Marc-André Weber
*: shared first/last authors
VERIDAH: Solving Enumeration Anomaly Aware Vertebra Labeling across Imaging Sequences
Jan 20, 2026Abstract:The human spine commonly consists of seven cervical, twelve thoracic, and five lumbar vertebrae. However, enumeration anomalies may result in individuals having eleven or thirteen thoracic vertebrae and four or six lumbar vertebrae. Although the identification of enumeration anomalies has potential clinical implications for chronic back pain and operation planning, the thoracolumbar junction is often poorly assessed and rarely described in clinical reports. Additionally, even though multiple deep-learning-based vertebra labeling algorithms exist, there is a lack of methods to automatically label enumeration anomalies. Our work closes that gap by introducing "Vertebra Identification with Anomaly Handling" (VERIDAH), a novel vertebra labeling algorithm based on multiple classification heads combined with a weighted vertebra sequence prediction algorithm. We show that our approach surpasses existing models on T2w TSE sagittal (98.30% vs. 94.24% of subjects with all vertebrae correctly labeled, p < 0.001) and CT imaging (99.18% vs. 77.26% of subjects with all vertebrae correctly labeled, p < 0.001) and works in arbitrary field-of-view images. VERIDAH correctly labeled the presence 2 Möller et al. of thoracic enumeration anomalies in 87.80% and 96.30% of T2w and CT images, respectively, and lumbar enumeration anomalies in 94.48% and 97.22% for T2w and CT, respectively. Our code and models are available at: https://github.com/Hendrik-code/spineps.
The Brain Tumor Segmentation Challenge 2023: Local Synthesis of Healthy Brain Tissue via Inpainting
May 15, 2023



Abstract:A myriad of algorithms for the automatic analysis of brain MR images is available to support clinicians in their decision-making. For brain tumor patients, the image acquisition time series typically starts with a scan that is already pathological. This poses problems, as many algorithms are designed to analyze healthy brains and provide no guarantees for images featuring lesions. Examples include but are not limited to algorithms for brain anatomy parcellation, tissue segmentation, and brain extraction. To solve this dilemma, we introduce the BraTS 2023 inpainting challenge. Here, the participants' task is to explore inpainting techniques to synthesize healthy brain scans from lesioned ones. The following manuscript contains the task formulation, dataset, and submission procedure. Later it will be updated to summarize the findings of the challenge. The challenge is organized as part of the BraTS 2023 challenge hosted at the MICCAI 2023 conference in Vancouver, Canada.
Improving 3D convolutional neural network comprehensibility via interactive visualization of relevance maps: Evaluation in Alzheimer's disease
Dec 18, 2020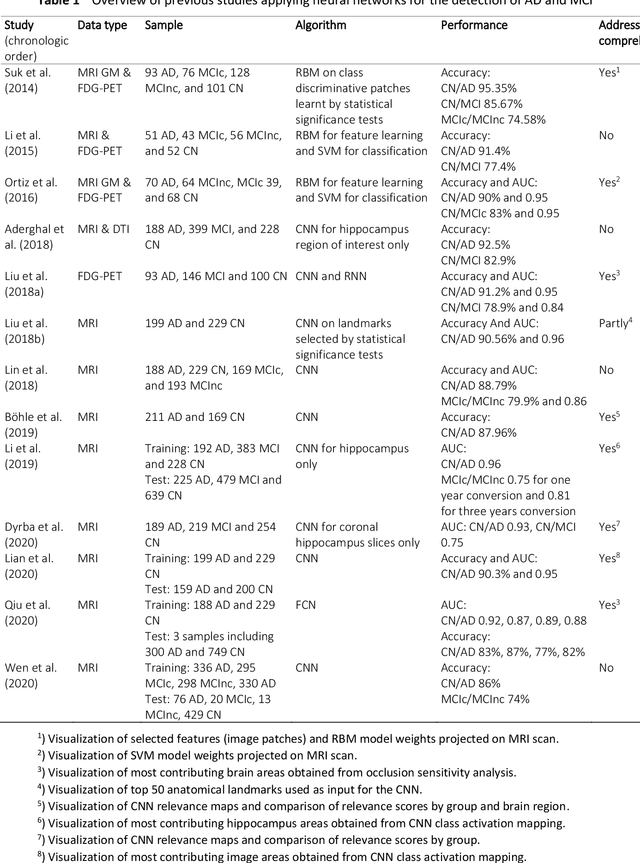
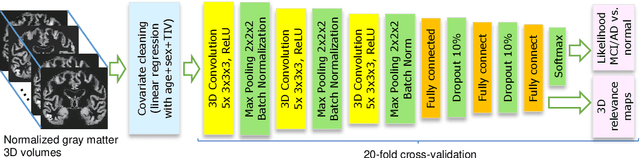
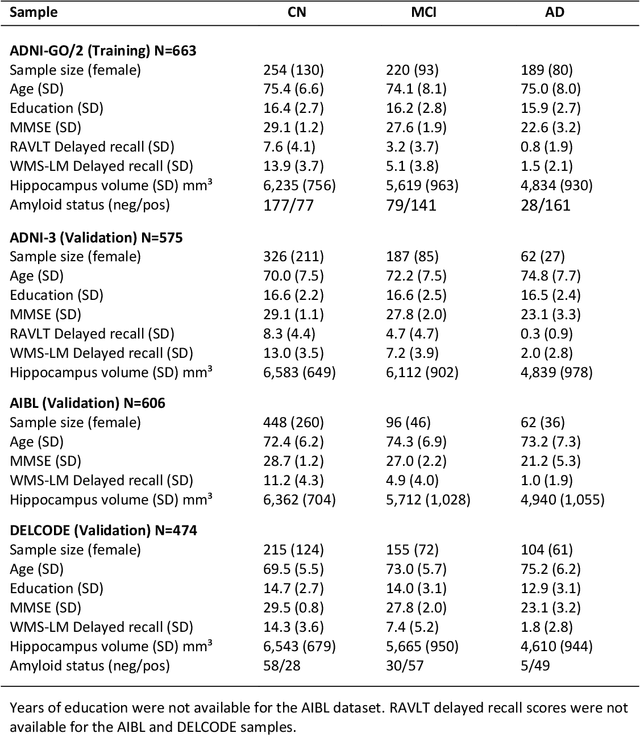
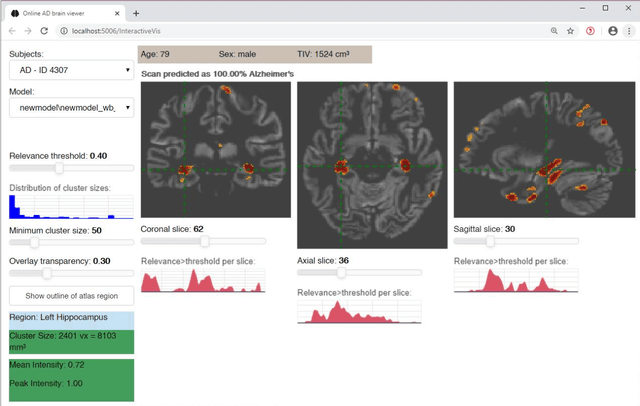
Abstract:Although convolutional neural networks (CNN) achieve high diagnostic accuracy for detecting Alzheimer's disease (AD) dementia based on magnetic resonance imaging (MRI) scans, they are not yet applied in clinical routine. One important reason for this is a lack of model comprehensibility. Recently developed visualization methods for deriving CNN relevance maps may help to fill this gap. We investigated whether models with higher accuracy also rely more on discriminative brain regions predefined by prior knowledge. We trained a CNN for the detection of AD in N=663 T1-weighted MRI scans of patients with dementia and amnestic mild cognitive impairment (MCI) and verified the accuracy of the models via cross-validation and in three independent samples including N=1655 cases. We evaluated the association of relevance scores and hippocampus volume to validate the clinical utility of this approach. To improve model comprehensibility, we implemented an interactive visualization of 3D CNN relevance maps. Across three independent datasets, group separation showed high accuracy for AD dementia vs. controls (AUC$\geq$0.92) and moderate accuracy for MCI vs. controls (AUC$\approx$0.75). Relevance maps indicated that hippocampal atrophy was considered as the most informative factor for AD detection, with additional contributions from atrophy in other cortical and subcortical regions. Relevance scores within the hippocampus were highly correlated with hippocampal volumes (Pearson's r$\approx$-0.81). The relevance maps highlighted atrophy in regions that we had hypothesized a priori. This strengthens the comprehensibility of the CNN models, which were trained in a purely data-driven manner based on the scans and diagnosis labels. The high hippocampus relevance scores and high performance achieved in independent samples support the validity of the CNN models in the detection of AD-related MRI abnormalities.
Is the winner really the best? A critical analysis of common research practice in biomedical image analysis competitions
Jun 06, 2018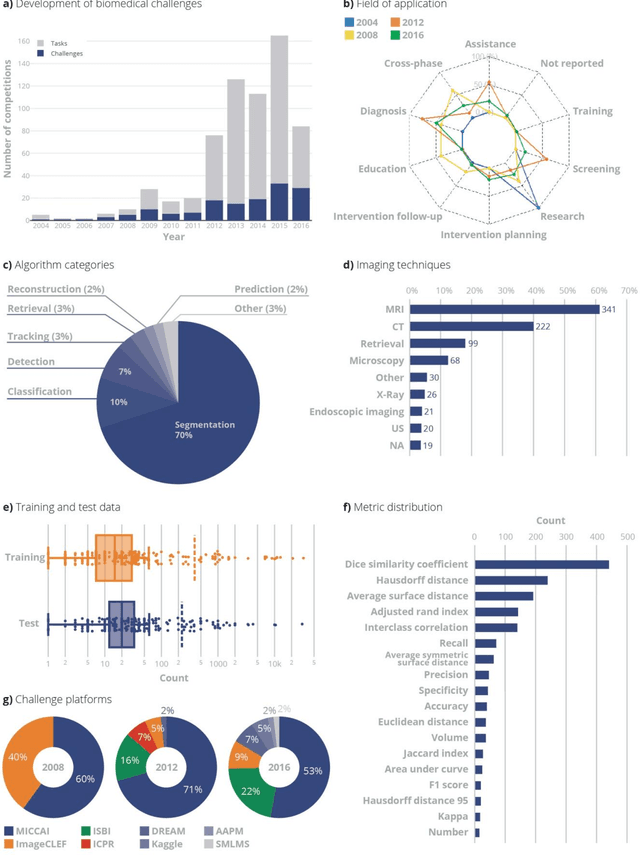
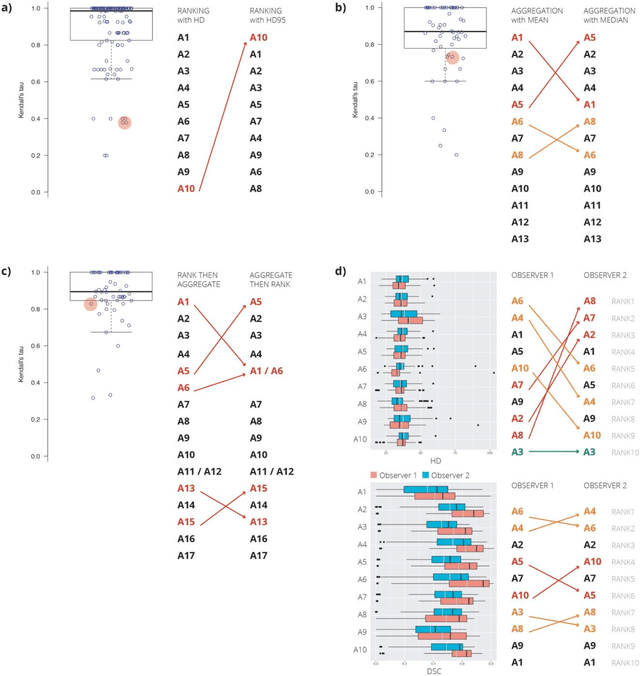
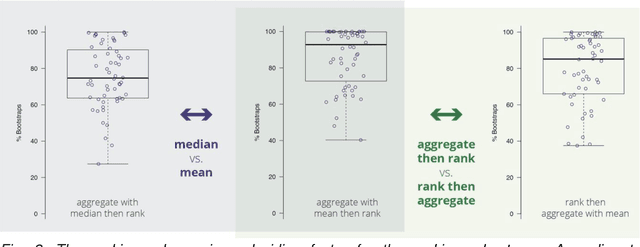
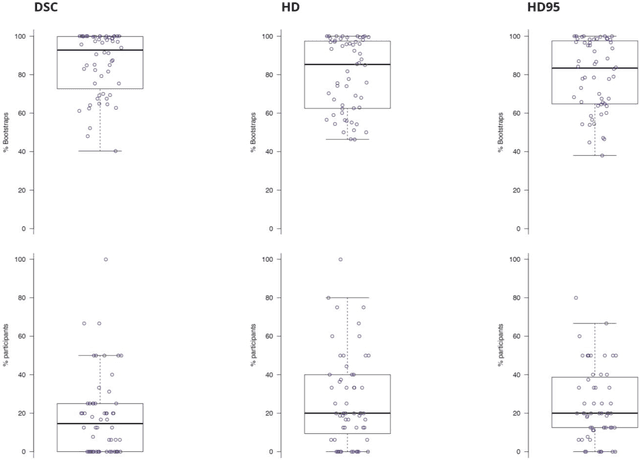
Abstract:International challenges have become the standard for validation of biomedical image analysis methods. Given their scientific impact, it is surprising that a critical analysis of common practices related to the organization of challenges has not yet been performed. In this paper, we present a comprehensive analysis of biomedical image analysis challenges conducted up to now. We demonstrate the importance of challenges and show that the lack of quality control has critical consequences. First, reproducibility and interpretation of the results is often hampered as only a fraction of relevant information is typically provided. Second, the rank of an algorithm is generally not robust to a number of variables such as the test data used for validation, the ranking scheme applied and the observers that make the reference annotations. To overcome these problems, we recommend best practice guidelines and define open research questions to be addressed in the future.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge