Mallikarjun Shankar
Scalable Artificial Intelligence for Science: Perspectives, Methods and Exemplars
Jun 24, 2024



Abstract:In a post-ChatGPT world, this paper explores the potential of leveraging scalable artificial intelligence for scientific discovery. We propose that scaling up artificial intelligence on high-performance computing platforms is essential to address such complex problems. This perspective focuses on scientific use cases like cognitive simulations, large language models for scientific inquiry, medical image analysis, and physics-informed approaches. The study outlines the methodologies needed to address such challenges at scale on supercomputers or the cloud and provides exemplars of such approaches applied to solve a variety of scientific problems.
Zero Coordinate Shift: Whetted Automatic Differentiation for Physics-informed Operator Learning
Nov 01, 2023



Abstract:Automatic differentiation (AD) is a critical step in physics-informed machine learning, required for computing the high-order derivatives of network output w.r.t. coordinates. In this paper, we present a novel and lightweight algorithm to conduct such AD for physics-informed operator learning, as we call the trick of Zero Coordinate Shift (ZCS). Instead of making all sampled coordinates leaf variables, ZCS introduces only one scalar-valued leaf variable for each spatial or temporal dimension, leading to a game-changing performance leap by simplifying the wanted derivatives from "many-roots-many-leaves" to "one-root-many-leaves". ZCS is easy to implement with current deep learning libraries; our own implementation is by extending the DeepXDE package. We carry out a comprehensive benchmark analysis and several case studies, training physics-informed DeepONets to solve partial differential equations (PDEs) without data. The results show that ZCS has persistently brought down GPU memory consumption and wall time for training by an order of magnitude, with the savings increasing with problem scale (i.e., number of functions, number of points and order of PDE). As a low-level optimisation, ZCS entails no restrictions on data, physics (PDEs) or network architecture and does not compromise training results from any aspect.
DeepSpeed4Science Initiative: Enabling Large-Scale Scientific Discovery through Sophisticated AI System Technologies
Oct 11, 2023



Abstract:In the upcoming decade, deep learning may revolutionize the natural sciences, enhancing our capacity to model and predict natural occurrences. This could herald a new era of scientific exploration, bringing significant advancements across sectors from drug development to renewable energy. To answer this call, we present DeepSpeed4Science initiative (deepspeed4science.ai) which aims to build unique capabilities through AI system technology innovations to help domain experts to unlock today's biggest science mysteries. By leveraging DeepSpeed's current technology pillars (training, inference and compression) as base technology enablers, DeepSpeed4Science will create a new set of AI system technologies tailored for accelerating scientific discoveries by addressing their unique complexity beyond the common technical approaches used for accelerating generic large language models (LLMs). In this paper, we showcase the early progress we made with DeepSpeed4Science in addressing two of the critical system challenges in structural biology research.
MLPerf HPC: A Holistic Benchmark Suite for Scientific Machine Learning on HPC Systems
Oct 26, 2021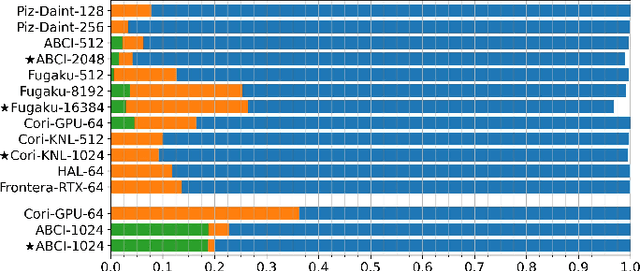
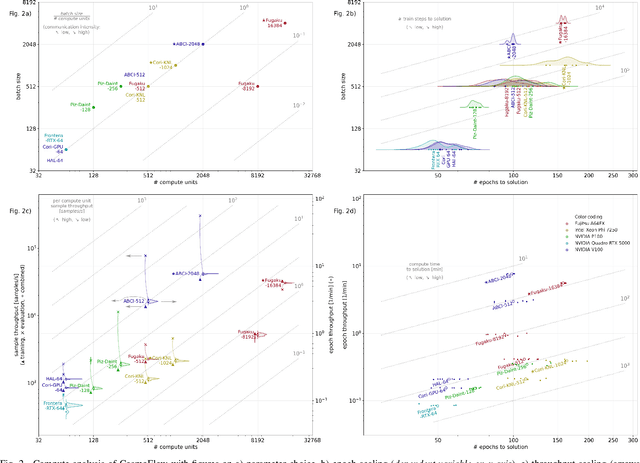
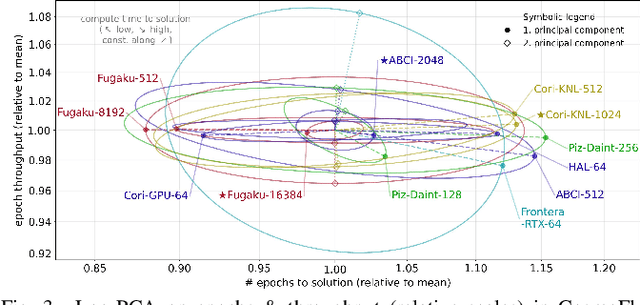
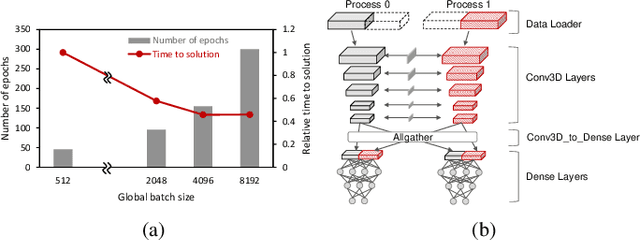
Abstract:Scientific communities are increasingly adopting machine learning and deep learning models in their applications to accelerate scientific insights. High performance computing systems are pushing the frontiers of performance with a rich diversity of hardware resources and massive scale-out capabilities. There is a critical need to understand fair and effective benchmarking of machine learning applications that are representative of real-world scientific use cases. MLPerf is a community-driven standard to benchmark machine learning workloads, focusing on end-to-end performance metrics. In this paper, we introduce MLPerf HPC, a benchmark suite of large-scale scientific machine learning training applications driven by the MLCommons Association. We present the results from the first submission round, including a diverse set of some of the world's largest HPC systems. We develop a systematic framework for their joint analysis and compare them in terms of data staging, algorithmic convergence, and compute performance. As a result, we gain a quantitative understanding of optimizations on different subsystems such as staging and on-node loading of data, compute-unit utilization, and communication scheduling, enabling overall $>10 \times$ (end-to-end) performance improvements through system scaling. Notably, our analysis shows a scale-dependent interplay between the dataset size, a system's memory hierarchy, and training convergence that underlines the importance of near-compute storage. To overcome the data-parallel scalability challenge at large batch sizes, we discuss specific learning techniques and hybrid data-and-model parallelism that are effective on large systems. We conclude by characterizing each benchmark with respect to low-level memory, I/O, and network behavior to parameterize extended roofline performance models in future rounds.
Scientific Machine Learning Benchmarks
Oct 25, 2021
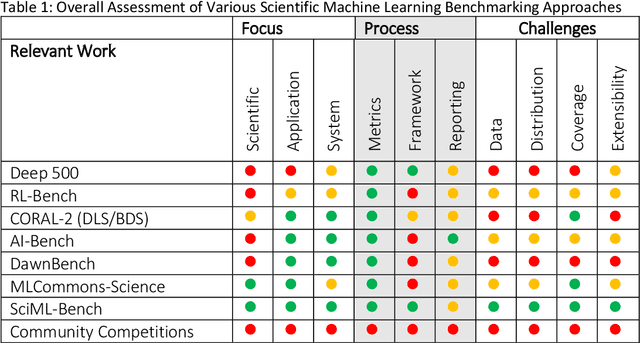
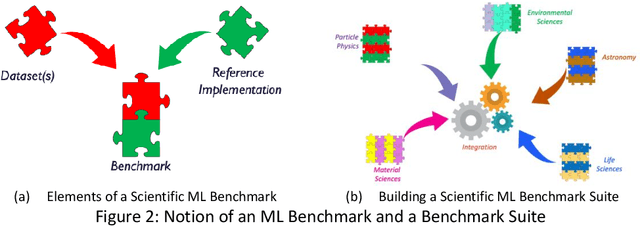
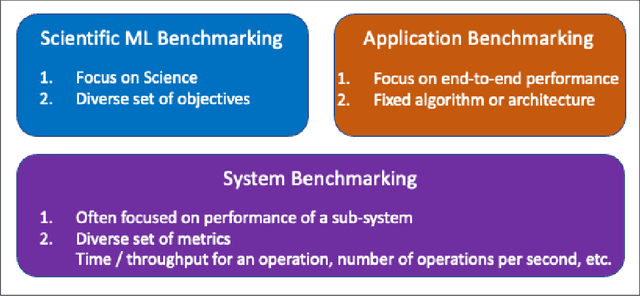
Abstract:The breakthrough in Deep Learning neural networks has transformed the use of AI and machine learning technologies for the analysis of very large experimental datasets. These datasets are typically generated by large-scale experimental facilities at national laboratories. In the context of science, scientific machine learning focuses on training machines to identify patterns, trends, and anomalies to extract meaningful scientific insights from such datasets. With a new generation of experimental facilities, the rate of data generation and the scale of data volumes will increasingly require the use of more automated data analysis. At present, identifying the most appropriate machine learning algorithm for the analysis of any given scientific dataset is still a challenge for scientists. This is due to many different machine learning frameworks, computer architectures, and machine learning models. Historically, for modelling and simulation on HPC systems such problems have been addressed through benchmarking computer applications, algorithms, and architectures. Extending such a benchmarking approach and identifying metrics for the application of machine learning methods to scientific datasets is a new challenge for both scientists and computer scientists. In this paper, we describe our approach to the development of scientific machine learning benchmarks and review other approaches to benchmarking scientific machine learning.
Data optimization for large batch distributed training of deep neural networks
Dec 18, 2020
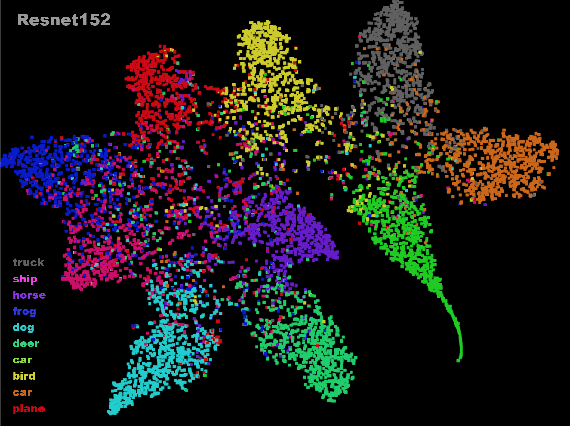

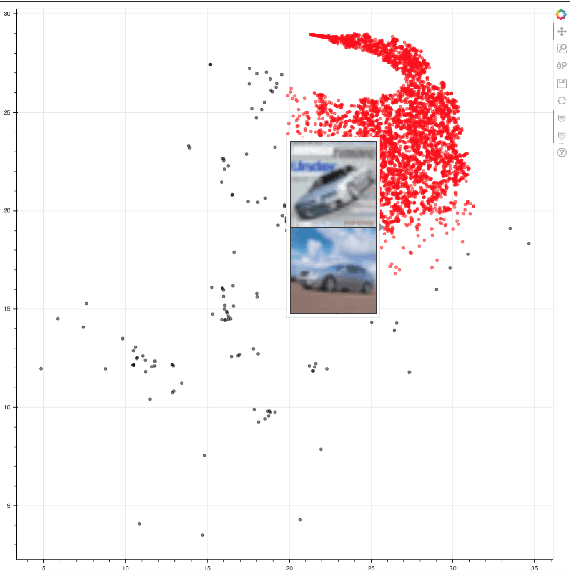
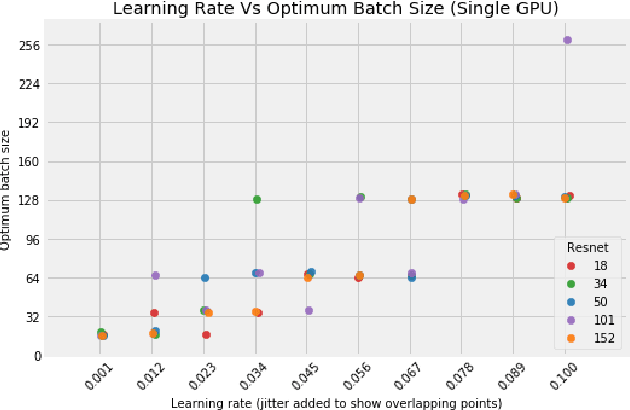
Abstract:Distributed training in deep learning (DL) is common practice as data and models grow. The current practice for distributed training of deep neural networks faces the challenges of communication bottlenecks when operating at scale, and model accuracy deterioration with an increase in global batch size. Present solutions focus on improving message exchange efficiency as well as implementing techniques to tweak batch sizes and models in the training process. The loss of training accuracy typically happens because the loss function gets trapped in a local minima. We observe that the loss landscape minimization is shaped by both the model and training data and propose a data optimization approach that utilizes machine learning to implicitly smooth out the loss landscape resulting in fewer local minima. Our approach filters out data points which are less important to feature learning, enabling us to speed up the training of models on larger batch sizes to improved accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge