Marleen de Bruijne
for the ALFA study
Mean Field Network based Graph Refinement with application to Airway Tree Extraction
Apr 10, 2018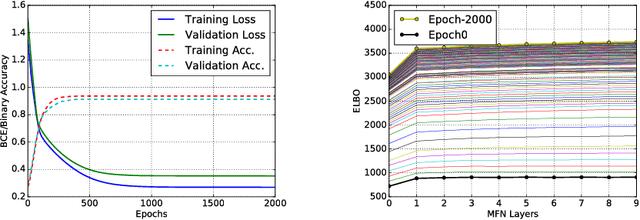

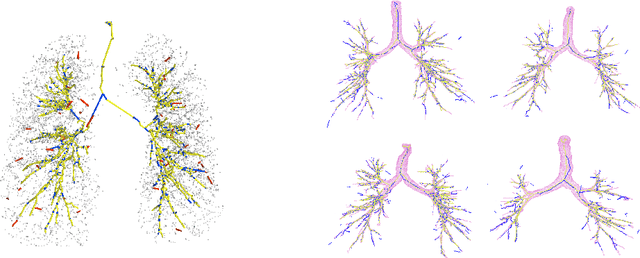
Abstract:We present tree extraction in 3D images as a graph refinement task, of obtaining a subgraph from an over-complete input graph. To this end, we formulate an approximate Bayesian inference framework on undirected graphs using mean field approximation (MFA). Mean field networks are used for inference based on the interpretation that iterations of MFA can be seen as feed-forward operations in a neural network. This allows us to learn the model parameters from training data using back-propagation algorithm. We demonstrate usefulness of the model to extract airway trees from 3D chest CT data. We first obtain probability images using a voxel classifier that distinguishes airways from background and use Bayesian smoothing to model individual airway branches. This yields us joint Gaussian density estimates of position, orientation and scale as node features of the input graph. Performance of the method is compared with two methods: the first uses probability images from a trained voxel classifier with region growing, which is similar to one of the best performing methods at EXACT'09 airway challenge, and the second method is based on Bayesian smoothing on these probability images. Using centerline distance as error measure the presented method shows significant improvement compared to these two methods.
Quantification of Lung Abnormalities in Cystic Fibrosis using Deep Networks
Mar 21, 2018Abstract:Cystic fibrosis is a genetic disease which may appear in early life with structural abnormalities in lung tissues. We propose to detect these abnormalities using a texture classification approach. Our method is a cascade of two convolutional neural networks. The first network detects the presence of abnormal tissues. The second network identifies the type of the structural abnormalities: bronchiectasis, atelectasis or mucus plugging.We also propose a network computing pixel-wise heatmaps of abnormality presence learning only from the patch-wise annotations. Our database consists of CT scans of 194 subjects. We use 154 subjects to train our algorithms and the 40 remaining ones as a test set. We compare our method with random forest and a single neural network approach. The first network reaches an accuracy of 0,94 for disease detection, 0,18 higher than the random forest classifier and 0,37 higher than the single neural network. Our cascade approach yields a final class-averaged F1-score of 0,33, outperforming the baseline method and the single network by 0,10 and 0,12.
* SPIE - Medical Imaging 2018: Image Processing
Transfer learning for multi-center classification of chronic obstructive pulmonary disease
Nov 23, 2017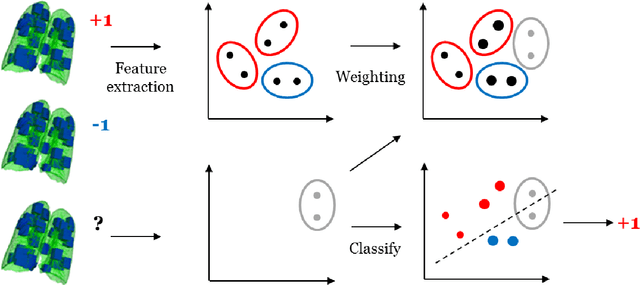
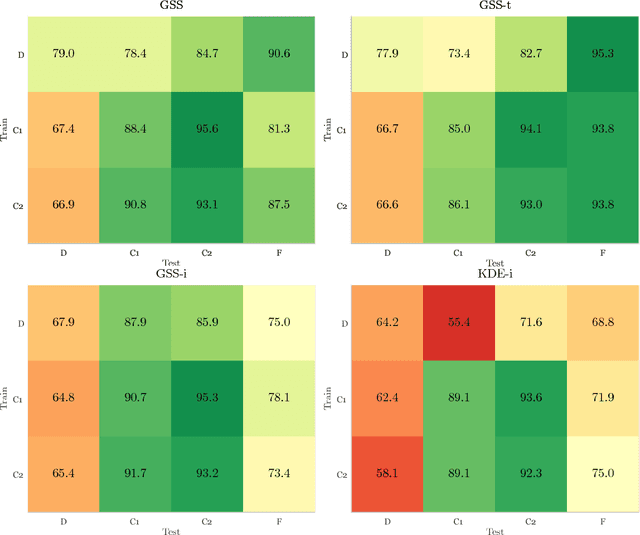
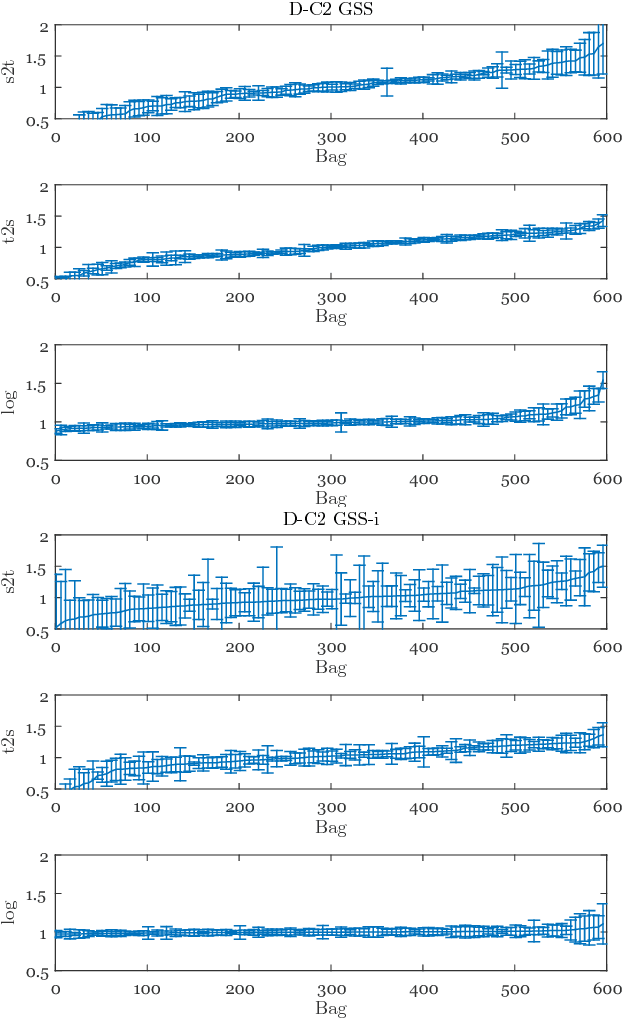
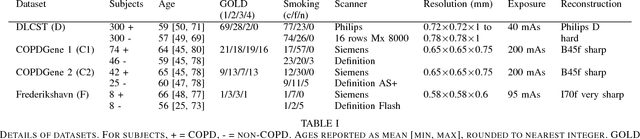
Abstract:Chronic obstructive pulmonary disease (COPD) is a lung disease which can be quantified using chest computed tomography (CT) scans. Recent studies have shown that COPD can be automatically diagnosed using weakly supervised learning of intensity and texture distributions. However, up till now such classifiers have only been evaluated on scans from a single domain, and it is unclear whether they would generalize across domains, such as different scanners or scanning protocols. To address this problem, we investigate classification of COPD in a multi-center dataset with a total of 803 scans from three different centers, four different scanners, with heterogenous subject distributions. Our method is based on Gaussian texture features, and a weighted logistic classifier, which increases the weights of samples similar to the test data. We show that Gaussian texture features outperform intensity features previously used in multi-center classification tasks. We also show that a weighting strategy based on a classifier that is trained to discriminate between scans from different domains, can further improve the results. To encourage further research into transfer learning methods for classification of COPD, upon acceptance of the paper we will release two feature datasets used in this study on http://bigr.nl/research/projects/copd
Extraction of Airways with Probabilistic State-space Models and Bayesian Smoothing
Aug 07, 2017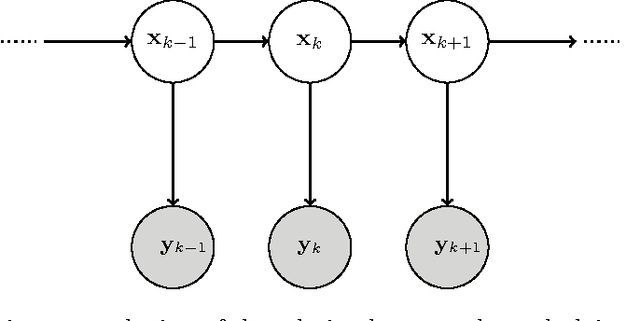
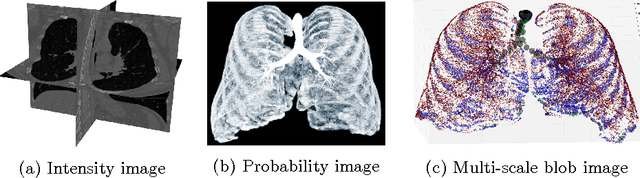
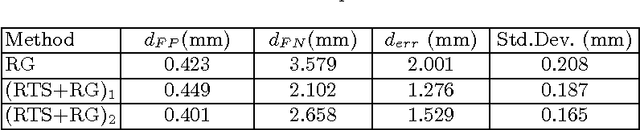
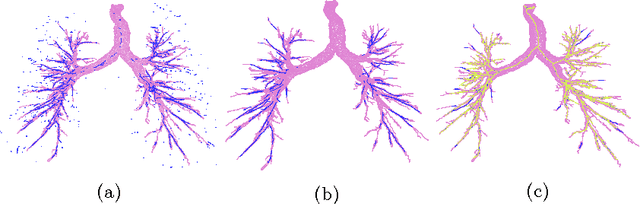
Abstract:Segmenting tree structures is common in several image processing applications. In medical image analysis, reliable segmentations of airways, vessels, neurons and other tree structures can enable important clinical applications. We present a framework for tracking tree structures comprising of elongated branches using probabilistic state-space models and Bayesian smoothing. Unlike most existing methods that proceed with sequential tracking of branches, we present an exploratory method, that is less sensitive to local anomalies in the data due to acquisition noise and/or interfering structures. The evolution of individual branches is modelled using a process model and the observed data is incorporated into the update step of the Bayesian smoother using a measurement model that is based on a multi-scale blob detector. Bayesian smoothing is performed using the RTS (Rauch-Tung-Striebel) smoother, which provides Gaussian density estimates of branch states at each tracking step. We select likely branch seed points automatically based on the response of the blob detection and track from all such seed points using the RTS smoother. We use covariance of the marginal posterior density estimated for each branch to discriminate false positive and true positive branches. The method is evaluated on 3D chest CT scans to track airways. We show that the presented method results in additional branches compared to a baseline method based on region growing on probability images.
Early Experiences with Crowdsourcing Airway Annotations in Chest CT
Jun 07, 2017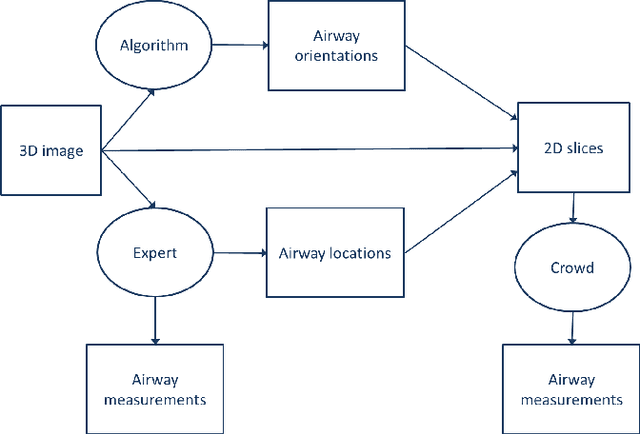
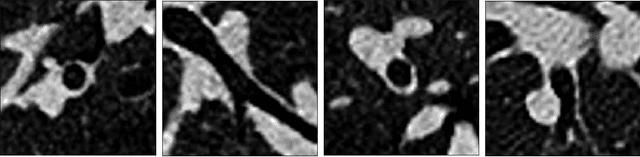
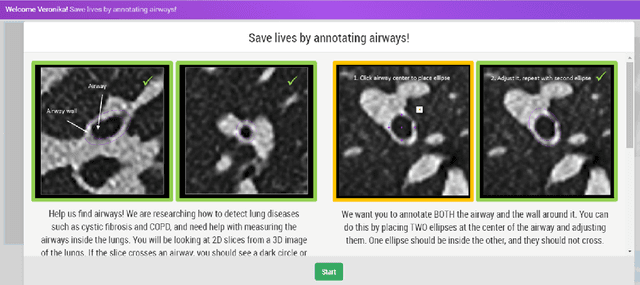
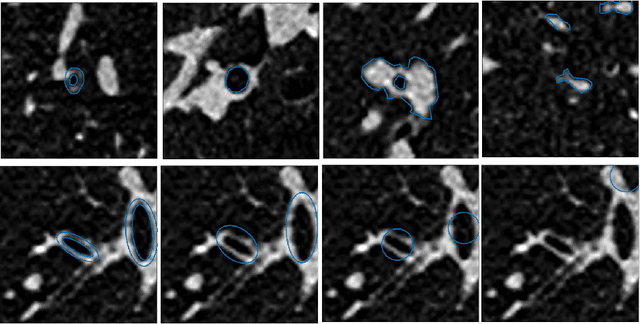
Abstract:Measuring airways in chest computed tomography (CT) images is important for characterizing diseases such as cystic fibrosis, yet very time-consuming to perform manually. Machine learning algorithms offer an alternative, but need large sets of annotated data to perform well. We investigate whether crowdsourcing can be used to gather airway annotations which can serve directly for measuring the airways, or as training data for the algorithms. We generate image slices at known locations of airways and request untrained crowd workers to outline the airway lumen and airway wall. Our results show that the workers are able to interpret the images, but that the instructions are too complex, leading to many unusable annotations. After excluding unusable annotations, quantitative results show medium to high correlations with expert measurements of the airways. Based on this positive experience, we describe a number of further research directions and provide insight into the challenges of crowdsourcing in medical images from the perspective of first-time users.
Segmentation of Intracranial Arterial Calcification with Deeply Supervised Residual Dropout Networks
Jun 04, 2017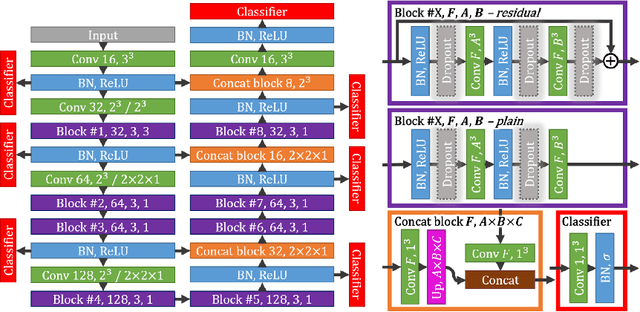
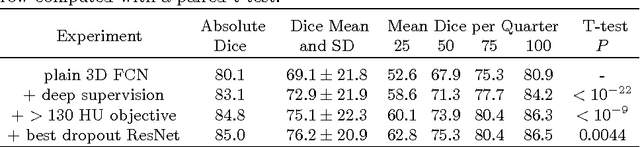
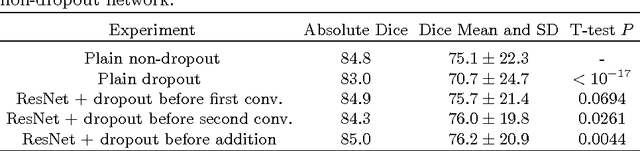
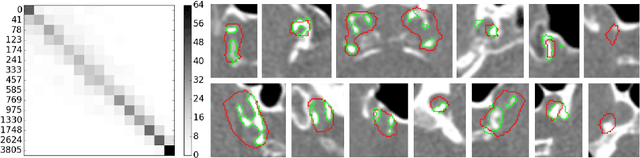
Abstract:Intracranial carotid artery calcification (ICAC) is a major risk factor for stroke, and might contribute to dementia and cognitive decline. Reliance on time-consuming manual annotation of ICAC hampers much demanded further research into the relationship between ICAC and neurological diseases. Automation of ICAC segmentation is therefore highly desirable, but difficult due to the proximity of the lesions to bony structures with a similar attenuation coefficient. In this paper, we propose a method for automatic segmentation of ICAC; the first to our knowledge. Our method is based on a 3D fully convolutional neural network that we extend with two regularization techniques. Firstly, we use deep supervision (hidden layers supervision) to encourage discriminative features in the hidden layers. Secondly, we augment the network with skip connections, as in the recently developed ResNet, and dropout layers, inserted in a way that skip connections circumvent them. We investigate the effect of skip connections and dropout. In addition, we propose a simple problem-specific modification of the network objective function that restricts the focus to the most important image regions and simplifies the optimization. We train and validate our model using 882 CT scans and test on 1,000. Our regularization techniques and objective improve the average Dice score by 7.1%, yielding an average Dice of 76.2% and 97.7% correlation between predicted ICAC volumes and manual annotations.
Label Stability in Multiple Instance Learning
Mar 15, 2017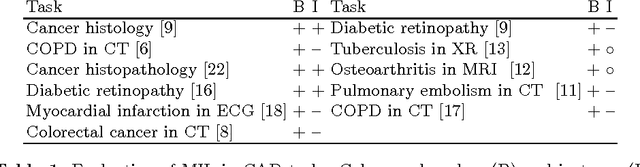
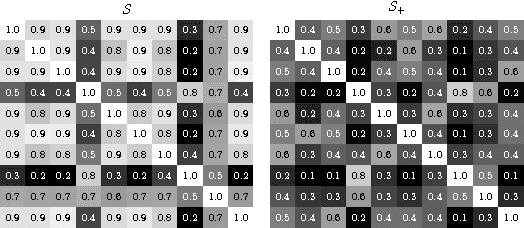

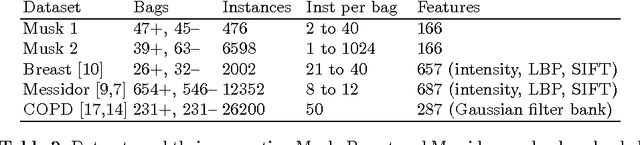
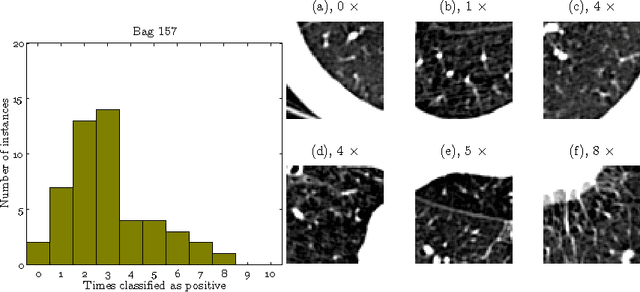
Abstract:We address the problem of \emph{instance label stability} in multiple instance learning (MIL) classifiers. These classifiers are trained only on globally annotated images (bags), but often can provide fine-grained annotations for image pixels or patches (instances). This is interesting for computer aided diagnosis (CAD) and other medical image analysis tasks for which only a coarse labeling is provided. Unfortunately, the instance labels may be unstable. This means that a slight change in training data could potentially lead to abnormalities being detected in different parts of the image, which is undesirable from a CAD point of view. Despite MIL gaining popularity in the CAD literature, this issue has not yet been addressed. We investigate the stability of instance labels provided by several MIL classifiers on 5 different datasets, of which 3 are medical image datasets (breast histopathology, diabetic retinopathy and computed tomography lung images). We propose an unsupervised measure to evaluate instance stability, and demonstrate that a performance-stability trade-off can be made when comparing MIL classifiers.
Transfer Learning by Asymmetric Image Weighting for Segmentation across Scanners
Mar 15, 2017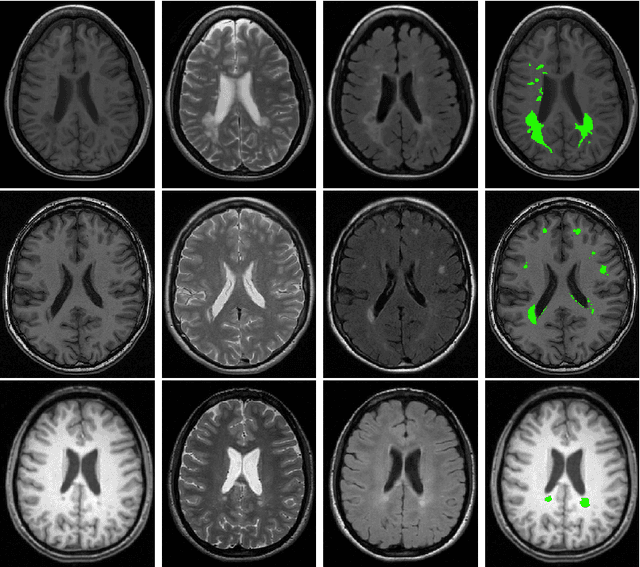
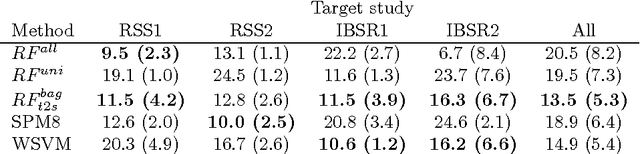
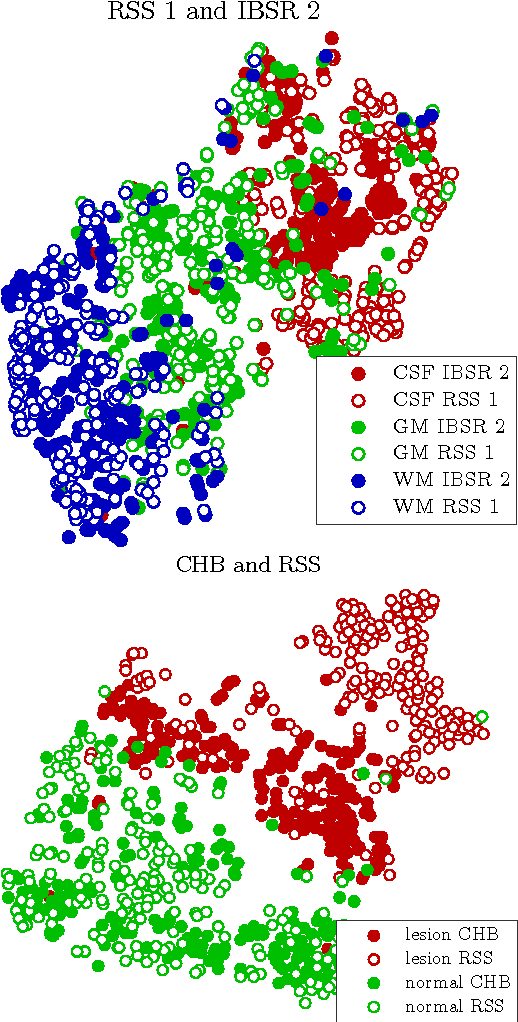
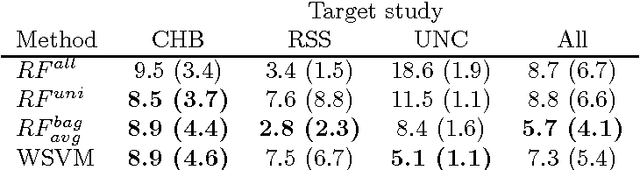
Abstract:Supervised learning has been very successful for automatic segmentation of images from a single scanner. However, several papers report deteriorated performances when using classifiers trained on images from one scanner to segment images from other scanners. We propose a transfer learning classifier that adapts to differences between training and test images. This method uses a weighted ensemble of classifiers trained on individual images. The weight of each classifier is determined by the similarity between its training image and the test image. We examine three unsupervised similarity measures, which can be used in scenarios where no labeled data from a newly introduced scanner or scanning protocol is available. The measures are based on a divergence, a bag distance, and on estimating the labels with a clustering procedure. These measures are asymmetric. We study whether the asymmetry can improve classification. Out of the three similarity measures, the bag similarity measure is the most robust across different studies and achieves excellent results on four brain tissue segmentation datasets and three white matter lesion segmentation datasets, acquired at different centers and with different scanners and scanning protocols. We show that the asymmetry can indeed be informative, and that computing the similarity from the test image to the training images is more appropriate than the opposite direction.
Classification of COPD with Multiple Instance Learning
Mar 15, 2017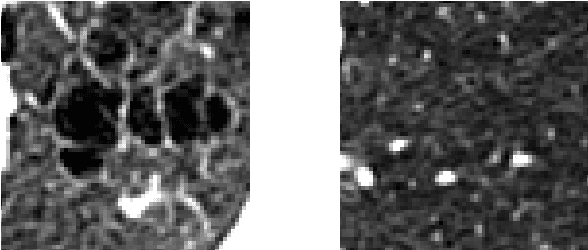
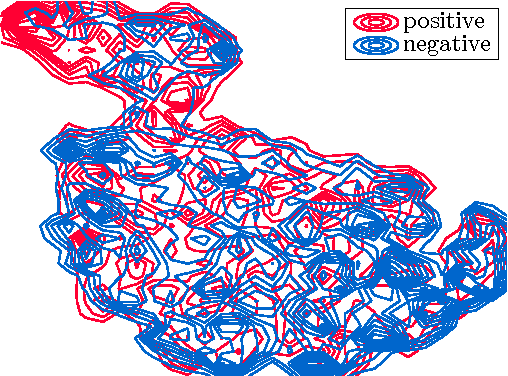

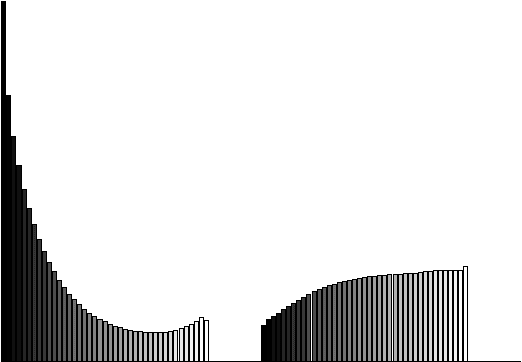
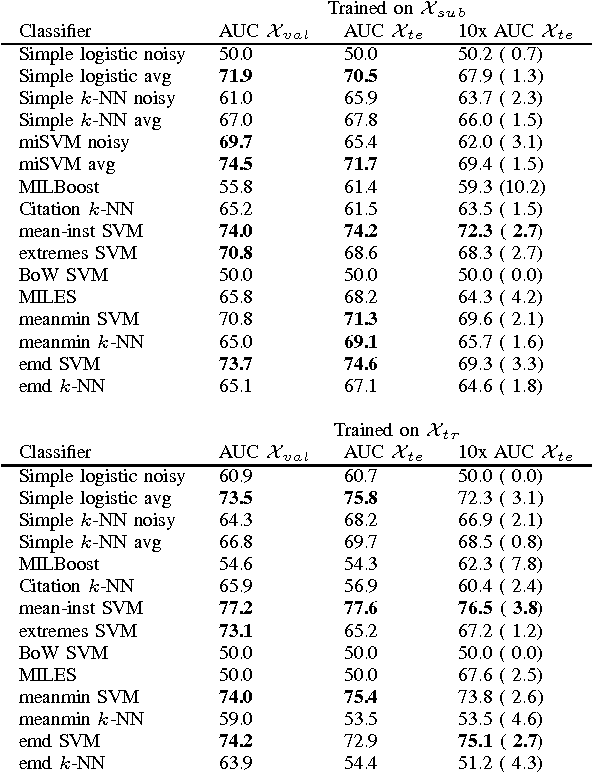
Abstract:Chronic obstructive pulmonary disease (COPD) is a lung disease where early detection benefits the survival rate. COPD can be quantified by classifying patches of computed tomography images, and combining patch labels into an overall diagnosis for the image. As labeled patches are often not available, image labels are propagated to the patches, incorrectly labeling healthy patches in COPD patients as being affected by the disease. We approach quantification of COPD from lung images as a multiple instance learning (MIL) problem, which is more suitable for such weakly labeled data. We investigate various MIL assumptions in the context of COPD and show that although a concept region with COPD-related disease patterns is present, considering the whole distribution of lung tissue patches improves the performance. The best method is based on averaging instances and obtains an AUC of 0.742, which is higher than the previously reported best of 0.713 on the same dataset. Using the full training set further increases performance to 0.776, which is significantly higher (DeLong test) than previous results.
Extraction of airway trees using multiple hypothesis tracking and template matching
Nov 24, 2016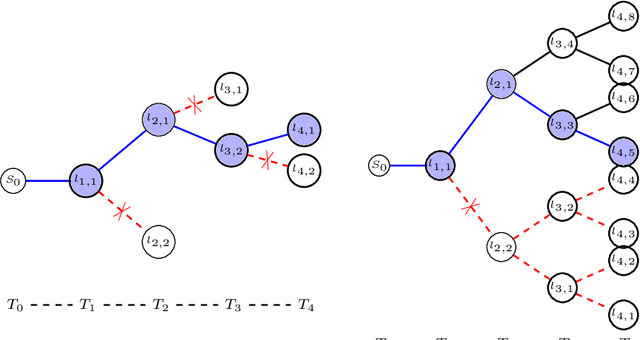
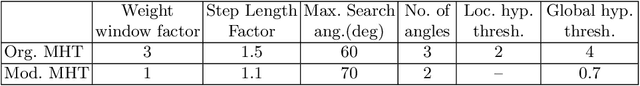
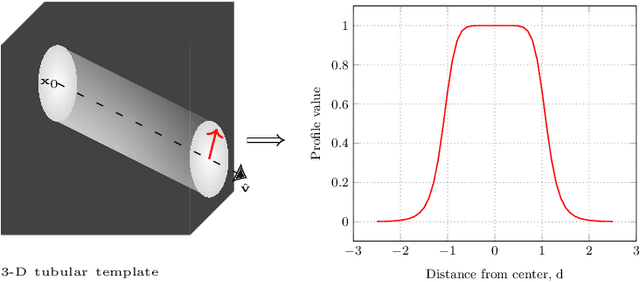
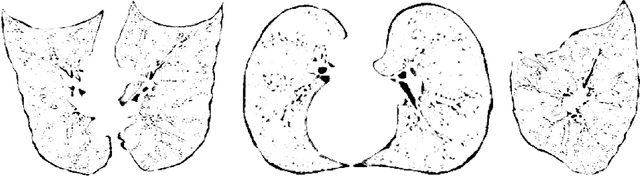
Abstract:Knowledge of airway tree morphology has important clinical applications in diagnosis of chronic obstructive pulmonary disease. We present an automatic tree extraction method based on multiple hypothesis tracking and template matching for this purpose and evaluate its performance on chest CT images. The method is adapted from a semi-automatic method devised for vessel segmentation. Idealized tubular templates are constructed that match airway probability obtained from a trained classifier and ranked based on their relative significance. Several such regularly spaced templates form the local hypotheses used in constructing a multiple hypothesis tree, which is then traversed to reach decisions. The proposed modifications remove the need for local thresholding of hypotheses as decisions are made entirely based on statistical comparisons involving the hypothesis tree. The results show improvements in performance when compared to the original method and region growing on intensity images. We also compare the method with region growing on the probability images, where the presented method does not show substantial improvement, but we expect it to be less sensitive to local anomalies in the data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge