Michael A. Riegler
DivergentNets: Medical Image Segmentation by Network Ensemble
Jul 01, 2021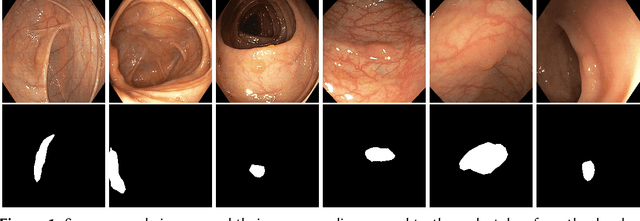
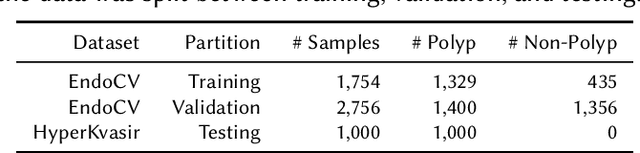
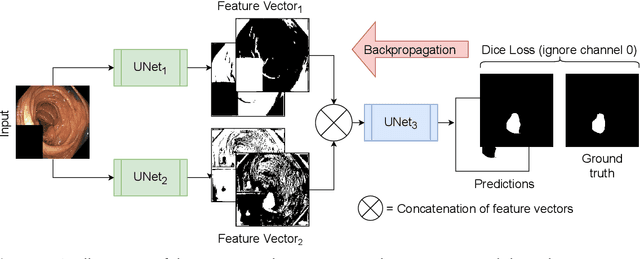
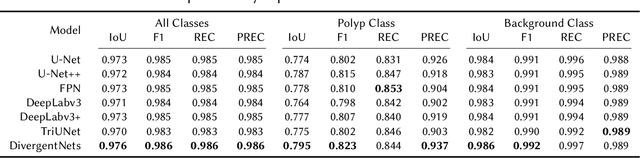
Abstract:Detection of colon polyps has become a trending topic in the intersecting fields of machine learning and gastrointestinal endoscopy. The focus has mainly been on per-frame classification. More recently, polyp segmentation has gained attention in the medical community. Segmentation has the advantage of being more accurate than per-frame classification or object detection as it can show the affected area in greater detail. For our contribution to the EndoCV 2021 segmentation challenge, we propose two separate approaches. First, a segmentation model named TriUNet composed of three separate UNet models. Second, we combine TriUNet with an ensemble of well-known segmentation models, namely UNet++, FPN, DeepLabv3, and DeepLabv3+, into a model called DivergentNets to produce more generalizable medical image segmentation masks. In addition, we propose a modified Dice loss that calculates loss only for a single class when performing multiclass segmentation, forcing the model to focus on what is most important. Overall, the proposed methods achieved the best average scores for each respective round in the challenge, with TriUNet being the winning model in Round I and DivergentNets being the winning model in Round II of the segmentation generalization challenge at EndoCV 2021. The implementation of our approach is made publicly available on GitHub.
* the winning model of the segmentation generalization challenge at EndoCV 2021
SinGAN-Seg: Synthetic Training Data Generation for Medical Image Segmentation
Jun 29, 2021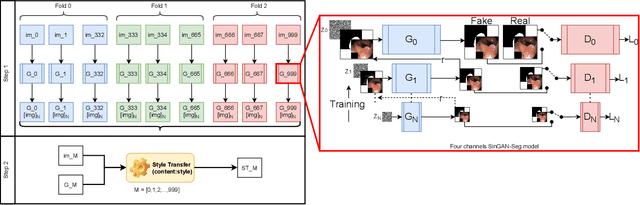
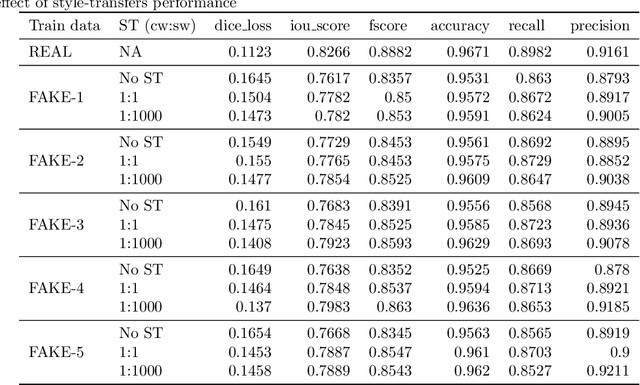
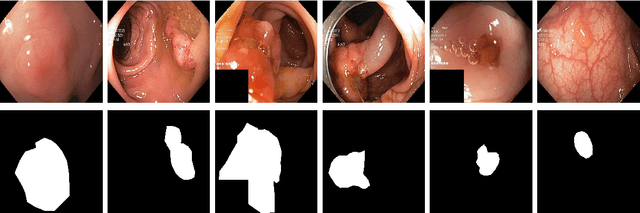
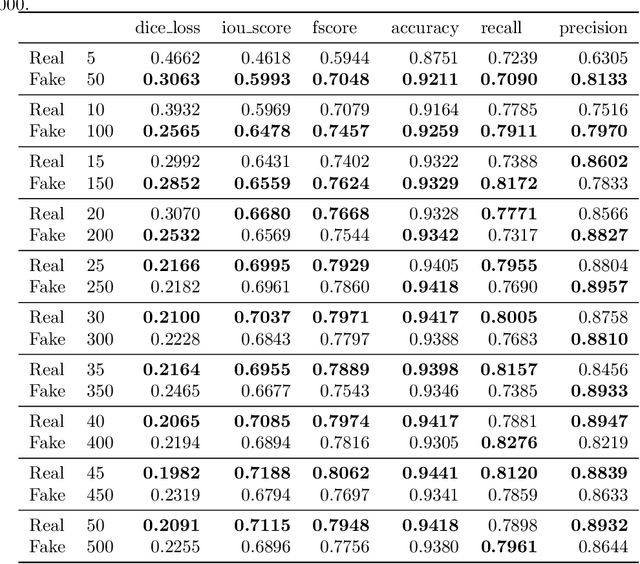
Abstract:Processing medical data to find abnormalities is a time-consuming and costly task, requiring tremendous efforts from medical experts. Therefore, Ai has become a popular tool for the automatic processing of medical data, acting as a supportive tool for doctors. AI tools highly depend on data for training the models. However, there are several constraints to access to large amounts of medical data to train machine learning algorithms in the medical domain, e.g., due to privacy concerns and the costly, time-consuming medical data annotation process. To address this, in this paper we present a novel synthetic data generation pipeline called SinGAN-Seg to produce synthetic medical data with the corresponding annotated ground truth masks. We show that these synthetic data generation pipelines can be used as an alternative to bypass privacy concerns and as an alternative way to produce artificial segmentation datasets with corresponding ground truth masks to avoid the tedious medical data annotation process. As a proof of concept, we used an open polyp segmentation dataset. By training UNet++ using both the real polyp segmentation dataset and the corresponding synthetic dataset generated from the SinGAN-Seg pipeline, we show that the synthetic data can achieve a very close performance to the real data when the real segmentation datasets are large enough. In addition, we show that synthetic data generated from the SinGAN-Seg pipeline improving the performance of segmentation algorithms when the training dataset is very small. Since our SinGAN-Seg pipeline is applicable for any medical dataset, this pipeline can be used with any other segmentation datasets.
PolypGen: A multi-center polyp detection and segmentation dataset for generalisability assessment
Jun 08, 2021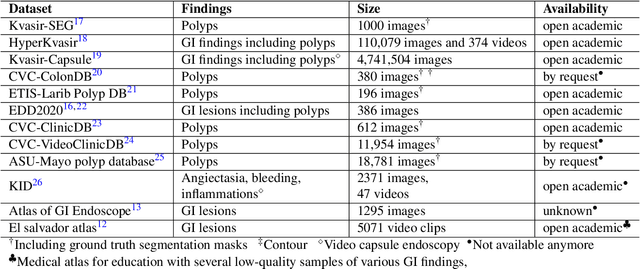
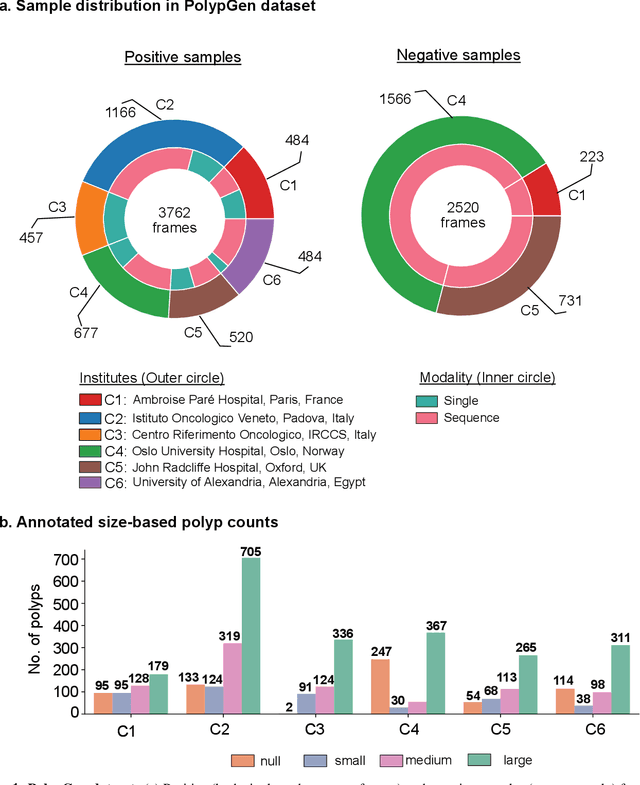
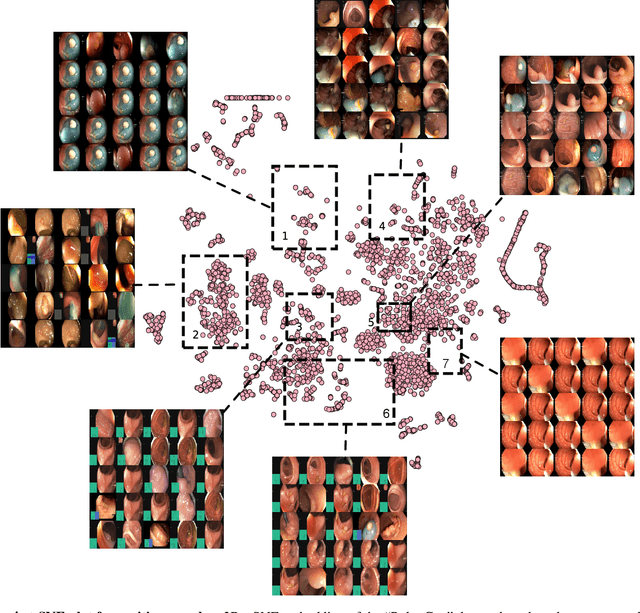
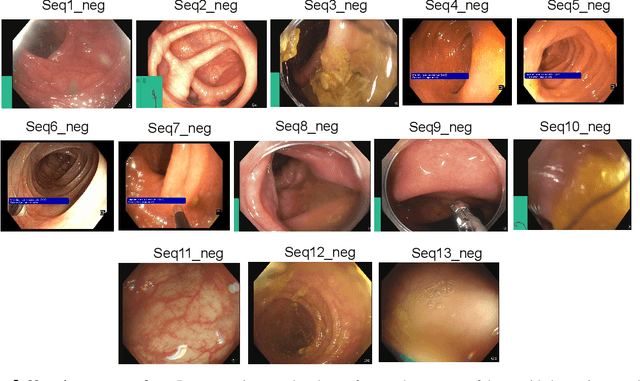
Abstract:Polyps in the colon are widely known as cancer precursors identified by colonoscopy either related to diagnostic work-up for symptoms, colorectal cancer screening or systematic surveillance of certain diseases. Whilst most polyps are benign, the number, size and the surface structure of the polyp are tightly linked to the risk of colon cancer. There exists a high missed detection rate and incomplete removal of colon polyps due to the variable nature, difficulties to delineate the abnormality, high recurrence rates and the anatomical topography of the colon. In the past, several methods have been built to automate polyp detection and segmentation. However, the key issue of most methods is that they have not been tested rigorously on a large multi-center purpose-built dataset. Thus, these methods may not generalise to different population datasets as they overfit to a specific population and endoscopic surveillance. To this extent, we have curated a dataset from 6 different centers incorporating more than 300 patients. The dataset includes both single frame and sequence data with 3446 annotated polyp labels with precise delineation of polyp boundaries verified by six senior gastroenterologists. To our knowledge, this is the most comprehensive detection and pixel-level segmentation dataset curated by a team of computational scientists and expert gastroenterologists. This dataset has been originated as the part of the Endocv2021 challenge aimed at addressing generalisability in polyp detection and segmentation. In this paper, we provide comprehensive insight into data construction and annotation strategies, annotation quality assurance and technical validation for our extended EndoCV2021 dataset which we refer to as PolypGen.
Few-shot segmentation of medical images based on meta-learning with implicit gradients
Jun 06, 2021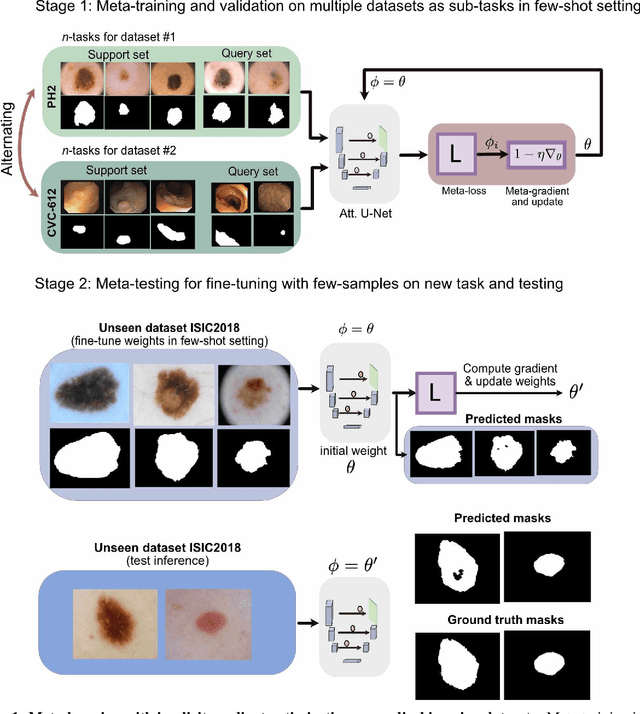
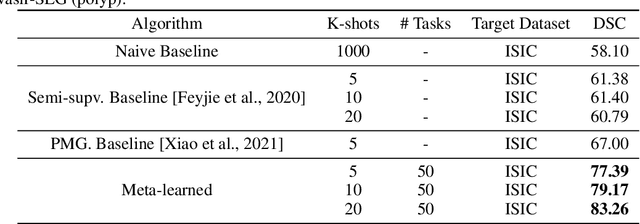
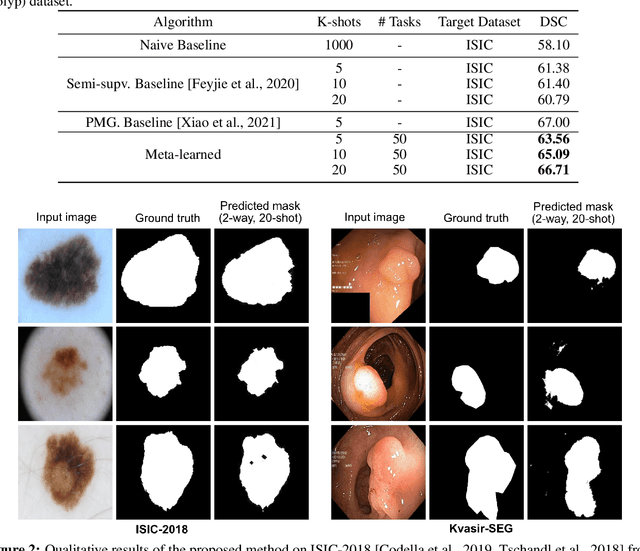
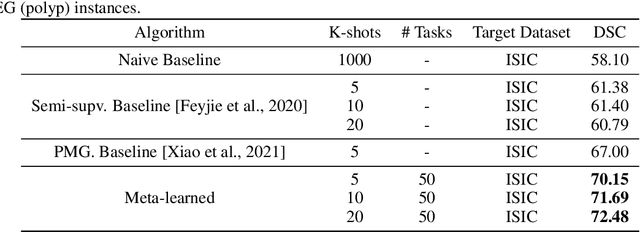
Abstract:Classical supervised methods commonly used often suffer from the requirement of an abudant number of training samples and are unable to generalize on unseen datasets. As a result, the broader application of any trained model is very limited in clinical settings. However, few-shot approaches can minimize the need for enormous reliable ground truth labels that are both labor intensive and expensive. To this end, we propose to exploit an optimization-based implicit model agnostic meta-learning {iMAML} algorithm in a few-shot setting for medical image segmentation. Our approach can leverage the learned weights from a diverse set of training samples and can be deployed on a new unseen dataset. We show that unlike classical few-shot learning approaches, our method has improved generalization capability. To our knowledge, this is the first work that exploits iMAML for medical image segmentation. Our quantitative results on publicly available skin and polyp datasets show that the proposed method outperforms the naive supervised baseline model and two recent few-shot segmentation approaches by large margins.
MSRF-Net: A Multi-Scale Residual Fusion Network for Biomedical Image Segmentation
May 16, 2021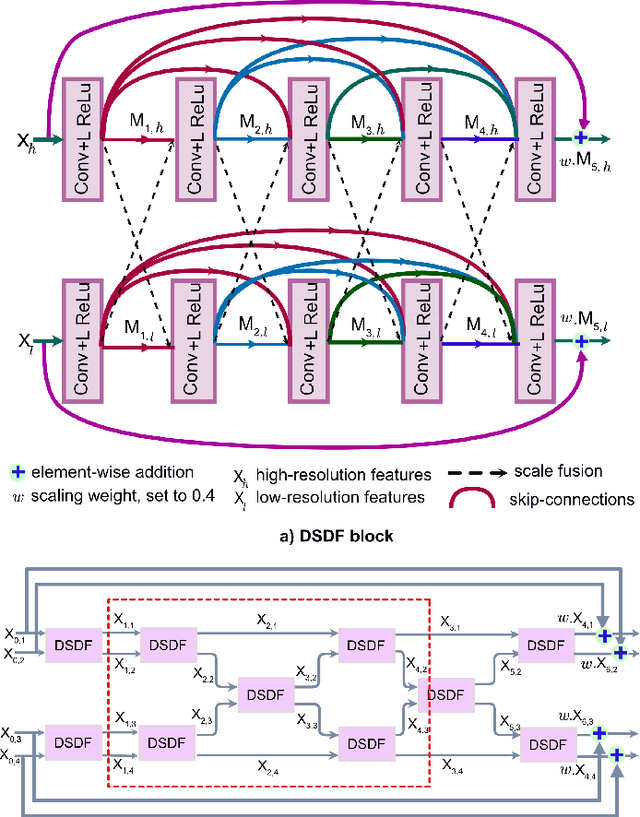
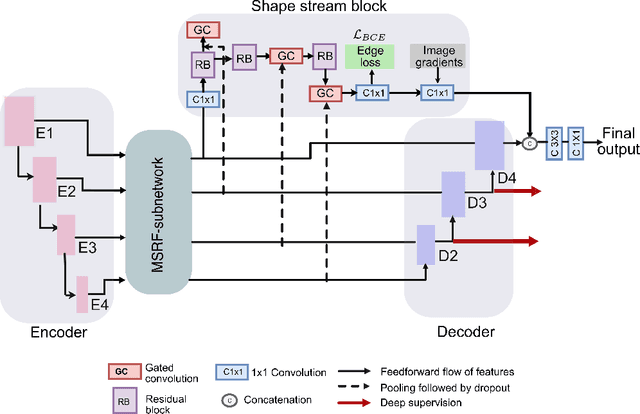
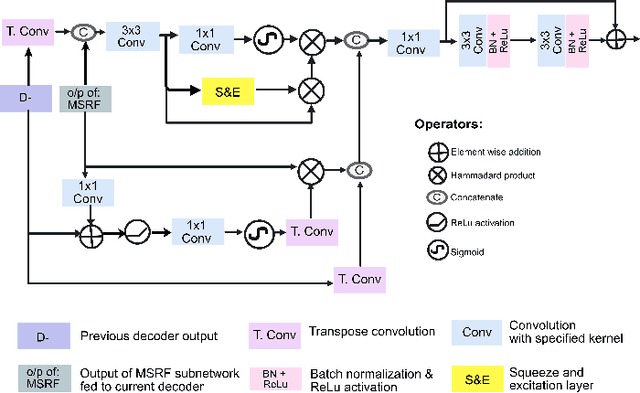
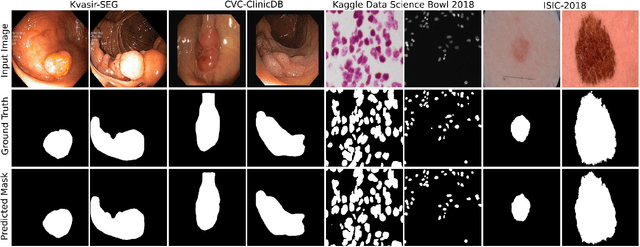
Abstract:Methods based on convolutional neural networks have improved the performance of biomedical image segmentation. However, most of these methods cannot efficiently segment objects of variable sizes and train on small and biased datasets, which are common in biomedical use cases. While methods exist that incorporate multi-scale fusion approaches to address the challenges arising with variable sizes, they usually use complex models that are more suitable for general semantic segmentation computer vision problems. In this paper, we propose a novel architecture called MSRF-Net, which is specially designed for medical image segmentation tasks. The proposed MSRF-Net is able to exchange multi-scale features of varying receptive fields using a dual-scale dense fusion block (DSDF). Our DSDF block can exchange information rigorously across two different resolution scales, and our MSRF sub-network uses multiple DSDF blocks in sequence to perform multi-scale fusion. This allows the preservation of resolution, improved information flow, and propagation of both high- and low-level features to obtain accurate segmentation maps. The proposed MSRF-Net allows to capture object variabilities and provides improved results on different biomedical datasets. Extensive experiments on MSRF-Net demonstrate that the proposed method outperforms most of the cutting-edge medical image segmentation state-of-the-art methods. MSRF-Net advances the performance on four publicly available datasets, and also, MSRF-Net is more generalizable as compared to state-of-the-art methods.
NanoNet: Real-Time Polyp Segmentation in Video Capsule Endoscopy and Colonoscopy
Apr 22, 2021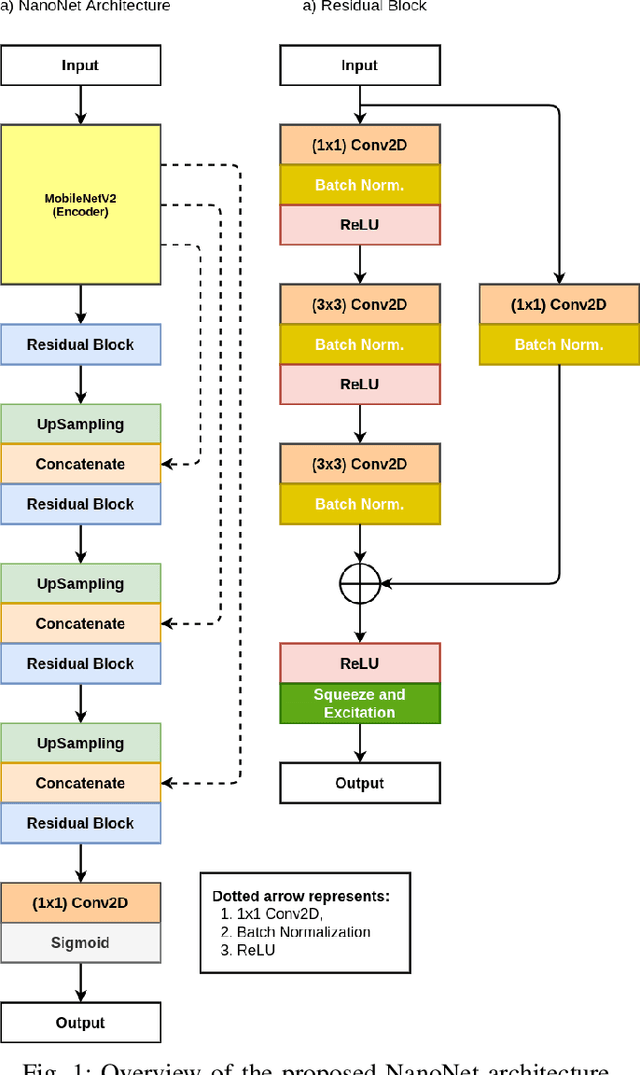
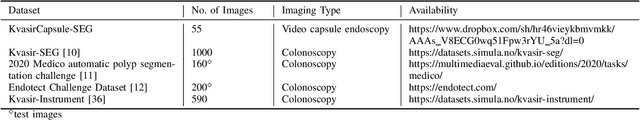
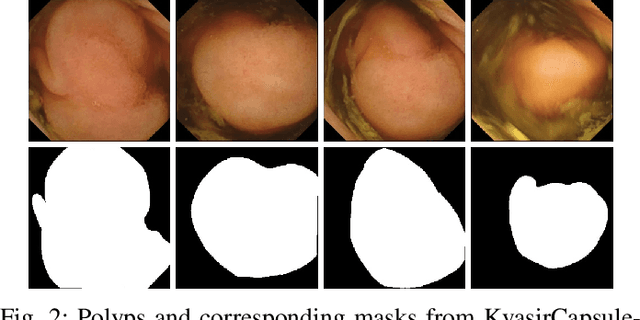
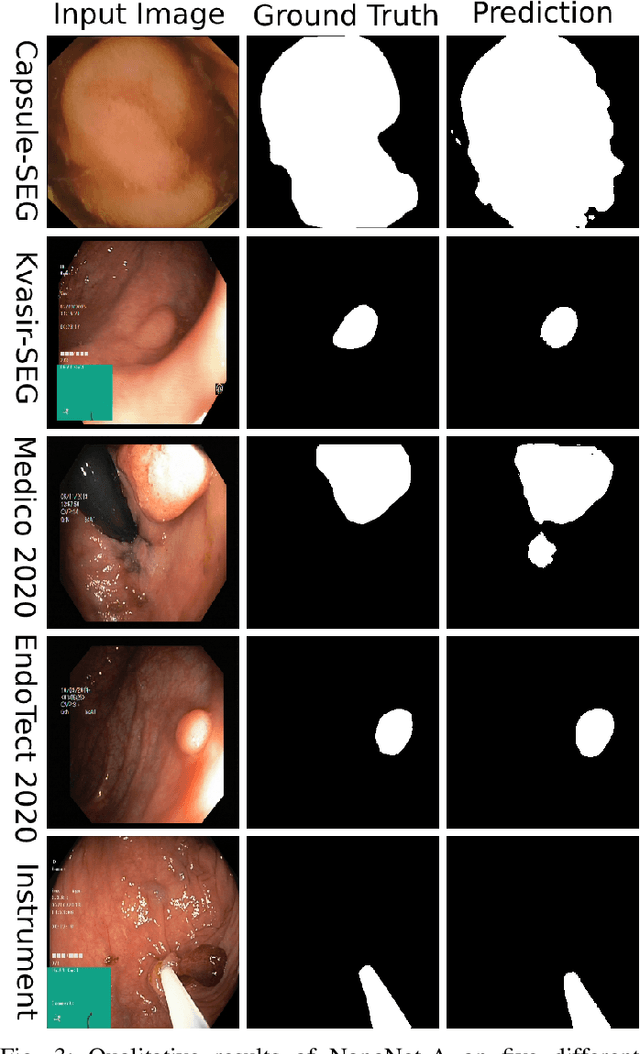

Abstract:Deep learning in gastrointestinal endoscopy can assist to improve clinical performance and be helpful to assess lesions more accurately. To this extent, semantic segmentation methods that can perform automated real-time delineation of a region-of-interest, e.g., boundary identification of cancer or precancerous lesions, can benefit both diagnosis and interventions. However, accurate and real-time segmentation of endoscopic images is extremely challenging due to its high operator dependence and high-definition image quality. To utilize automated methods in clinical settings, it is crucial to design lightweight models with low latency such that they can be integrated with low-end endoscope hardware devices. In this work, we propose NanoNet, a novel architecture for the segmentation of video capsule endoscopy and colonoscopy images. Our proposed architecture allows real-time performance and has higher segmentation accuracy compared to other more complex ones. We use video capsule endoscopy and standard colonoscopy datasets with polyps, and a dataset consisting of endoscopy biopsies and surgical instruments, to evaluate the effectiveness of our approach. Our experiments demonstrate the increased performance of our architecture in terms of a trade-off between model complexity, speed, model parameters, and metric performances. Moreover, the resulting model size is relatively tiny, with only nearly 36,000 parameters compared to traditional deep learning approaches having millions of parameters.
FANet: A Feedback Attention Network for Improved Biomedical Image Segmentation
Mar 31, 2021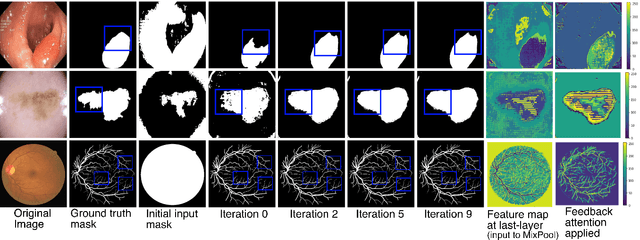
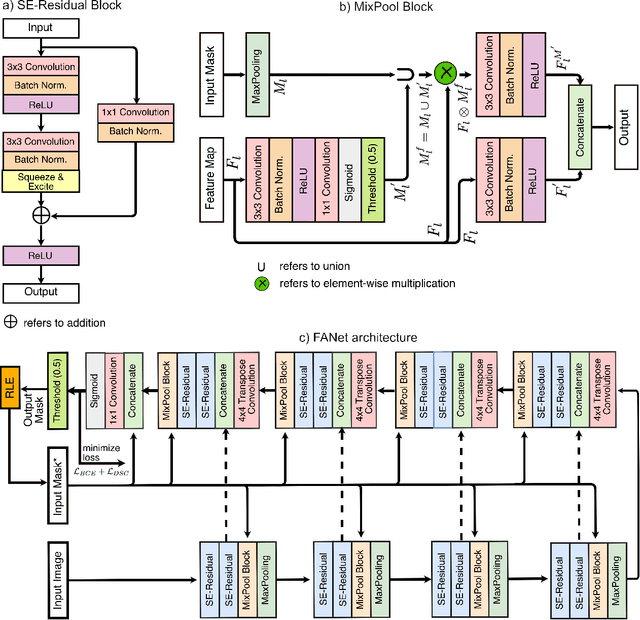
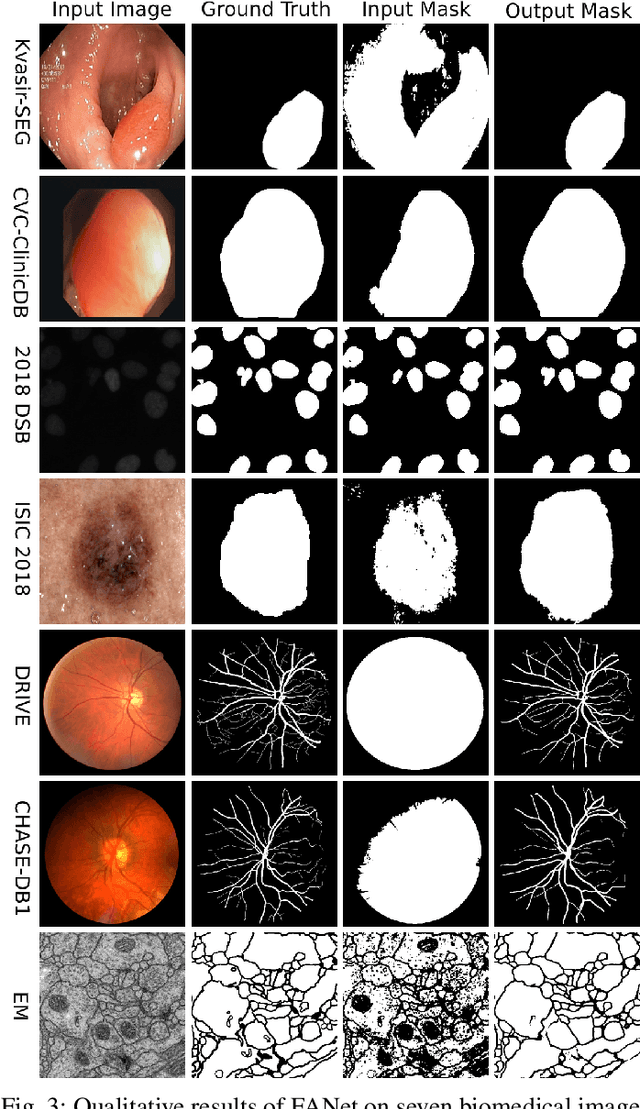
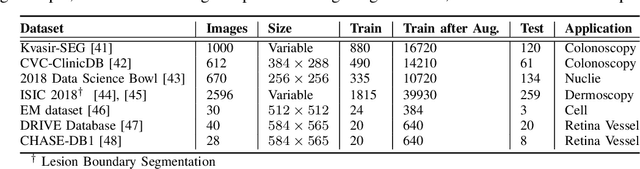
Abstract:With the increase in available large clinical and experimental datasets, there has been substantial amount of work being done on addressing the challenges in the area of biomedical image analysis. Image segmentation, which is crucial for any quantitative analysis, has especially attracted attention. Recent hardware advancement has led to the success of deep learning approaches. However, although deep learning models are being trained on large datasets, existing methods do not use the information from different learning epochs effectively. In this work, we leverage the information of each training epoch to prune the prediction maps of the subsequent epochs. We propose a novel architecture called feedback attention network (FANet) that unifies the previous epoch mask with the feature map of the current training epoch. The previous epoch mask is then used to provide a hard attention to the learnt feature maps at different convolutional layers. The network also allows to rectify the predictions in an iterative fashion during the test time. We show that our proposed feedback attention model provides a substantial improvement on most segmentation metrics tested on seven publicly available biomedical imaging datasets demonstrating the effectiveness of the proposed FANet.
LightLayers: Parameter Efficient Dense and Convolutional Layers for Image Classification
Jan 06, 2021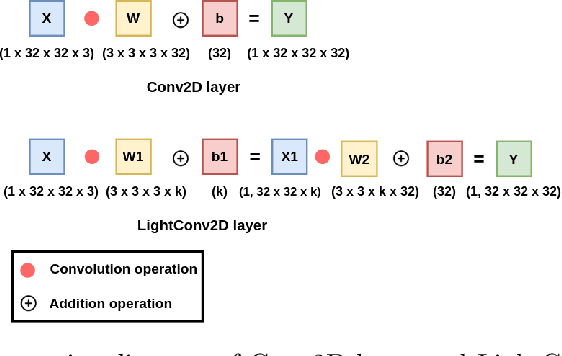
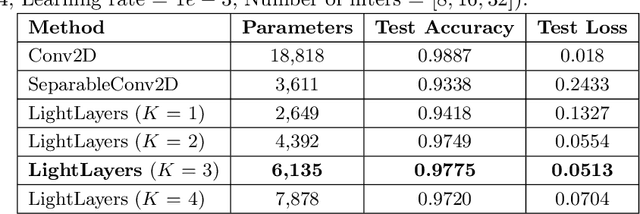
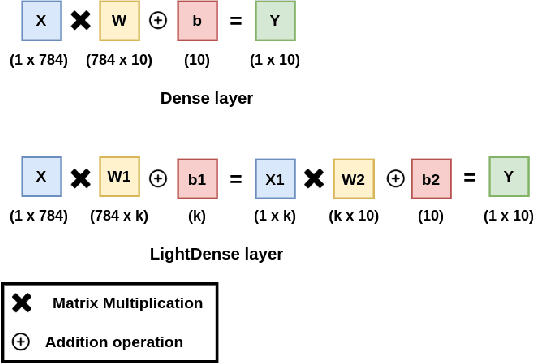
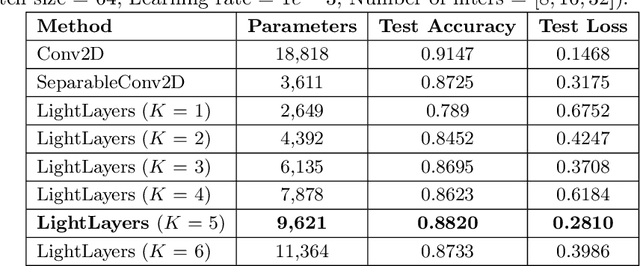
Abstract:Deep Neural Networks (DNNs) have become the de-facto standard in computer vision, as well as in many other pattern recognition tasks. A key drawback of DNNs is that the training phase can be very computationally expensive. Organizations or individuals that cannot afford purchasing state-of-the-art hardware or tapping into cloud-hosted infrastructures may face a long waiting time before the training completes or might not be able to train a model at all. Investigating novel ways to reduce the training time could be a potential solution to alleviate this drawback, and thus enabling more rapid development of new algorithms and models. In this paper, we propose LightLayers, a method for reducing the number of trainable parameters in deep neural networks (DNN). The proposed LightLayers consists of LightDense andLightConv2D layer that are as efficient as regular Conv2D and Dense layers, but uses less parameters. We resort to Matrix Factorization to reduce the complexity of the DNN models resulting into lightweight DNNmodels that require less computational power, without much loss in the accuracy. We have tested LightLayers on MNIST, Fashion MNIST, CI-FAR 10, and CIFAR 100 datasets. Promising results are obtained for MNIST, Fashion MNIST, CIFAR-10 datasets whereas CIFAR 100 shows acceptable performance by using fewer parameters.
DDANet: Dual Decoder Attention Network for Automatic Polyp Segmentation
Dec 30, 2020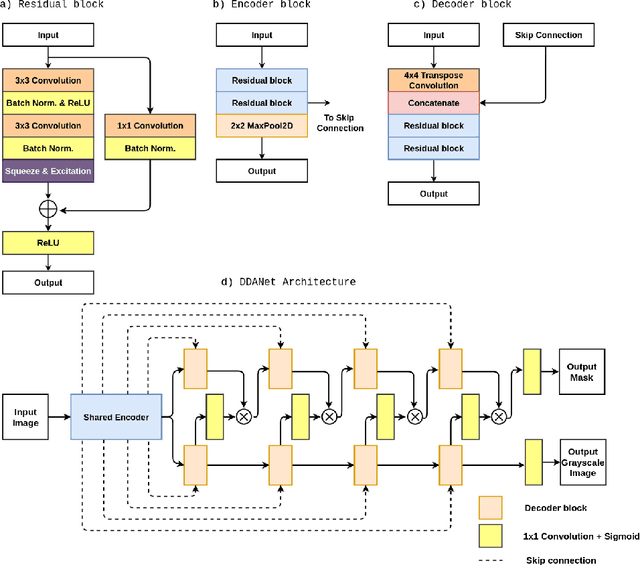
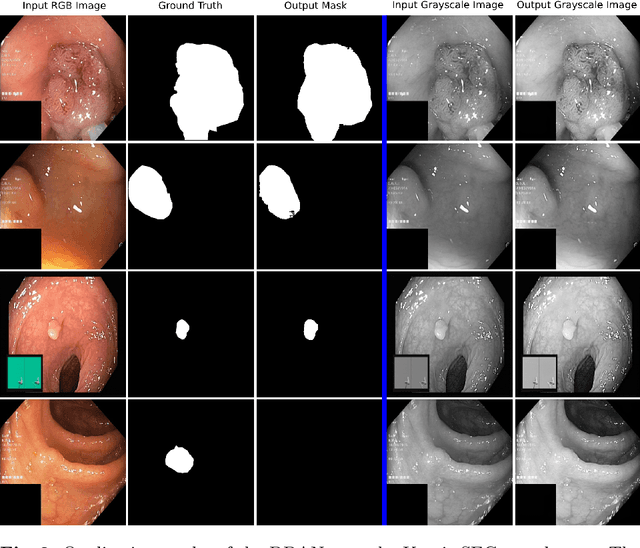
Abstract:Colonoscopy is the gold standard for examination and detection of colorectal polyps. Localization and delineation of polyps can play a vital role in treatment (e.g., surgical planning) and prognostic decision making. Polyp segmentation can provide detailed boundary information for clinical analysis. Convolutional neural networks have improved the performance in colonoscopy. However, polyps usually possess various challenges, such as intra-and inter-class variation and noise. While manual labeling for polyp assessment requires time from experts and is prone to human error (e.g., missed lesions), an automated, accurate, and fast segmentation can improve the quality of delineated lesion boundaries and reduce missed rate. The Endotect challenge provides an opportunity to benchmark computer vision methods by training on the publicly available Hyperkvasir and testing on a separate unseen dataset. In this paper, we propose a novel architecture called ``DDANet'' based on a dual decoder attention network. Our experiments demonstrate that the model trained on the Kvasir-SEG dataset and tested on an unseen dataset achieves a dice coefficient of 0.7874, mIoU of 0.7010, recall of 0.7987, and a precision of 0.8577, demonstrating the generalization ability of our model.
Medico Multimedia Task at MediaEval 2020: Automatic Polyp Segmentation
Dec 30, 2020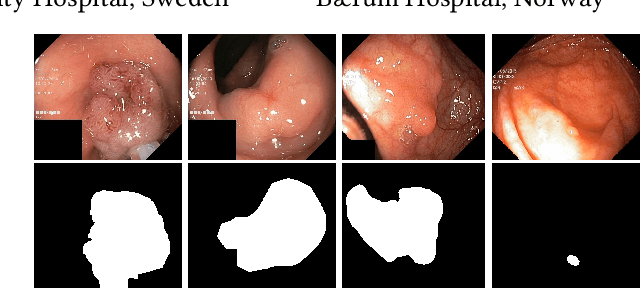

Abstract:Colorectal cancer is the third most common cause of cancer worldwide. According to Global cancer statistics 2018, the incidence of colorectal cancer is increasing in both developing and developed countries. Early detection of colon anomalies such as polyps is important for cancer prevention, and automatic polyp segmentation can play a crucial role for this. Regardless of the recent advancement in early detection and treatment options, the estimated polyp miss rate is still around 20\%. Support via an automated computer-aided diagnosis system could be one of the potential solutions for the overlooked polyps. Such detection systems can help low-cost design solutions and save doctors time, which they could for example use to perform more patient examinations. In this paper, we introduce the 2020 Medico challenge, provide some information on related work and the dataset, describe the task and evaluation metrics, and discuss the necessity of organizing the Medico challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge