Hugo Lewi Hammer
Knowledge-Guided Retrieval-Augmented Generation for Zero-Shot Psychiatric Data: Privacy Preserving Synthetic Data Generation
Mar 26, 2026Abstract:AI systems in healthcare research have shown potential to increase patient throughput and assist clinicians, yet progress is constrained by limited access to real patient data. To address this issue, we present a zero-shot, knowledge-guided framework for psychiatric tabular data in which large language models (LLMs) are steered via Retrieval-Augmented Generation using the Diagnostic and Statistical Manual of Mental Disorders (DSM-5) and the International Classification of Diseases (ICD-10). We conducted experiments using different combinations of knowledge bases to generate privacy-preserving synthetic data. The resulting models were benchmarked against two state-of-the-art deep learning models for synthetic tabular data generation, namely CTGAN and TVAE, both of which rely on real data and therefore entail potential privacy risks. Evaluation was performed on six anxiety-related disorders: specific phobia, social anxiety disorder, agoraphobia, generalized anxiety disorder, separation anxiety disorder, and panic disorder. CTGAN typically achieves the best marginals and multivariate structure, while the knowledge-augmented LLM is competitive on pairwise structure and attains the lowest pairwise error in separation anxiety and social anxiety. An ablation study shows that clinical retrieval reliably improves univariate and pairwise fidelity over a no-retrieval LLM. Privacy analyses indicate that the real data-free LLM yields modest overlaps and a low average linkage risk comparable to CTGAN, whereas TVAE exhibits extensive duplication despite a low k-map score. Overall, grounding an LLM in clinical knowledge enables high-quality, privacy-preserving synthetic psychiatric data when real datasets are unavailable or cannot be shared.
Classifying Dry Eye Disease Patients from Healthy Controls Using Machine Learning and Metabolomics Data
Jun 20, 2024



Abstract:Dry eye disease is a common disorder of the ocular surface, leading patients to seek eye care. Clinical signs and symptoms are currently used to diagnose dry eye disease. Metabolomics, a method for analyzing biological systems, has been found helpful in identifying distinct metabolites in patients and in detecting metabolic profiles that may indicate dry eye disease at early stages. In this study, we explored using machine learning and metabolomics information to identify which cataract patients suffered from dry eye disease. As there is no one-size-fits-all machine learning model for metabolomics data, choosing the most suitable model can significantly affect the quality of predictions and subsequent metabolomics analyses. To address this challenge, we conducted a comparative analysis of nine machine learning models on three metabolomics data sets from cataract patients with and without dry eye disease. The models were evaluated and optimized using nested k-fold cross-validation. To assess the performance of these models, we selected a set of suitable evaluation metrics tailored to the data set's challenges. The logistic regression model overall performed the best, achieving the highest area under the curve score of 0.8378, balanced accuracy of 0.735, Matthew's correlation coefficient of 0.5147, an F1-score of 0.8513, and a specificity of 0.5667. Additionally, following the logistic regression, the XGBoost and Random Forest models also demonstrated good performance.
An Extensive Study on Cross-Dataset Bias and Evaluation Metrics Interpretation for Machine Learning applied to Gastrointestinal Tract Abnormality Classification
May 08, 2020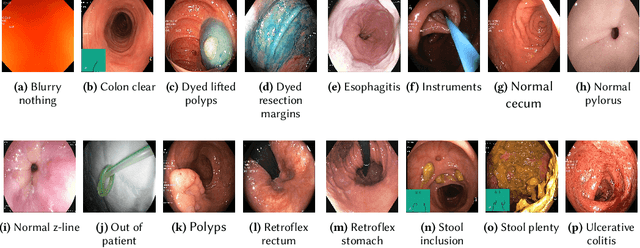
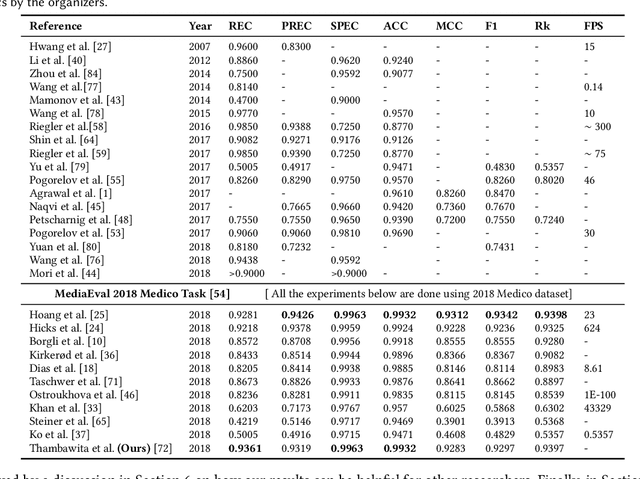
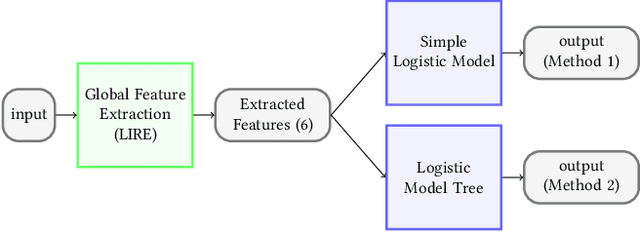
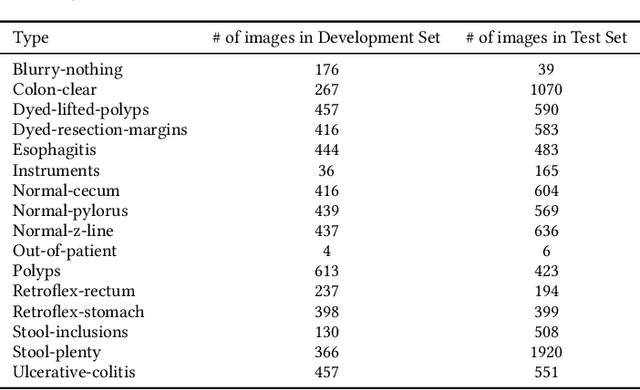
Abstract:Precise and efficient automated identification of Gastrointestinal (GI) tract diseases can help doctors treat more patients and improve the rate of disease detection and identification. Currently, automatic analysis of diseases in the GI tract is a hot topic in both computer science and medical-related journals. Nevertheless, the evaluation of such an automatic analysis is often incomplete or simply wrong. Algorithms are often only tested on small and biased datasets, and cross-dataset evaluations are rarely performed. A clear understanding of evaluation metrics and machine learning models with cross datasets is crucial to bring research in the field to a new quality level. Towards this goal, we present comprehensive evaluations of five distinct machine learning models using Global Features and Deep Neural Networks that can classify 16 different key types of GI tract conditions, including pathological findings, anatomical landmarks, polyp removal conditions, and normal findings from images captured by common GI tract examination instruments. In our evaluation, we introduce performance hexagons using six performance metrics such as recall, precision, specificity, accuracy, F1-score, and Matthews Correlation Coefficient to demonstrate how to determine the real capabilities of models rather than evaluating them shallowly. Furthermore, we perform cross-dataset evaluations using different datasets for training and testing. With these cross-dataset evaluations, we demonstrate the challenge of actually building a generalizable model that could be used across different hospitals. Our experiments clearly show that more sophisticated performance metrics and evaluation methods need to be applied to get reliable models rather than depending on evaluations of the splits of the same dataset, i.e., the performance metrics should always be interpreted together rather than relying on a single metric.
The Medico-Task 2018: Disease Detection in the Gastrointestinal Tract using Global Features and Deep Learning
Oct 31, 2018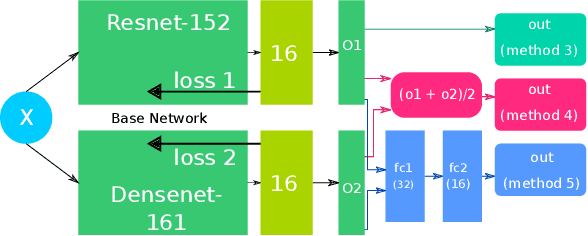
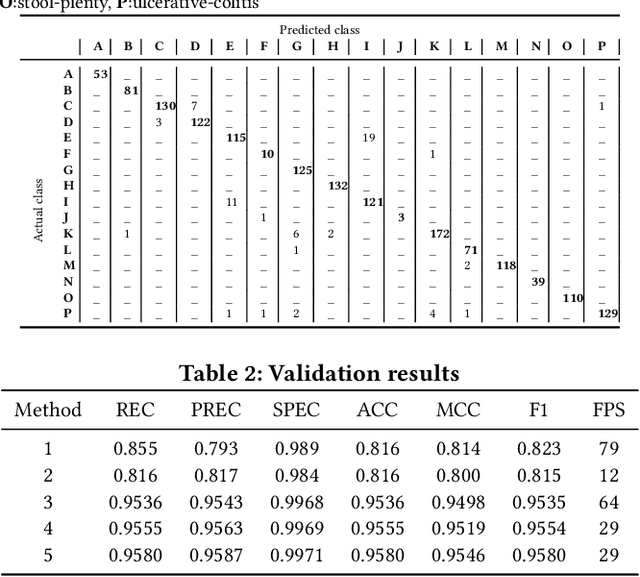
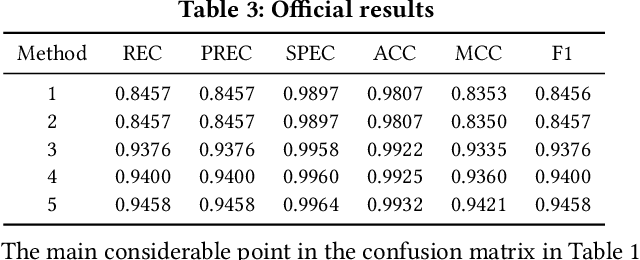
Abstract:In this paper, we present our approach for the 2018 Medico Task classifying diseases in the gastrointestinal tract. We have proposed a system based on global features and deep neural networks. The best approach combines two neural networks, and the reproducible experimental results signify the efficiency of the proposed model with an accuracy rate of 95.80%, a precision of 95.87%, and an F1-score of 95.80%.
* 2 pages + 1 page for references, 1 figure, Conference paper
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge