Md. Kamrul Hasan
Systems and Software Lab, Department of Computer Science and Engineering, Islamic University of Technology
Assorted, Archetypal and Annotated Two Million (3A2M) Cooking Recipes Dataset based on Active Learning
Mar 27, 2023Abstract:Cooking recipes allow individuals to exchange culinary ideas and provide food preparation instructions. Due to a lack of adequate labeled data, categorizing raw recipes found online to the appropriate food genres is a challenging task in this domain. Utilizing the knowledge of domain experts to categorize recipes could be a solution. In this study, we present a novel dataset of two million culinary recipes labeled in respective categories leveraging the knowledge of food experts and an active learning technique. To construct the dataset, we collect the recipes from the RecipeNLG dataset. Then, we employ three human experts whose trustworthiness score is higher than 86.667% to categorize 300K recipe by their Named Entity Recognition (NER) and assign it to one of the nine categories: bakery, drinks, non-veg, vegetables, fast food, cereals, meals, sides and fusion. Finally, we categorize the remaining 1900K recipes using Active Learning method with a blend of Query-by-Committee and Human In The Loop (HITL) approaches. There are more than two million recipes in our dataset, each of which is categorized and has a confidence score linked with it. For the 9 genres, the Fleiss Kappa score of this massive dataset is roughly 0.56026. We believe that the research community can use this dataset to perform various machine learning tasks such as recipe genre classification, recipe generation of a specific genre, new recipe creation, etc. The dataset can also be used to train and evaluate the performance of various NLP tasks such as named entity recognition, part-of-speech tagging, semantic role labeling, and so on. The dataset will be available upon publication: https://tinyurl.com/3zu4778y.
Hybrid Parallel Imaging and Compressed Sensing MRI Reconstruction with GRAPPA Integrated Multi-loss Supervised GAN
Sep 19, 2022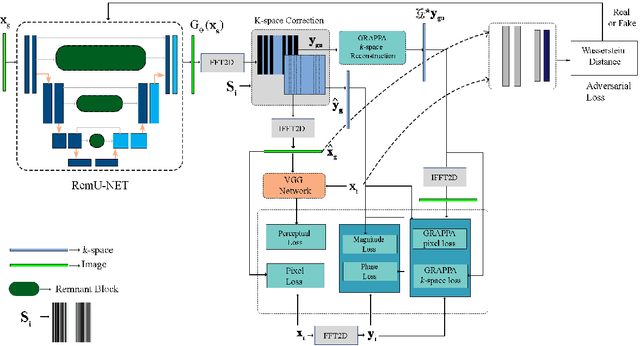
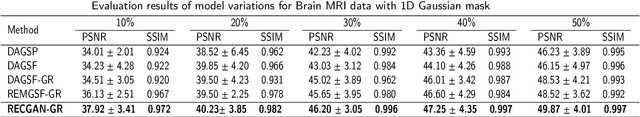

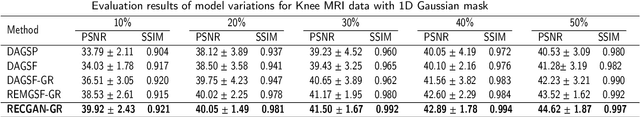
Abstract:Objective: Parallel imaging accelerates the acquisition of magnetic resonance imaging (MRI) data by acquiring additional sensitivity information with an array of receiver coils resulting in reduced phase encoding steps. Compressed sensing magnetic resonance imaging (CS-MRI) has achieved popularity in the field of medical imaging because of its less data requirement than parallel imaging. Parallel imaging and compressed sensing (CS) both speed up traditional MRI acquisition by minimizing the amount of data captured in the k-space. As acquisition time is inversely proportional to the number of samples, the inverse formation of an image from reduced k-space samples leads to faster acquisition but with aliasing artifacts. This paper proposes a novel Generative Adversarial Network (GAN) namely RECGAN-GR supervised with multi-modal losses for de-aliasing the reconstructed image. Methods: In contrast to existing GAN networks, our proposed method introduces a novel generator network namely RemU-Net integrated with dual-domain loss functions including weighted magnitude and phase loss functions along with parallel imaging-based loss i.e., GRAPPA consistency loss. A k-space correction block is proposed as refinement learning to make the GAN network self-resistant to generating unnecessary data which drives the convergence of the reconstruction process faster. Results: Comprehensive results show that the proposed RECGAN-GR achieves a 4 dB improvement in the PSNR among the GAN-based methods and a 2 dB improvement among conventional state-of-the-art CNN methods available in the literature. Conclusion and significance: The proposed work contributes to significant improvement in the image quality for low retained data leading to 5x or 10x faster acquisition.
Skin Lesion Analysis: A State-of-the-Art Survey, Systematic Review, and Future Trends
Aug 25, 2022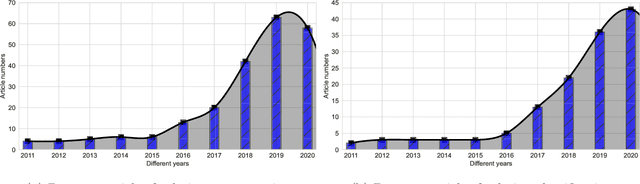
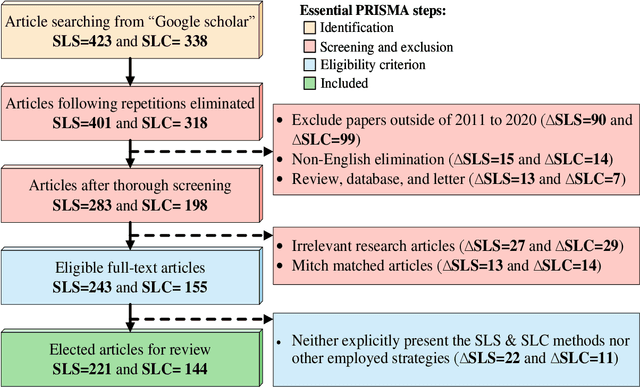
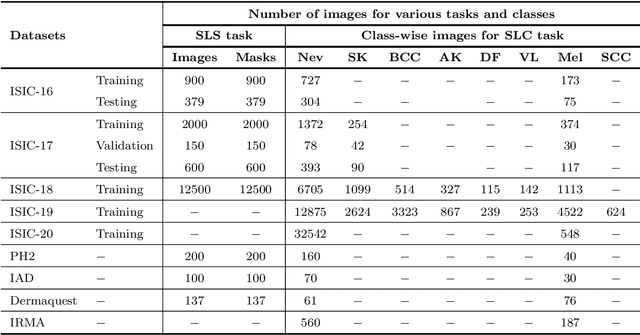
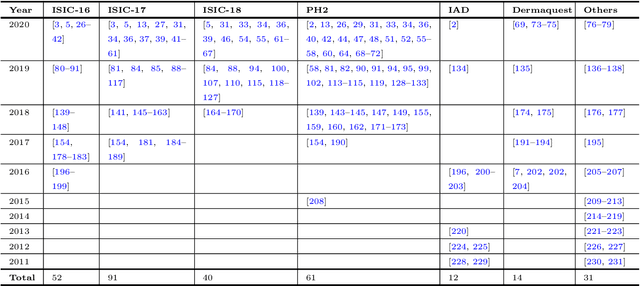
Abstract:The Computer-aided Diagnosis (CAD) system for skin lesion analysis is an emerging field of research that has the potential to relieve the burden and cost of skin cancer screening. Researchers have recently indicated increasing interest in developing such CAD systems, with the intention of providing a user-friendly tool to dermatologists in order to reduce the challenges that are raised by manual inspection. The purpose of this article is to provide a complete literature review of cutting-edge CAD techniques published between 2011 and 2020. The Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) method was used to identify a total of 365 publications, 221 for skin lesion segmentation and 144 for skin lesion classification. These articles are analyzed and summarized in a number of different ways so that we can contribute vital information about the methods for the evolution of CAD systems. These ways include: relevant and essential definitions and theories, input data (datasets utilization, preprocessing, augmentations, and fixing imbalance problems), method configuration (techniques, architectures, module frameworks, and losses), training tactics (hyperparameter settings), and evaluation criteria (metrics). We also intend to investigate a variety of performance-enhancing methods, including ensemble and post-processing. In addition, in this survey, we highlight the primary problems associated with evaluating skin lesion segmentation and classification systems using minimal datasets, as well as the potential solutions to these plights. In conclusion, enlightening findings, recommendations, and trends are discussed for the purpose of future research surveillance in related fields of interest. It is foreseen that it will guide researchers of all levels, from beginners to experts, in the process of developing an automated and robust CAD system for skin lesion analysis.
Learning Audio Representations with MLPs
Mar 16, 2022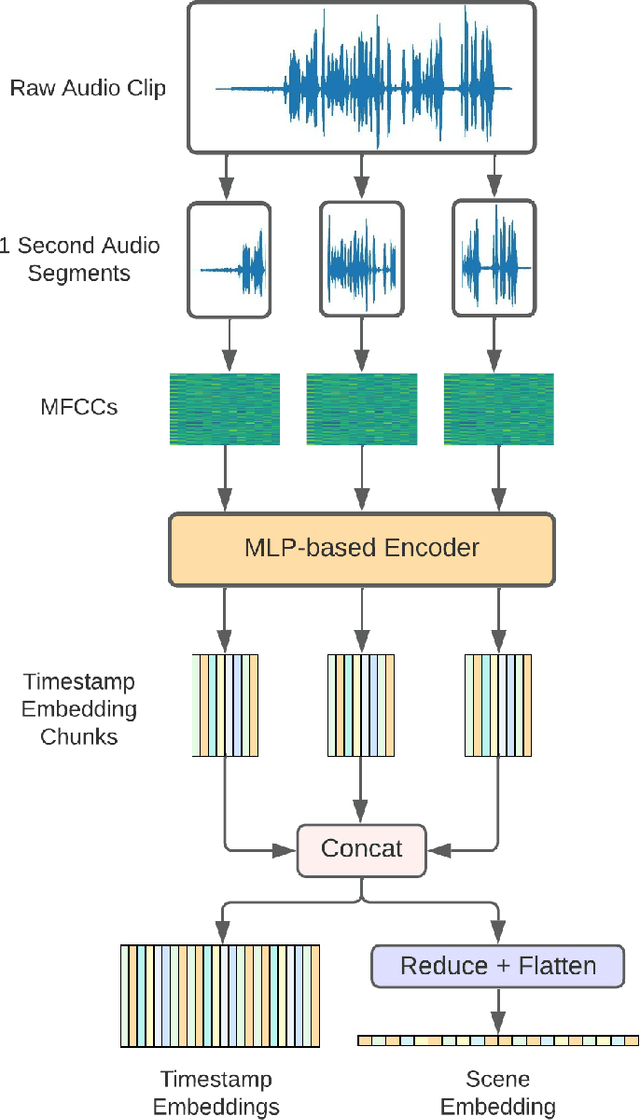
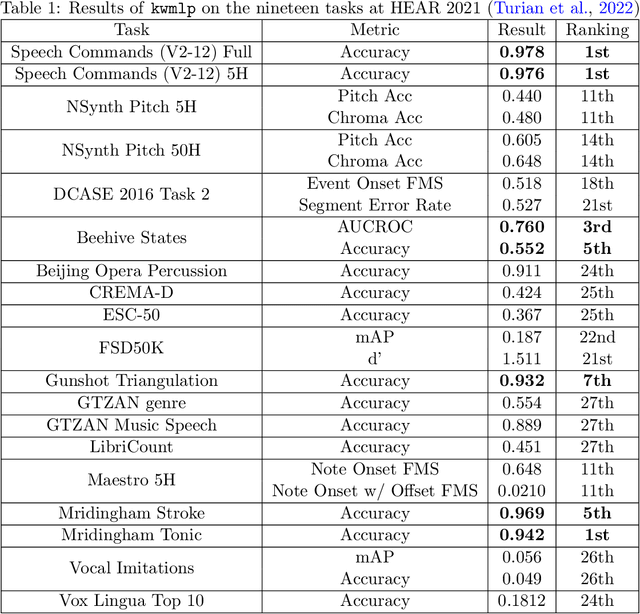
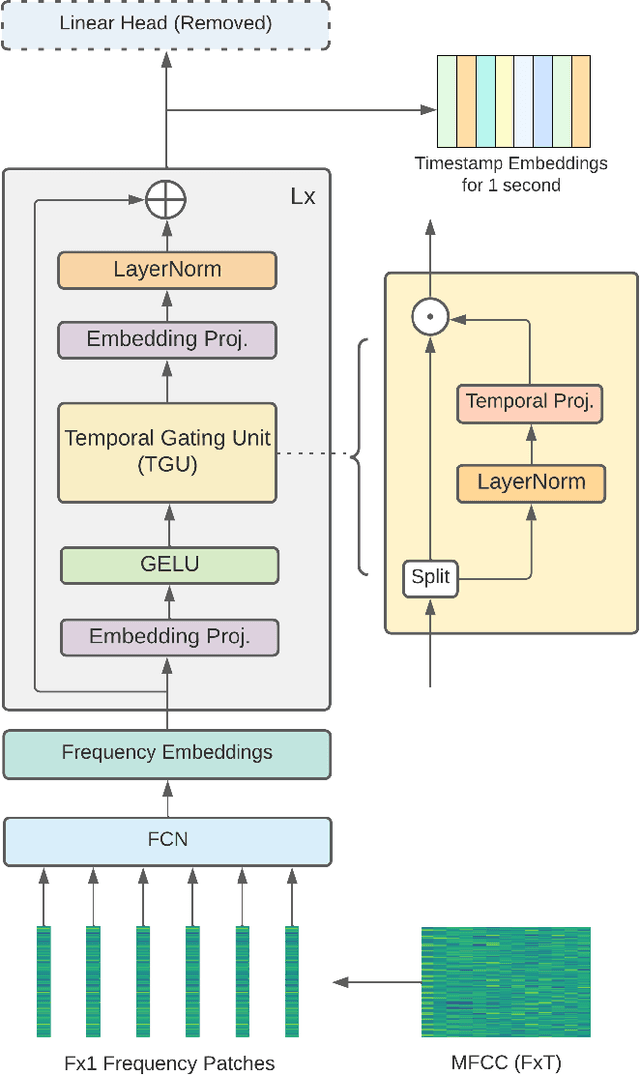
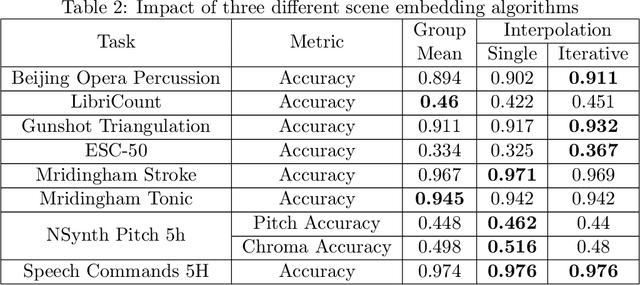
Abstract:In this paper, we propose an efficient MLP-based approach for learning audio representations, namely timestamp and scene-level audio embeddings. We use an encoder consisting of sequentially stacked gated MLP blocks, which accept 2D MFCCs as inputs. In addition, we also provide a simple temporal interpolation-based algorithm for computing scene-level embeddings from timestamp embeddings. The audio representations generated by our method are evaluated across a diverse set of benchmarks at the Holistic Evaluation of Audio Representations (HEAR) challenge, hosted at the NeurIPS 2021 competition track. We achieved first place on the Speech Commands (full), Speech Commands (5 hours), and the Mridingham Tonic benchmarks. Furthermore, our approach is also the most resource-efficient among all the submitted methods, in terms of both the number of model parameters and the time required to compute embeddings.
Grasp-and-Lift Detection from EEG Signal Using Convolutional Neural Network
Feb 12, 2022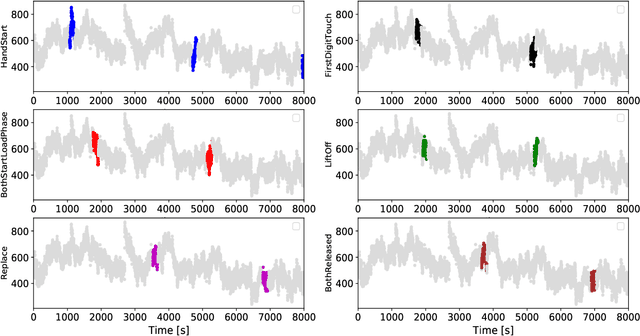
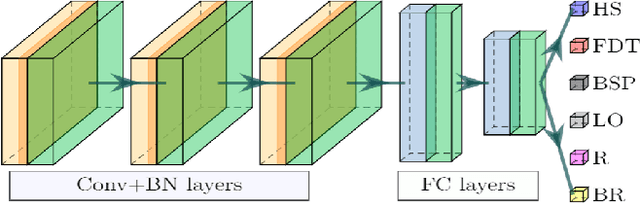
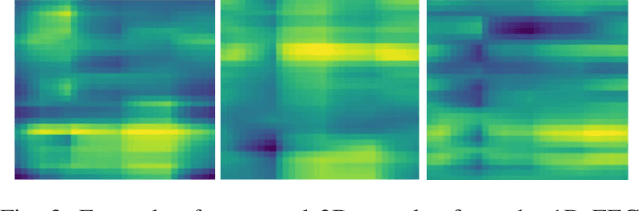
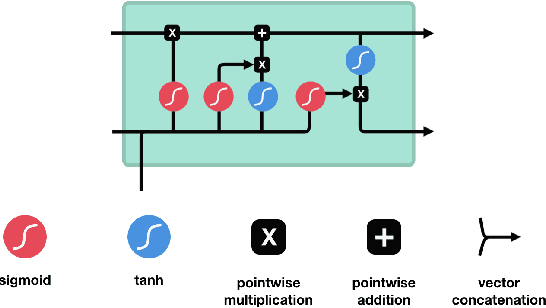
Abstract:People undergoing neuromuscular dysfunctions and amputated limbs require automatic prosthetic appliances. In developing such prostheses, the precise detection of brain motor actions is imperative for the Grasp-and-Lift (GAL) tasks. Because of the low-cost and non-invasive essence of Electroencephalography (EEG), it is widely preferred for detecting motor actions during the controls of prosthetic tools. This article has automated the hand movement activity viz GAL detection method from the 32-channel EEG signals. The proposed pipeline essentially combines preprocessing and end-to-end detection steps, eliminating the requirement of hand-crafted feature engineering. Preprocessing action consists of raw signal denoising, using either Discrete Wavelet Transform (DWT) or highpass or bandpass filtering and data standardization. The detection step consists of Convolutional Neural Network (CNN)- or Long Short Term Memory (LSTM)-based model. All the investigations utilize the publicly available WAY-EEG-GAL dataset, having six different GAL events. The best experiment reveals that the proposed framework achieves an average area under the ROC curve of 0.944, employing the DWT-based denoising filter, data standardization, and CNN-based detection model. The obtained outcome designates an excellent achievement of the introduced method in detecting GAL events from the EEG signals, turning it applicable to prosthetic appliances, brain-computer interfaces, robotic arms, etc.
VIS-iTrack: Visual Intention through Gaze Tracking using Low-Cost Webcam
Feb 05, 2022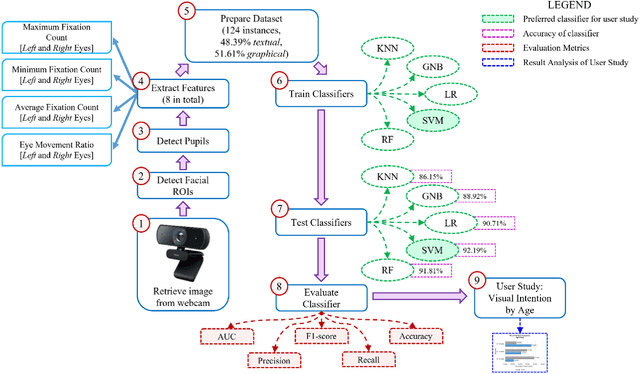
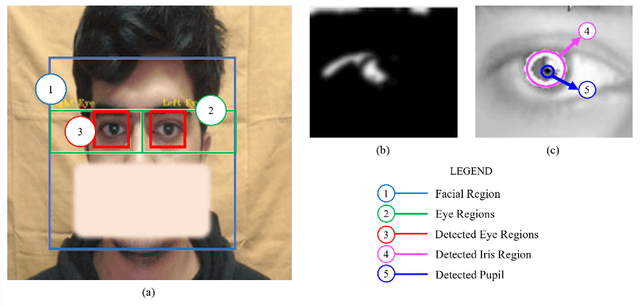
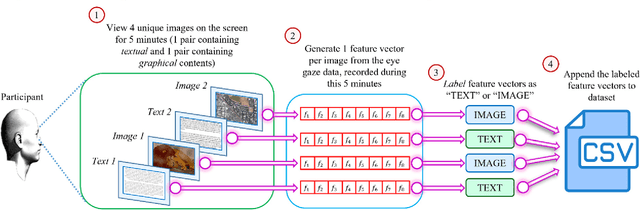
Abstract:Human intention is an internal, mental characterization for acquiring desired information. From interactive interfaces containing either textual or graphical information, intention to perceive desired information is subjective and strongly connected with eye gaze. In this work, we determine such intention by analyzing real-time eye gaze data with a low-cost regular webcam. We extracted unique features (e.g., Fixation Count, Eye Movement Ratio) from the eye gaze data of 31 participants to generate a dataset containing 124 samples of visual intention for perceiving textual or graphical information, labeled as either TEXT or IMAGE, having 48.39% and 51.61% distribution, respectively. Using this dataset, we analyzed 5 classifiers, including Support Vector Machine (SVM) (Accuracy: 92.19%). Using the trained SVM, we investigated the variation of visual intention among 30 participants, distributed in 3 age groups, and found out that young users were more leaned towards graphical contents whereas older adults felt more interested in textual ones. This finding suggests that real-time eye gaze data can be a potential source of identifying visual intention, analyzing which intention aware interactive interfaces can be designed and developed to facilitate human cognition.
Lung-Originated Tumor Segmentation from Computed Tomography Scan (LOTUS) Benchmark
Jan 03, 2022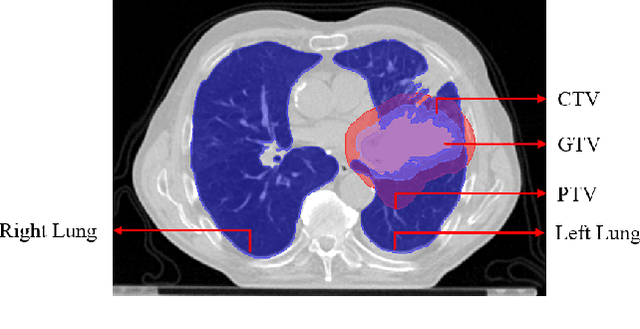
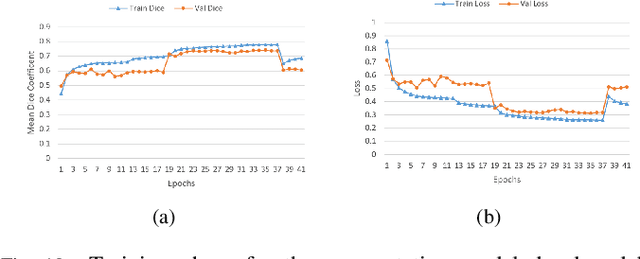
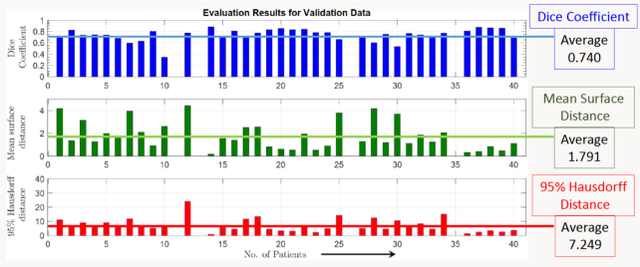
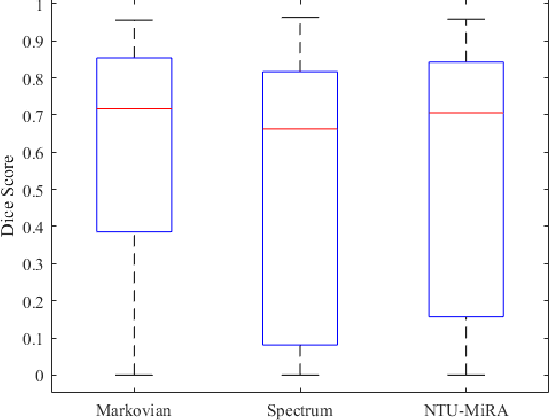
Abstract:Lung cancer is one of the deadliest cancers, and in part its effective diagnosis and treatment depend on the accurate delineation of the tumor. Human-centered segmentation, which is currently the most common approach, is subject to inter-observer variability, and is also time-consuming, considering the fact that only experts are capable of providing annotations. Automatic and semi-automatic tumor segmentation methods have recently shown promising results. However, as different researchers have validated their algorithms using various datasets and performance metrics, reliably evaluating these methods is still an open challenge. The goal of the Lung-Originated Tumor Segmentation from Computed Tomography Scan (LOTUS) Benchmark created through 2018 IEEE Video and Image Processing (VIP) Cup competition, is to provide a unique dataset and pre-defined metrics, so that different researchers can develop and evaluate their methods in a unified fashion. The 2018 VIP Cup started with a global engagement from 42 countries to access the competition data. At the registration stage, there were 129 members clustered into 28 teams from 10 countries, out of which 9 teams made it to the final stage and 6 teams successfully completed all the required tasks. In a nutshell, all the algorithms proposed during the competition, are based on deep learning models combined with a false positive reduction technique. Methods developed by the three finalists show promising results in tumor segmentation, however, more effort should be put into reducing the false positive rate. This competition manuscript presents an overview of the VIP-Cup challenge, along with the proposed algorithms and results.
Is Speech Emotion Recognition Language-Independent? Analysis of English and Bangla Languages using Language-Independent Vocal Features
Nov 21, 2021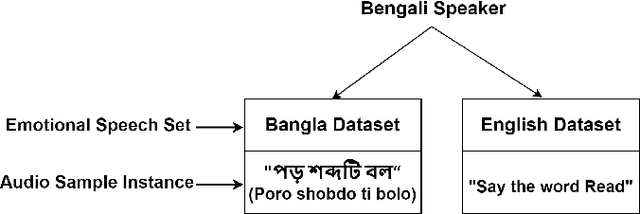
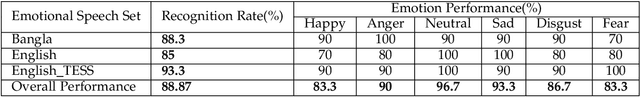
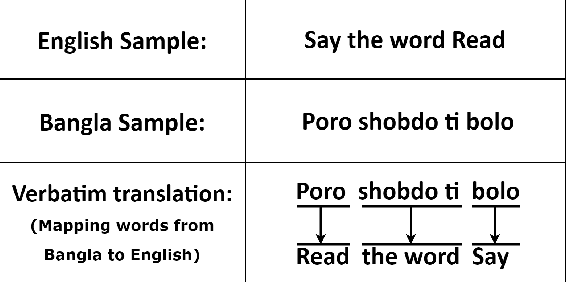
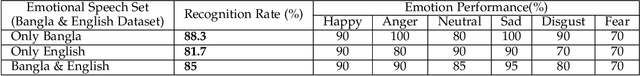
Abstract:A language agnostic approach to recognizing emotions from speech remains an incomplete and challenging task. In this paper, we used Bangla and English languages to assess whether distinguishing emotions from speech is independent of language. The following emotions were categorized for this study: happiness, anger, neutral, sadness, disgust, and fear. We employed three Emotional Speech Sets, of which the first two were developed by native Bengali speakers in Bangla and English languages separately. The third was the Toronto Emotional Speech Set (TESS), which was developed by native English speakers from Canada. We carefully selected language-independent prosodic features, adopted a Support Vector Machine (SVM) model, and conducted three experiments to carry out our proposition. In the first experiment, we measured the performance of the three speech sets individually. This was followed by the second experiment, where we recorded the classification rate by combining the speech sets. Finally, in the third experiment we measured the recognition rate by training and testing the model with different speech sets. Although this study reveals that Speech Emotion Recognition (SER) is mostly language-independent, there is some disparity while recognizing emotional states like disgust and fear in these two languages. Moreover, our investigations inferred that non-native speakers convey emotions through speech, much like expressing themselves in their native tongue.
A deep-learning--based multimodal depth-aware dynamic hand gesture recognition system
Jul 06, 2021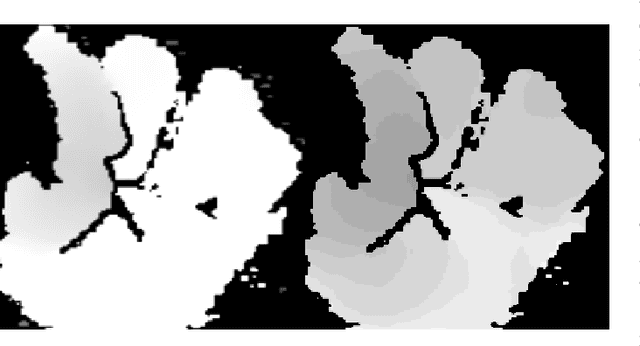
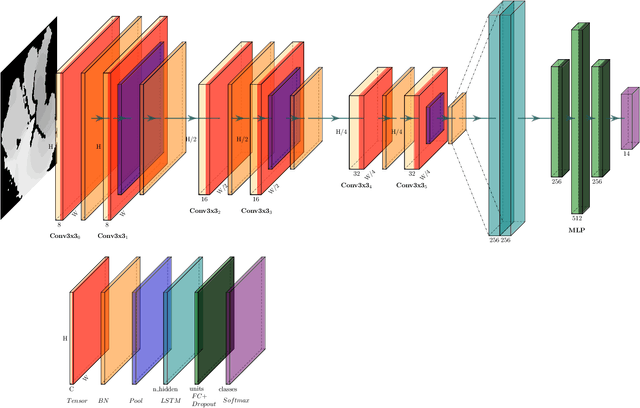

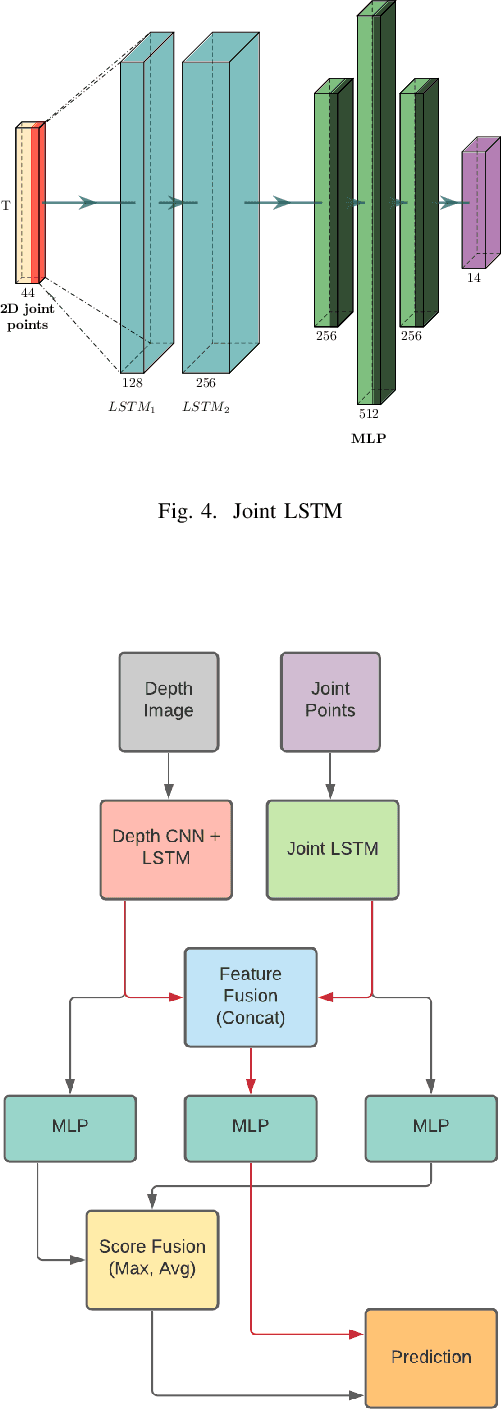
Abstract:Any spatio-temporal movement or reorientation of the hand, done with the intention of conveying a specific meaning, can be considered as a hand gesture. Inputs to hand gesture recognition systems can be in several forms, such as depth images, monocular RGB, or skeleton joint points. We observe that raw depth images possess low contrasts in the hand regions of interest (ROI). They do not highlight important details to learn, such as finger bending information (whether a finger is overlapping the palm, or another finger). Recently, in deep-learning--based dynamic hand gesture recognition, researchers are tying to fuse different input modalities (e.g. RGB or depth images and hand skeleton joint points) to improve the recognition accuracy. In this paper, we focus on dynamic hand gesture (DHG) recognition using depth quantized image features and hand skeleton joint points. In particular, we explore the effect of using depth-quantized features in Convolutional Neural Network (CNN) and Recurrent Neural Network (RNN) based multi-modal fusion networks. We find that our method improves existing results on the SHREC-DHG-14 dataset. Furthermore, using our method, we show that it is possible to reduce the resolution of the input images by more than four times and still obtain comparable or better accuracy to that of the resolutions used in previous methods.
Acute Lymphoblastic Leukemia Detection from Microscopic Images Using Weighted Ensemble of Convolutional Neural Networks
May 09, 2021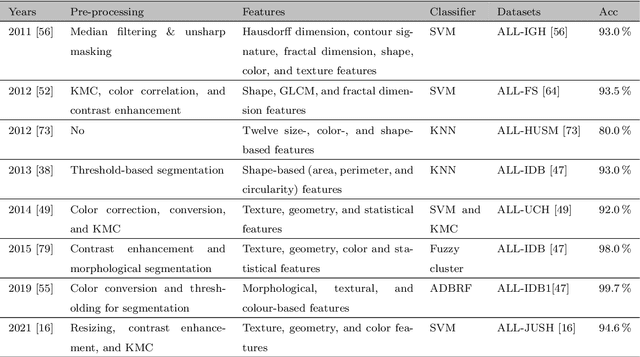
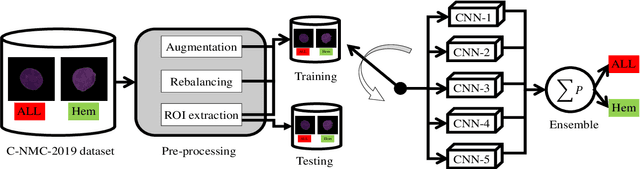
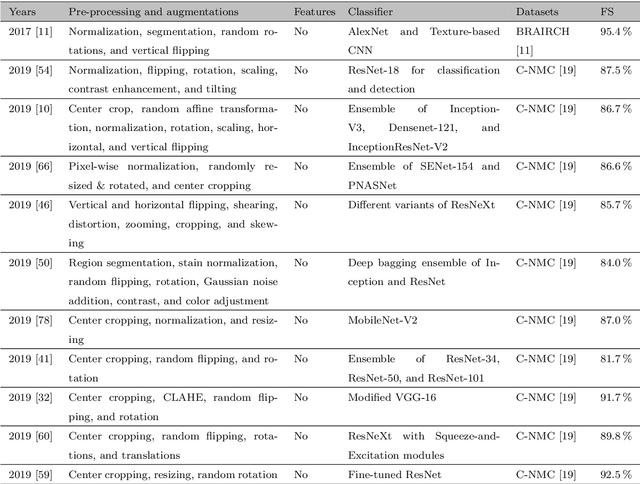
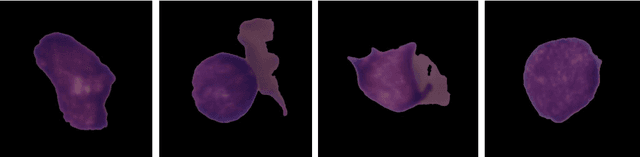
Abstract:Acute Lymphoblastic Leukemia (ALL) is a blood cell cancer characterized by numerous immature lymphocytes. Even though automation in ALL prognosis is an essential aspect of cancer diagnosis, it is challenging due to the morphological correlation between malignant and normal cells. The traditional ALL classification strategy demands experienced pathologists to carefully read the cell images, which is arduous, time-consuming, and often suffers inter-observer variations. This article has automated the ALL detection task from microscopic cell images, employing deep Convolutional Neural Networks (CNNs). We explore the weighted ensemble of different deep CNNs to recommend a better ALL cell classifier. The weights for the ensemble candidate models are estimated from their corresponding metrics, such as accuracy, F1-score, AUC, and kappa values. Various data augmentations and pre-processing are incorporated for achieving a better generalization of the network. We utilize the publicly available C-NMC-2019 ALL dataset to conduct all the comprehensive experiments. Our proposed weighted ensemble model, using the kappa values of the ensemble candidates as their weights, has outputted a weighted F1-score of 88.6 %, a balanced accuracy of 86.2 %, and an AUC of 0.941 in the preliminary test set. The qualitative results displaying the gradient class activation maps confirm that the introduced model has a concentrated learned region. In contrast, the ensemble candidate models, such as Xception, VGG-16, DenseNet-121, MobileNet, and InceptionResNet-V2, separately produce coarse and scatter learned areas for most example cases. Since the proposed kappa value-based weighted ensemble yields a better result for the aimed task in this article, it can experiment in other domains of medical diagnostic applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge