Aasa Feragen
Fair Lung Disease Diagnosis from Chest CT via Gender-Adversarial Attention Multiple Instance Learning
Mar 13, 2026Abstract:We present a fairness-aware framework for multi-class lung disease diagnosis from chest CT volumes, developed for the Fair Disease Diagnosis Challenge at the PHAROS-AIF-MIH Workshop (CVPR 2026). The challenge requires classifying CT scans into four categories -- Healthy, COVID-19, Adenocarcinoma, and Squamous Cell Carcinoma -- with performance measured as the average of per-gender macro F1 scores, explicitly penalizing gender-inequitable predictions. Our approach addresses two core difficulties: the sparse pathological signal across hundreds of slices, and a severe demographic imbalance compounded across disease class and gender. We propose an attention-based Multiple Instance Learning (MIL) model on a ConvNeXt backbone that learns to identify diagnostically relevant slices without slice-level supervision, augmented with a Gradient Reversal Layer (GRL) that adversarially suppresses gender-predictive structure in the learned scan representation. Training incorporates focal loss with label smoothing, stratified cross-validation over joint (class, gender) strata, and targeted oversampling of the most underrepresented subgroup. At inference, all five-fold checkpoints are ensembled with horizontal-flip test-time augmentation via soft logit voting and out-of-the-fold threshold optimization for robustness. Our model achieves a mean validation competition score of 0.685 (std - 0.030), with the best single fold reaching 0.759. All training and inference code is publicly available at https://github.com/ADE-17/cvpr-fair-chest-ct
Measuring What VLMs Don't Say: Validation Metrics Hide Clinical Terminology Erasure in Radiology Report Generation
Mar 02, 2026Abstract:Reliable deployment of Vision-Language Models (VLMs) in radiology requires validation metrics that go beyond surface-level text similarity to ensure clinical fidelity and demographic fairness. This paper investigates a critical blind spot in current model evaluation: the use of decoding strategies that lead to high aggregate token-overlap scores despite succumbing to template collapse, in which models generate only repetitive, safe generic text and omit clinical terminology. Unaddressed, this blind spot can lead to metric gaming, where models that perform well on benchmarks prove clinically uninformative. Instead, we advocate for lexical diversity measures to check model generations for clinical specificity. We introduce Clinical Association Displacement (CAD), a vocabulary-level framework that quantifies shifts in demographic-based word associations in generated reports. Weighted Association Erasure (WAE) aggregates these shifts to measure the clinical signal loss across demographic groups. We show that deterministic decoding produces high levels of semantic erasure, while stochastic sampling generates diverse outputs but risks introducing new bias, motivating a fundamental rethink of how "optimal" reporting is defined.
Weight Space Correlation Analysis: Quantifying Feature Utilization in Deep Learning Models
Dec 15, 2025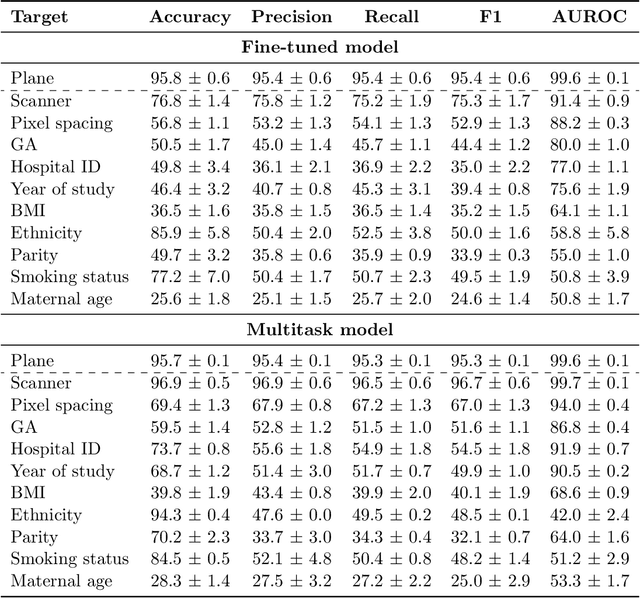
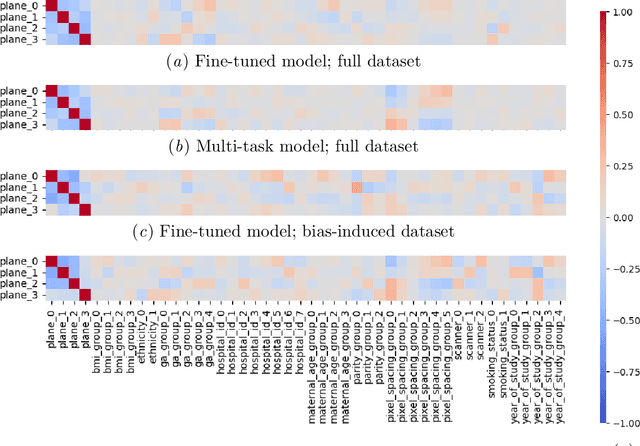
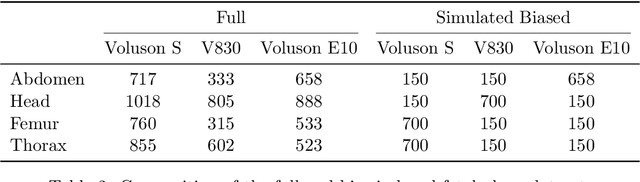

Abstract:Deep learning models in medical imaging are susceptible to shortcut learning, relying on confounding metadata (e.g., scanner model) that is often encoded in image embeddings. The crucial question is whether the model actively utilizes this encoded information for its final prediction. We introduce Weight Space Correlation Analysis, an interpretable methodology that quantifies feature utilization by measuring the alignment between the classification heads of a primary clinical task and auxiliary metadata tasks. We first validate our method by successfully detecting artificially induced shortcut learning. We then apply it to probe the feature utilization of an SA-SonoNet model trained for Spontaneous Preterm Birth (sPTB) prediction. Our analysis confirmed that while the embeddings contain substantial metadata, the sPTB classifier's weight vectors were highly correlated with clinically relevant factors (e.g., birth weight) but decoupled from clinically irrelevant acquisition factors (e.g. scanner). Our methodology provides a tool to verify model trustworthiness, demonstrating that, in the absence of induced bias, the clinical model selectively utilizes features related to the genuine clinical signal.
Who Does Your Algorithm Fail? Investigating Age and Ethnic Bias in the MAMA-MIA Dataset
Oct 31, 2025Abstract:Deep learning models aim to improve diagnostic workflows, but fairness evaluation remains underexplored beyond classification, e.g., in image segmentation. Unaddressed segmentation bias can lead to disparities in the quality of care for certain populations, potentially compounded across clinical decision points and amplified through iterative model development. Here, we audit the fairness of the automated segmentation labels provided in the breast cancer tumor segmentation dataset MAMA-MIA. We evaluate automated segmentation quality across age, ethnicity, and data source. Our analysis reveals an intrinsic age-related bias against younger patients that continues to persist even after controlling for confounding factors, such as data source. We hypothesize that this bias may be linked to physiological factors, a known challenge for both radiologists and automated systems. Finally, we show how aggregating data from multiple data sources influences site-specific ethnic biases, underscoring the necessity of investigating data at a granular level.
General Methods Make Great Domain-specific Foundation Models: A Case-study on Fetal Ultrasound
Jun 24, 2025Abstract:With access to large-scale, unlabeled medical datasets, researchers are confronted with two questions: Should they attempt to pretrain a custom foundation model on this medical data, or use transfer-learning from an existing generalist model? And, if a custom model is pretrained, are novel methods required? In this paper we explore these questions by conducting a case-study, in which we train a foundation model on a large regional fetal ultrasound dataset of 2M images. By selecting the well-established DINOv2 method for pretraining, we achieve state-of-the-art results on three fetal ultrasound datasets, covering data from different countries, classification, segmentation, and few-shot tasks. We compare against a series of models pretrained on natural images, ultrasound images, and supervised baselines. Our results demonstrate two key insights: (i) Pretraining on custom data is worth it, even if smaller models are trained on less data, as scaling in natural image pretraining does not translate to ultrasound performance. (ii) Well-tuned methods from computer vision are making it feasible to train custom foundation models for a given medical domain, requiring no hyperparameter tuning and little methodological adaptation. Given these findings, we argue that a bias towards methodological innovation should be avoided when developing domain specific foundation models under common computational resource constraints.
Patronus: Bringing Transparency to Diffusion Models with Prototypes
Mar 28, 2025



Abstract:Diffusion-based generative models, such as Denoising Diffusion Probabilistic Models (DDPMs), have achieved remarkable success in image generation, but their step-by-step denoising process remains opaque, leaving critical aspects of the generation mechanism unexplained. To address this, we introduce \emph{Patronus}, an interpretable diffusion model inspired by ProtoPNet. Patronus integrates a prototypical network into DDPMs, enabling the extraction of prototypes and conditioning of the generation process on their prototype activation vector. This design enhances interpretability by showing the learned prototypes and how they influence the generation process. Additionally, the model supports downstream tasks like image manipulation, enabling more transparent and controlled modifications. Moreover, Patronus could reveal shortcut learning in the generation process by detecting unwanted correlations between learned prototypes. Notably, Patronus operates entirely without any annotations or text prompts. This work opens new avenues for understanding and controlling diffusion models through prototype-based interpretability. Our code is available at \href{https://github.com/nina-weng/patronus}{https://github.com/nina-weng/patronus}.
Are generative models fair? A study of racial bias in dermatological image generation
Jan 20, 2025



Abstract:Racial bias in medicine, particularly in dermatology, presents significant ethical and clinical challenges. It often results from the underrepresentation of darker skin tones in training datasets for machine learning models. While efforts to address bias in dermatology have focused on improving dataset diversity and mitigating disparities in discriminative models, the impact of racial bias on generative models remains underexplored. Generative models, such as Variational Autoencoders (VAEs), are increasingly used in healthcare applications, yet their fairness across diverse skin tones is currently not well understood. In this study, we evaluate the fairness of generative models in clinical dermatology with respect to racial bias. For this purpose, we first train a VAE with a perceptual loss to generate and reconstruct high-quality skin images across different skin tones. We utilize the Fitzpatrick17k dataset to examine how racial bias influences the representation and performance of these models. Our findings indicate that the VAE is influenced by the diversity of skin tones in the training dataset, with better performance observed for lighter skin tones. Additionally, the uncertainty estimates produced by the VAE are ineffective in assessing the model's fairness. These results highlight the need for improved uncertainty quantification mechanisms to detect and address racial bias in generative models for trustworthy healthcare technologies.
In the Picture: Medical Imaging Datasets, Artifacts, and their Living Review
Jan 18, 2025


Abstract:Datasets play a critical role in medical imaging research, yet issues such as label quality, shortcuts, and metadata are often overlooked. This lack of attention may harm the generalizability of algorithms and, consequently, negatively impact patient outcomes. While existing medical imaging literature reviews mostly focus on machine learning (ML) methods, with only a few focusing on datasets for specific applications, these reviews remain static -- they are published once and not updated thereafter. This fails to account for emerging evidence, such as biases, shortcuts, and additional annotations that other researchers may contribute after the dataset is published. We refer to these newly discovered findings of datasets as research artifacts. To address this gap, we propose a living review that continuously tracks public datasets and their associated research artifacts across multiple medical imaging applications. Our approach includes a framework for the living review to monitor data documentation artifacts, and an SQL database to visualize the citation relationships between research artifact and dataset. Lastly, we discuss key considerations for creating medical imaging datasets, review best practices for data annotation, discuss the significance of shortcuts and demographic diversity, and emphasize the importance of managing datasets throughout their entire lifecycle. Our demo is publicly available at http://130.226.140.142.
Graph Counterfactual Explainable AI via Latent Space Traversal
Jan 15, 2025Abstract:Explaining the predictions of a deep neural network is a nontrivial task, yet high-quality explanations for predictions are often a prerequisite for practitioners to trust these models. Counterfactual explanations aim to explain predictions by finding the ''nearest'' in-distribution alternative input whose prediction changes in a pre-specified way. However, it remains an open question how to define this nearest alternative input, whose solution depends on both the domain (e.g. images, graphs, tabular data, etc.) and the specific application considered. For graphs, this problem is complicated i) by their discrete nature, as opposed to the continuous nature of state-of-the-art graph classifiers; and ii) by the node permutation group acting on the graphs. We propose a method to generate counterfactual explanations for any differentiable black-box graph classifier, utilizing a case-specific permutation equivariant graph variational autoencoder. We generate counterfactual explanations in a continuous fashion by traversing the latent space of the autoencoder across the classification boundary of the classifier, allowing for seamless integration of discrete graph structure and continuous graph attributes. We empirically validate the approach on three graph datasets, showing that our model is consistently high-performing and more robust than the baselines.
* Published at Northern Lights Deep Learning Conference 2025
Counterfactual Explanations via Riemannian Latent Space Traversal
Nov 04, 2024Abstract:The adoption of increasingly complex deep models has fueled an urgent need for insight into how these models make predictions. Counterfactual explanations form a powerful tool for providing actionable explanations to practitioners. Previously, counterfactual explanation methods have been designed by traversing the latent space of generative models. Yet, these latent spaces are usually greatly simplified, with most of the data distribution complexity contained in the decoder rather than the latent embedding. Thus, traversing the latent space naively without taking the nonlinear decoder into account can lead to unnatural counterfactual trajectories. We introduce counterfactual explanations obtained using a Riemannian metric pulled back via the decoder and the classifier under scrutiny. This metric encodes information about the complex geometric structure of the data and the learned representation, enabling us to obtain robust counterfactual trajectories with high fidelity, as demonstrated by our experiments in real-world tabular datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge