead data additions that target weak points of the model domain, such as images from different microscopes, imaging conditions, irradiation environments, and alloy types. Finally, we discuss the first phase of an effort to provide an easy-to-use, open-source object detection tool to the broader community for identifying defects in new images.
Multi defect detection and analysis of electron microscopy images with deep learning
Aug 19, 2021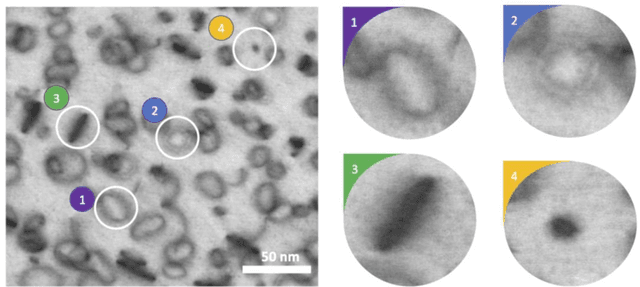
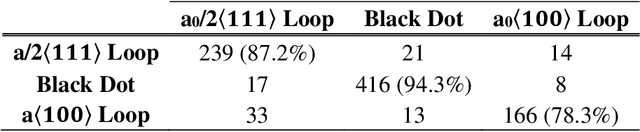

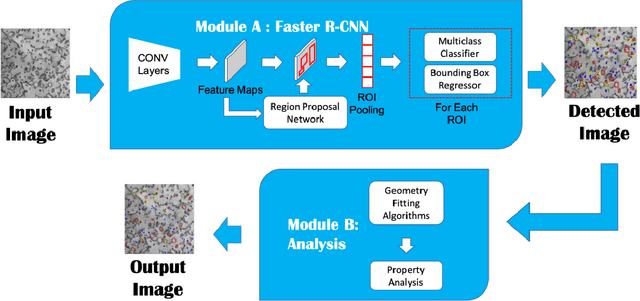

Abstract:Electron microscopy is widely used to explore defects in crystal structures, but human detecting of defects is often time-consuming, error-prone, and unreliable, and is not scalable to large numbers of images or real-time analysis. In this work, we discuss the application of machine learning approaches to find the location and geometry of different defect clusters in irradiated steels. We show that a deep learning based Faster R-CNN analysis system has a performance comparable to human analysis with relatively small training data sets. This study proves the promising ability to apply deep learning to assist the development of automated microscopy data analysis even when multiple features are present and paves the way for fast, scalable, and reliable analysis systems for massive amounts of modern electron microscopy data.
A Deep Learning Based Automatic Defect Analysis Framework for In-situ TEM Ion Irradiations
Aug 19, 2021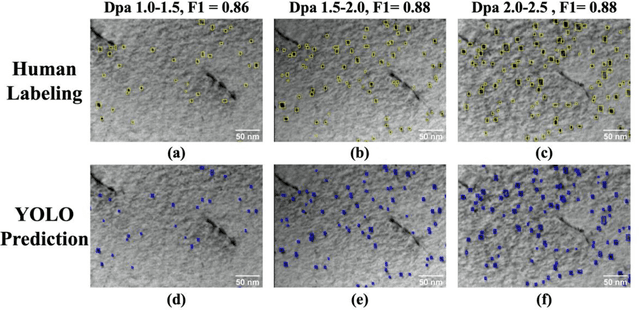
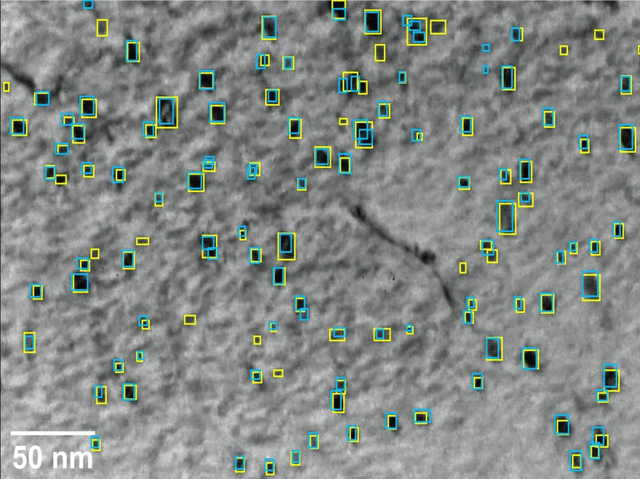


Abstract:Videos captured using Transmission Electron Microscopy (TEM) can encode details regarding the morphological and temporal evolution of a material by taking snapshots of the microstructure sequentially. However, manual analysis of such video is tedious, error-prone, unreliable, and prohibitively time-consuming if one wishes to analyze a significant fraction of frames for even videos of modest length. In this work, we developed an automated TEM video analysis system for microstructural features based on the advanced object detection model called YOLO and tested the system on an in-situ ion irradiation TEM video of dislocation loops formed in a FeCrAl alloy. The system provides analysis of features observed in TEM including both static and dynamic properties using the YOLO-based defect detection module coupled to a geometry analysis module and a dynamic tracking module. Results show that the system can achieve human comparable performance with an F1 score of 0.89 for fast, consistent, and scalable frame-level defect analysis. This result is obtained on a real but exceptionally clean and stable data set and more challenging data sets may not achieve this performance. The dynamic tracking also enabled evaluation of individual defect evolution like per defect growth rate at a fidelity never before achieved using common human analysis methods. Our work shows that automatically detecting and tracking interesting microstructures and properties contained in TEM videos is viable and opens new doors for evaluating materials dynamics.
Exploring Generative Adversarial Networks for Image-to-Image Translation in STEM Simulation
Oct 29, 2020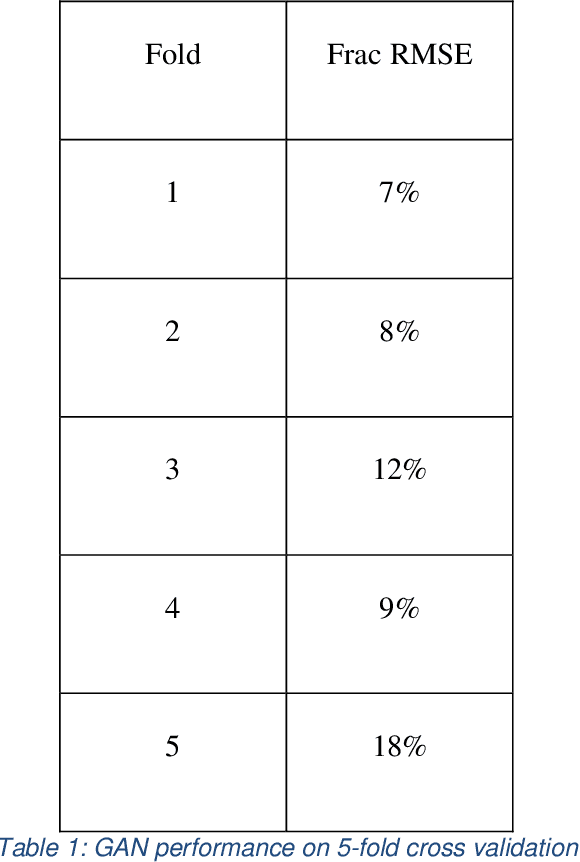
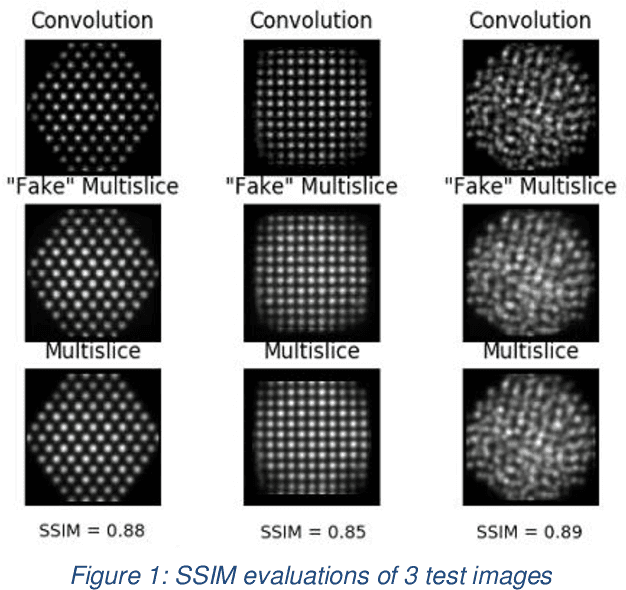

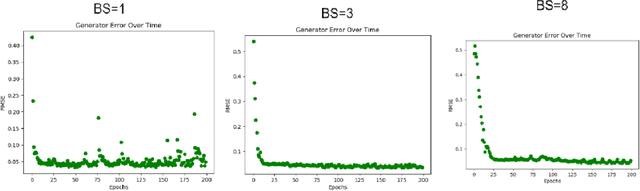
Abstract:The use of accurate scanning transmission electron microscopy (STEM) image simulation methods require large computation times that can make their use infeasible for the simulation of many images. Other simulation methods based on linear imaging models, such as the convolution method, are much faster but are too inaccurate to be used in application. In this paper, we explore deep learning models that attempt to translate a STEM image produced by the convolution method to a prediction of the high accuracy multislice image. We then compare our results to those of regression methods. We find that using the deep learning model Generative Adversarial Network (GAN) provides us with the best results and performs at a similar accuracy level to previous regression models on the same dataset. Codes and data for this project can be found in this GitHub repository, https://github.com/uw-cmg/GAN-STEM-Conv2MultiSlice.
Assessing Graph-based Deep Learning Models for Predicting Flash Point
Feb 26, 2020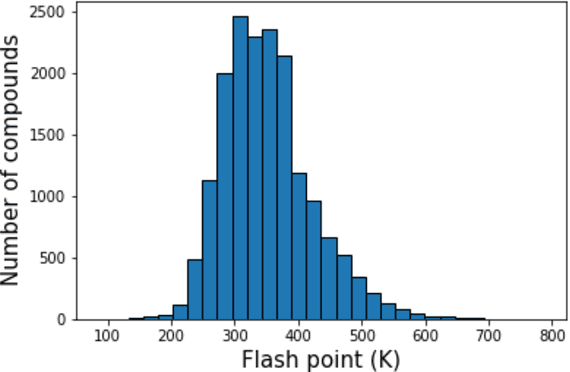
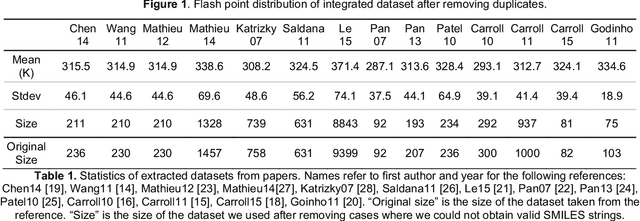
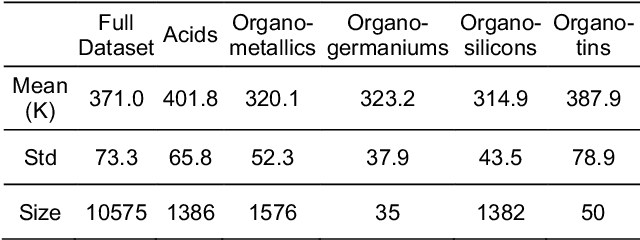
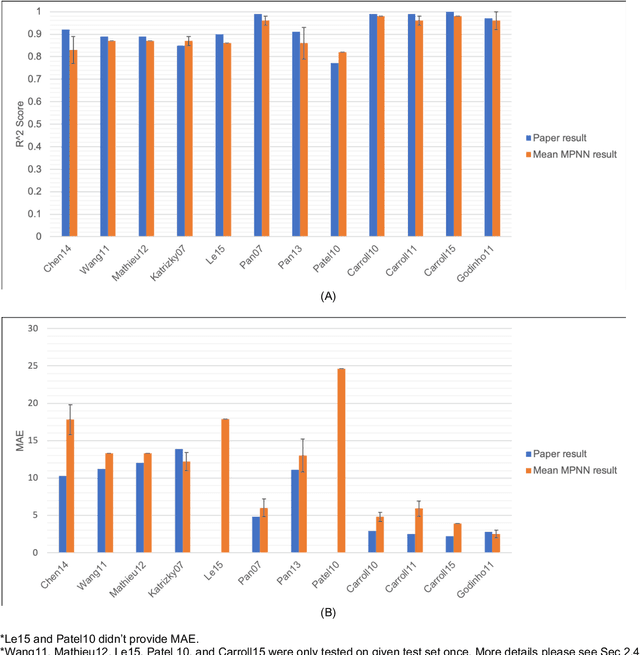
Abstract:Flash points of organic molecules play an important role in preventing flammability hazards and large databases of measured values exist, although millions of compounds remain unmeasured. To rapidly extend existing data to new compounds many researchers have used quantitative structure-property relationship (QSPR) analysis to effectively predict flash points. In recent years graph-based deep learning (GBDL) has emerged as a powerful alternative method to traditional QSPR. In this paper, GBDL models were implemented in predicting flash point for the first time. We assessed the performance of two GBDL models, message-passing neural network (MPNN) and graph convolutional neural network (GCNN), by comparing methods. Our result shows that MPNN both outperforms GCNN and yields slightly worse but comparable performance with previous QSPR studies. The average R2 and Mean Absolute Error (MAE) scores of MPNN are, respectively, 2.3% lower and 2.0 K higher than previous comparable studies. To further explore GBDL models, we collected the largest flash point dataset to date, which contains 10575 unique molecules. The optimized MPNN gives a test data R2 of 0.803 and MAE of 17.8 K on the complete dataset. We also extracted 5 datasets from our integrated dataset based on molecular types (acids, organometallics, organogermaniums, organosilicons, and organotins) and explore the quality of the model in these classes.against 12 previous QSPR studies using more traditional
* 26 pages, 6 tabels, 3 figures
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge