Megan Leszczynski
Beyond Single Items: Exploring User Preferences in Item Sets with the Conversational Playlist Curation Dataset
Mar 13, 2023Abstract:Users in consumption domains, like music, are often able to more efficiently provide preferences over a set of items (e.g. a playlist or radio) than over single items (e.g. songs). Unfortunately, this is an underexplored area of research, with most existing recommendation systems limited to understanding preferences over single items. Curating an item set exponentiates the search space that recommender systems must consider (all subsets of items!): this motivates conversational approaches-where users explicitly state or refine their preferences and systems elicit preferences in natural language-as an efficient way to understand user needs. We call this task conversational item set curation and present a novel data collection methodology that efficiently collects realistic preferences about item sets in a conversational setting by observing both item-level and set-level feedback. We apply this methodology to music recommendation to build the Conversational Playlist Curation Dataset (CPCD), where we show that it leads raters to express preferences that would not be otherwise expressed. Finally, we propose a wide range of conversational retrieval models as baselines for this task and evaluate them on the dataset.
Generating Synthetic Data for Conversational Music Recommendation Using Random Walks and Language Models
Jan 27, 2023



Abstract:Conversational recommendation systems (CRSs) enable users to use natural language feedback to control their recommendations, overcoming many of the challenges of traditional recommendation systems. However, the practical adoption of CRSs remains limited due to a lack of rich and diverse conversational training data that pairs user utterances with recommendations. To address this problem, we introduce a new method to generate synthetic training data by transforming curated item collections, such as playlists or movie watch lists, into item-seeking conversations. First, we use a biased random walk to generate a sequence of slates, or sets of item recommendations; then, we use a language model to generate corresponding user utterances. We demonstrate our approach by generating a conversational music recommendation dataset with over one million conversations, which were found to be consistent with relevant recommendations by a crowdsourced evaluation. Using the synthetic data to train a CRS, we significantly outperform standard retrieval baselines in offline and online evaluations.
TABi: Type-Aware Bi-Encoders for Open-Domain Entity Retrieval
Apr 18, 2022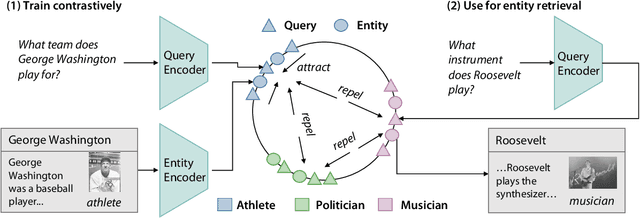
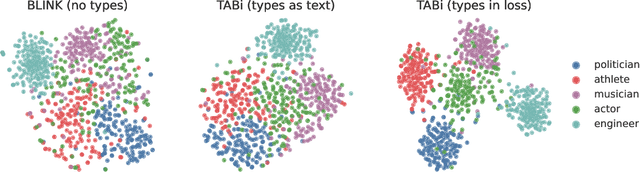
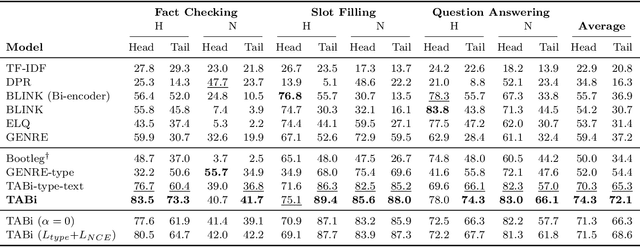

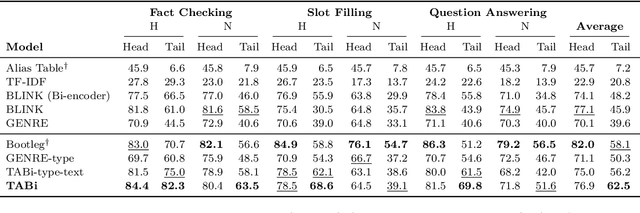
Abstract:Entity retrieval--retrieving information about entity mentions in a query--is a key step in open-domain tasks, such as question answering or fact checking. However, state-of-the-art entity retrievers struggle to retrieve rare entities for ambiguous mentions due to biases towards popular entities. Incorporating knowledge graph types during training could help overcome popularity biases, but there are several challenges: (1) existing type-based retrieval methods require mention boundaries as input, but open-domain tasks run on unstructured text, (2) type-based methods should not compromise overall performance, and (3) type-based methods should be robust to noisy and missing types. In this work, we introduce TABi, a method to jointly train bi-encoders on knowledge graph types and unstructured text for entity retrieval for open-domain tasks. TABi leverages a type-enforced contrastive loss to encourage entities and queries of similar types to be close in the embedding space. TABi improves retrieval of rare entities on the Ambiguous Entity Retrieval (AmbER) sets, while maintaining strong overall retrieval performance on open-domain tasks in the KILT benchmark compared to state-of-the-art retrievers. TABi is also robust to incomplete type systems, improving rare entity retrieval over baselines with only 5% type coverage of the training dataset. We make our code publicly available at https://github.com/HazyResearch/tabi.
Cross-Domain Data Integration for Named Entity Disambiguation in Biomedical Text
Oct 15, 2021Abstract:Named entity disambiguation (NED), which involves mapping textual mentions to structured entities, is particularly challenging in the medical domain due to the presence of rare entities. Existing approaches are limited by the presence of coarse-grained structural resources in biomedical knowledge bases as well as the use of training datasets that provide low coverage over uncommon resources. In this work, we address these issues by proposing a cross-domain data integration method that transfers structural knowledge from a general text knowledge base to the medical domain. We utilize our integration scheme to augment structural resources and generate a large biomedical NED dataset for pretraining. Our pretrained model with injected structural knowledge achieves state-of-the-art performance on two benchmark medical NED datasets: MedMentions and BC5CDR. Furthermore, we improve disambiguation of rare entities by up to 57 accuracy points.
Managing ML Pipelines: Feature Stores and the Coming Wave of Embedding Ecosystems
Aug 11, 2021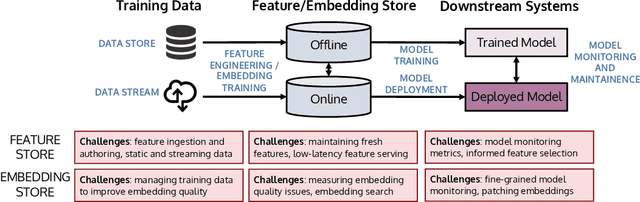
Abstract:The industrial machine learning pipeline requires iterating on model features, training and deploying models, and monitoring deployed models at scale. Feature stores were developed to manage and standardize the engineer's workflow in this end-to-end pipeline, focusing on traditional tabular feature data. In recent years, however, model development has shifted towards using self-supervised pretrained embeddings as model features. Managing these embeddings and the downstream systems that use them introduces new challenges with respect to managing embedding training data, measuring embedding quality, and monitoring downstream models that use embeddings. These challenges are largely unaddressed in standard feature stores. Our goal in this tutorial is to introduce the feature store system and discuss the challenges and current solutions to managing these new embedding-centric pipelines.
Kaleidoscope: An Efficient, Learnable Representation For All Structured Linear Maps
Jan 05, 2021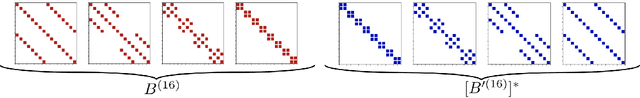
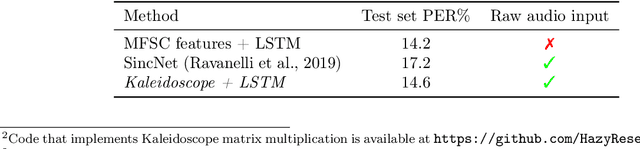
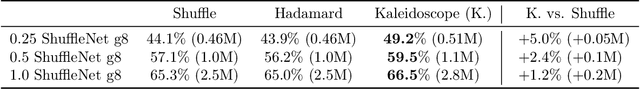

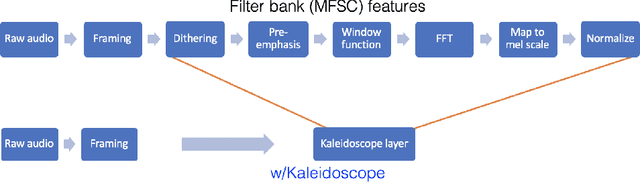

Abstract:Modern neural network architectures use structured linear transformations, such as low-rank matrices, sparse matrices, permutations, and the Fourier transform, to improve inference speed and reduce memory usage compared to general linear maps. However, choosing which of the myriad structured transformations to use (and its associated parameterization) is a laborious task that requires trading off speed, space, and accuracy. We consider a different approach: we introduce a family of matrices called kaleidoscope matrices (K-matrices) that provably capture any structured matrix with near-optimal space (parameter) and time (arithmetic operation) complexity. We empirically validate that K-matrices can be automatically learned within end-to-end pipelines to replace hand-crafted procedures, in order to improve model quality. For example, replacing channel shuffles in ShuffleNet improves classification accuracy on ImageNet by up to 5%. K-matrices can also simplify hand-engineered pipelines -- we replace filter bank feature computation in speech data preprocessing with a learnable kaleidoscope layer, resulting in only 0.4% loss in accuracy on the TIMIT speech recognition task. In addition, K-matrices can capture latent structure in models: for a challenging permuted image classification task, a K-matrix based representation of permutations is able to learn the right latent structure and improves accuracy of a downstream convolutional model by over 9%. We provide a practically efficient implementation of our approach, and use K-matrices in a Transformer network to attain 36% faster end-to-end inference speed on a language translation task.
Bootleg: Chasing the Tail with Self-Supervised Named Entity Disambiguation
Oct 23, 2020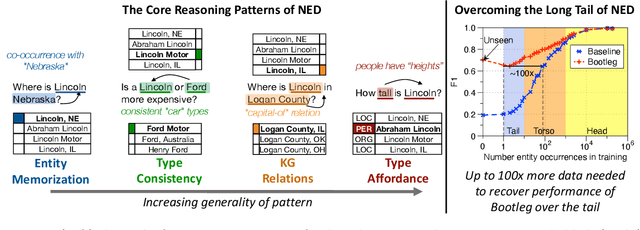
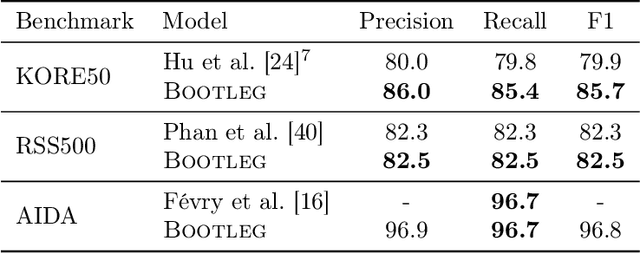
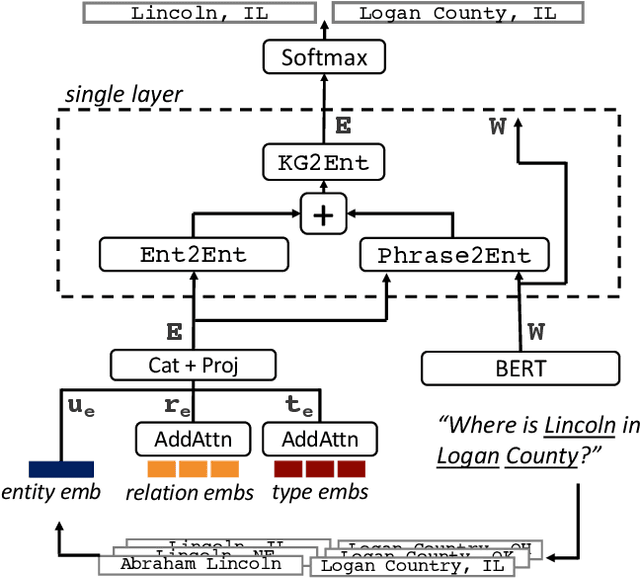
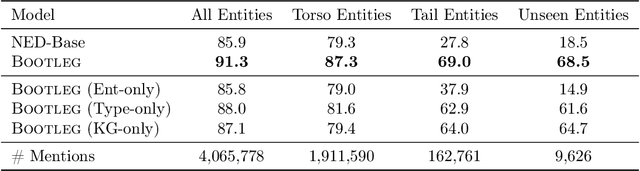
Abstract:A challenge for named entity disambiguation (NED), the task of mapping textual mentions to entities in a knowledge base, is how to disambiguate entities that appear rarely in the training data, termed tail entities. Humans use subtle reasoning patterns based on knowledge of entity facts, relations, and types to disambiguate unfamiliar entities. Inspired by these patterns, we introduce Bootleg, a self-supervised NED system that is explicitly grounded in reasoning patterns for disambiguation. We define core reasoning patterns for disambiguation, create a learning procedure to encourage the self-supervised model to learn the patterns, and show how to use weak supervision to enhance the signals in the training data. Encoding the reasoning patterns in a simple Transformer architecture, Bootleg meets or exceeds state-of-the-art on three NED benchmarks. We further show that the learned representations from Bootleg successfully transfer to other non-disambiguation tasks that require entity-based knowledge: we set a new state-of-the-art in the popular TACRED relation extraction task by 1.0 F1 points and demonstrate up to 8% performance lift in highly optimized production search and assistant tasks at a major technology company
Understanding the Downstream Instability of Word Embeddings
Feb 29, 2020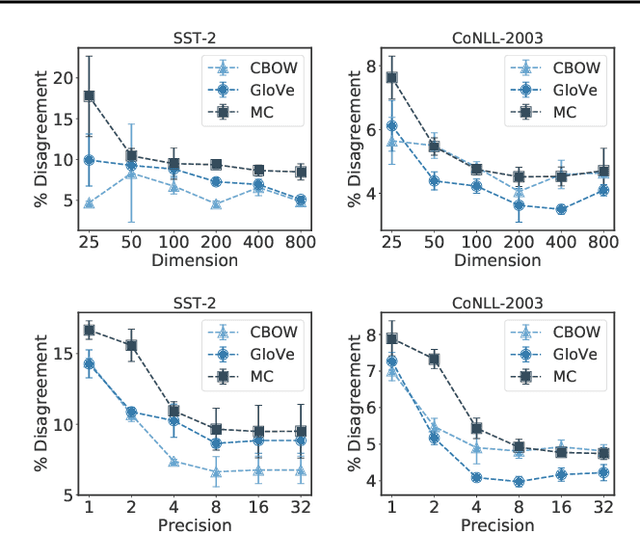
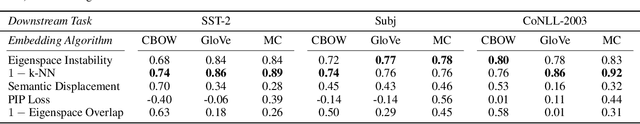
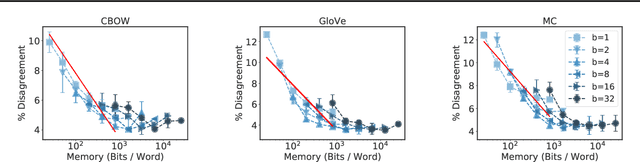
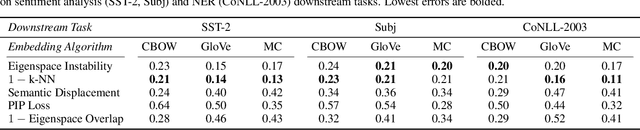
Abstract:Many industrial machine learning (ML) systems require frequent retraining to keep up-to-date with constantly changing data. This retraining exacerbates a large challenge facing ML systems today: model training is unstable, i.e., small changes in training data can cause significant changes in the model's predictions. In this paper, we work on developing a deeper understanding of this instability, with a focus on how a core building block of modern natural language processing (NLP) pipelines---pre-trained word embeddings---affects the instability of downstream NLP models. We first empirically reveal a tradeoff between stability and memory: increasing the embedding memory 2x can reduce the disagreement in predictions due to small changes in training data by 5% to 37% (relative). To theoretically explain this tradeoff, we introduce a new measure of embedding instability---the eigenspace instability measure---which we prove bounds the disagreement in downstream predictions introduced by the change in word embeddings. Practically, we show that the eigenspace instability measure can be a cost-effective way to choose embedding parameters to minimize instability without training downstream models, outperforming other embedding distance measures and performing competitively with a nearest neighbor-based measure. Finally, we demonstrate that the observed stability-memory tradeoffs extend to other types of embeddings as well, including knowledge graph and contextual word embeddings.
Low-Memory Neural Network Training: A Technical Report
Apr 24, 2019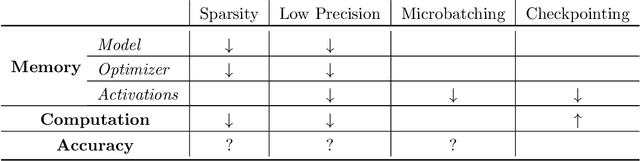
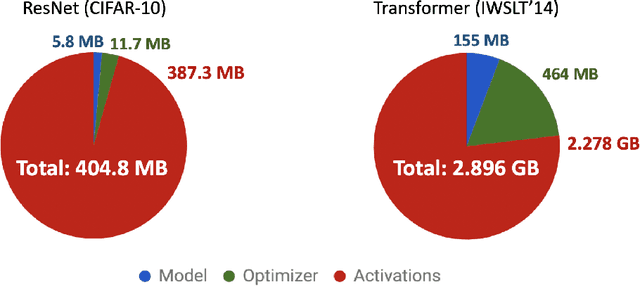
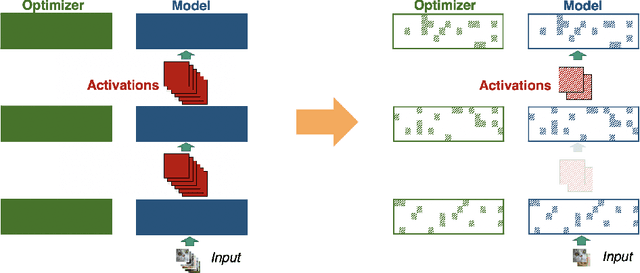

Abstract:Memory is increasingly often the bottleneck when training neural network models. Despite this, techniques to lower the overall memory requirements of training have been less widely studied compared to the extensive literature on reducing the memory requirements of inference. In this paper we study a fundamental question: How much memory is actually needed to train a neural network? To answer this question, we profile the overall memory usage of training on two representative deep learning benchmarks -- the WideResNet model for image classification and the DynamicConv Transformer model for machine translation -- and comprehensively evaluate four standard techniques for reducing the training memory requirements: (1) imposing sparsity on the model, (2) using low precision, (3) microbatching, and (4) gradient checkpointing. We explore how each of these techniques in isolation affects both the peak memory usage of training and the quality of the end model, and explore the memory, accuracy, and computation tradeoffs incurred when combining these techniques. Using appropriate combinations of these techniques, we show that it is possible to the reduce the memory required to train a WideResNet-28-2 on CIFAR-10 by up to 60.7x with a 0.4% loss in accuracy, and reduce the memory required to train a DynamicConv model on IWSLT'14 German to English translation by up to 8.7x with a BLEU score drop of 0.15.
High-Accuracy Low-Precision Training
Mar 09, 2018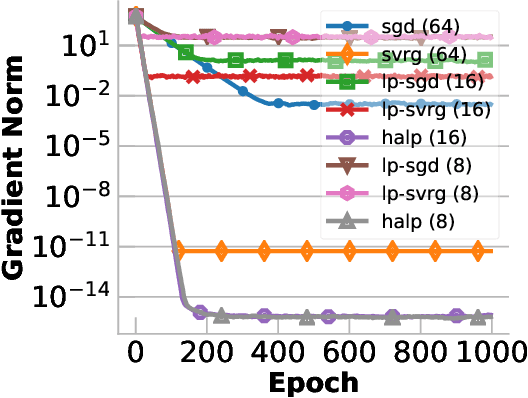
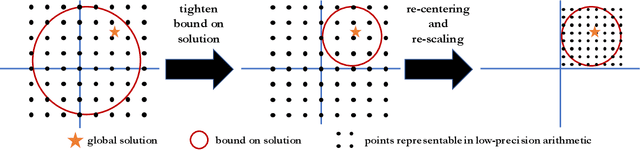
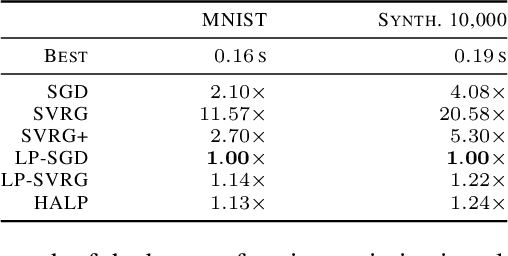
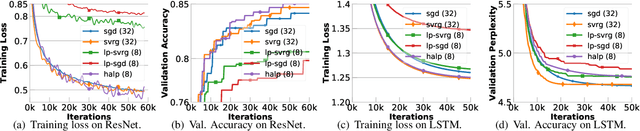
Abstract:Low-precision computation is often used to lower the time and energy cost of machine learning, and recently hardware accelerators have been developed to support it. Still, it has been used primarily for inference - not training. Previous low-precision training algorithms suffered from a fundamental tradeoff: as the number of bits of precision is lowered, quantization noise is added to the model, which limits statistical accuracy. To address this issue, we describe a simple low-precision stochastic gradient descent variant called HALP. HALP converges at the same theoretical rate as full-precision algorithms despite the noise introduced by using low precision throughout execution. The key idea is to use SVRG to reduce gradient variance, and to combine this with a novel technique called bit centering to reduce quantization error. We show that on the CPU, HALP can run up to $4 \times$ faster than full-precision SVRG and can match its convergence trajectory. We implemented HALP in TensorQuant, and show that it exceeds the validation performance of plain low-precision SGD on two deep learning tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge