Lizhen Lin
Bayesian community detection for networks with covariates
Mar 04, 2022
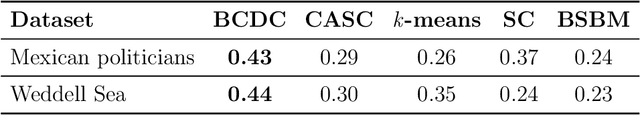
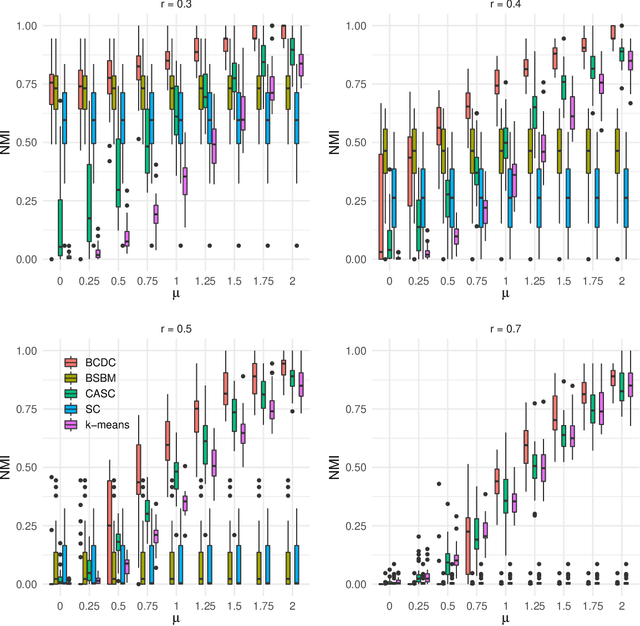
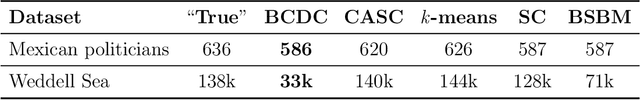
Abstract:The increasing prevalence of network data in a vast variety of fields and the need to extract useful information out of them have spurred fast developments in related models and algorithms. Among the various learning tasks with network data, community detection, the discovery of node clusters or "communities," has arguably received the most attention in the scientific community. In many real-world applications, the network data often come with additional information in the form of node or edge covariates that should ideally be leveraged for inference. In this paper, we add to a limited literature on community detection for networks with covariates by proposing a Bayesian stochastic block model with a covariate-dependent random partition prior. Under our prior, the covariates are explicitly expressed in specifying the prior distribution on the cluster membership. Our model has the flexibility of modeling uncertainties of all the parameter estimates including the community membership. Importantly, and unlike the majority of existing methods, our model has the ability to learn the number of the communities via posterior inference without having to assume it to be known. Our model can be applied to community detection in both dense and sparse networks, with both categorical and continuous covariates, and our MCMC algorithm is very efficient with good mixing properties. We demonstrate the superior performance of our model over existing models in a comprehensive simulation study and an application to two real datasets.
Robustness against Adversarial Attacks in Neural Networks using Incremental Dissipativity
Nov 25, 2021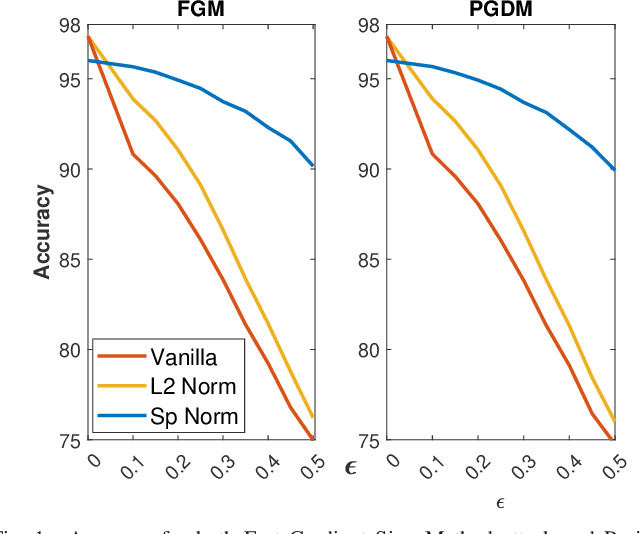
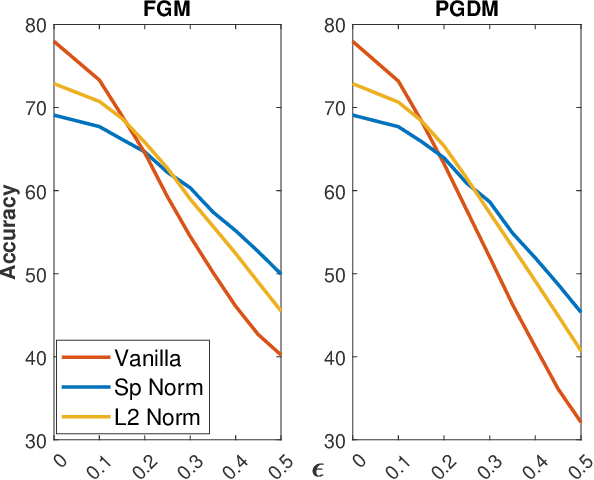
Abstract:Adversarial examples can easily degrade the classification performance in neural networks. Empirical methods for promoting robustness to such examples have been proposed, but often lack both analytical insights and formal guarantees. Recently, some robustness certificates have appeared in the literature based on system theoretic notions. This work proposes an incremental dissipativity-based robustness certificate for neural networks in the form of a linear matrix inequality for each layer. We also propose an equivalent spectral norm bound for this certificate which is scalable to neural networks with multiple layers. We demonstrate the improved performance against adversarial attacks on a feed-forward neural network trained on MNIST and an Alexnet trained using CIFAR-10.
Adaptive variational Bayes: Optimality, computation and applications
Sep 07, 2021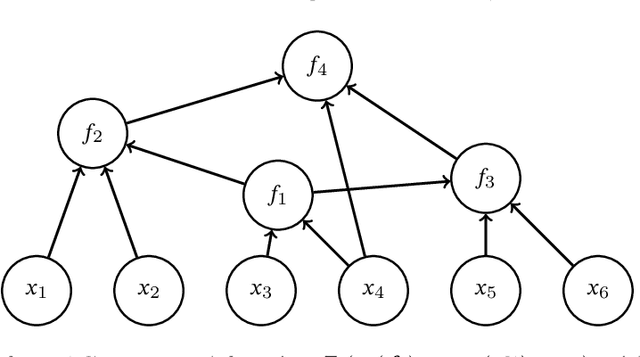
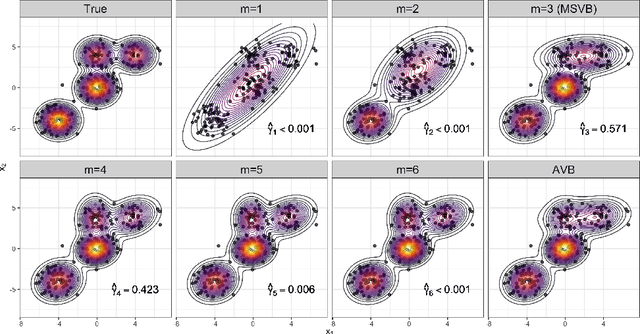
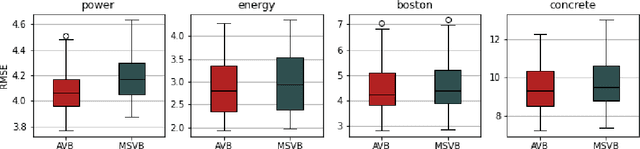
Abstract:In this paper, we explore adaptive inference based on variational Bayes. Although a number of studies have been conducted to analyze contraction properties of variational posteriors, there is still a lack of a general and computationally tractable variational Bayes method that can achieve adaptive optimal contraction of the variational posterior. We propose a novel variational Bayes framework, called adaptive variational Bayes, which can operate on a collection of models with varying dimensions and structures. The proposed framework combines variational posteriors over individual models with certain weights to obtain a variational posterior over the entire model. It turns out that this combined variational posterior minimizes the Kullback-Leibler divergence to the original posterior distribution. We show that the proposed variational posterior achieves optimal contraction rates adaptively under very general conditions and attains model selection consistency when the true model structure exists. We apply the general results obtained for the adaptive variational Bayes to several examples including deep learning models and derive some new and adaptive inference results. Moreover, we consider the use of quasi-likelihood in our framework. We formulate conditions on the quasi-likelihood to ensure the adaptive optimality and discuss specific applications to stochastic block models and nonparametric regression with sub-Gaussian errors.
Training Graph Neural Networks by Graphon Estimation
Sep 04, 2021
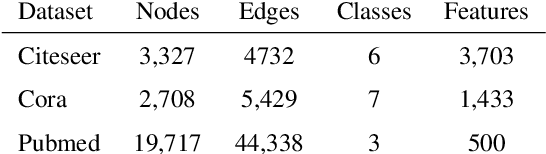
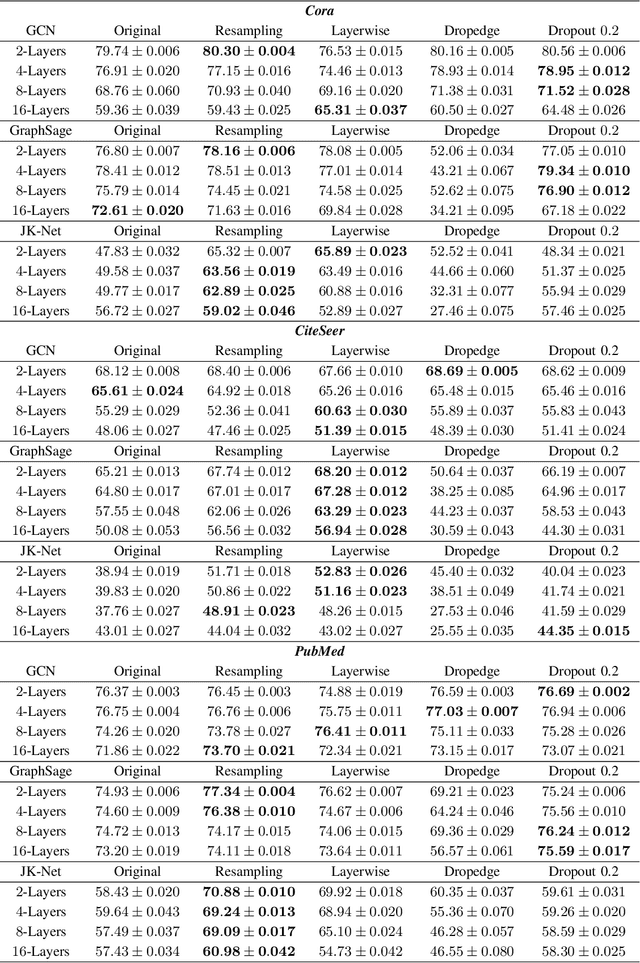
Abstract:In this work, we propose to train a graph neural network via resampling from a graphon estimate obtained from the underlying network data. More specifically, the graphon or the link probability matrix of the underlying network is first obtained from which a new network will be resampled and used during the training process at each layer. Due to the uncertainty induced from the resampling, it helps mitigate the well-known issue of over-smoothing in a graph neural network (GNN) model. Our framework is general, computationally efficient, and conceptually simple. Another appealing feature of our method is that it requires minimal additional tuning during the training process. Extensive numerical results show that our approach is competitive with and in many cases outperform the other over-smoothing reducing GNN training methods.
A likelihood approach to nonparametric estimation of a singular distribution using deep generative models
May 09, 2021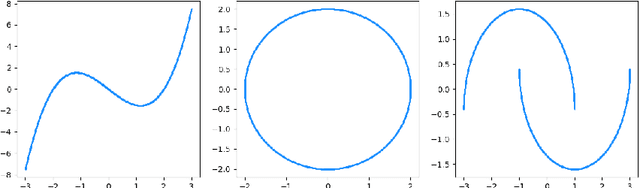
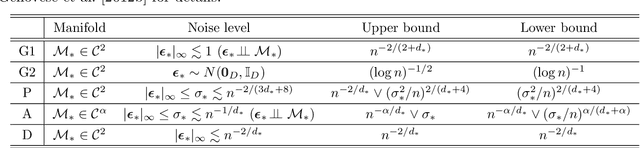

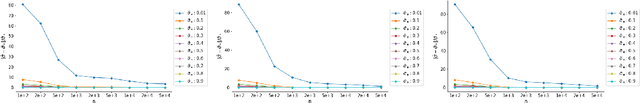
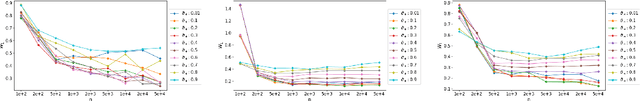
Abstract:We investigate statistical properties of a likelihood approach to nonparametric estimation of a singular distribution using deep generative models. More specifically, a deep generative model is used to model high-dimensional data that are assumed to concentrate around some low-dimensional structure. Estimating the distribution supported on this low-dimensional structure such as a low-dimensional manifold is challenging due to its singularity with respect to the Lebesgue measure in the ambient space. In the considered model, a usual likelihood approach can fail to estimate the target distribution consistently due to the singularity. We prove that a novel and effective solution exists by perturbing the data with an instance noise which leads to consistent estimation of the underlying distribution with desirable convergence rates. We also characterize the class of distributions that can be efficiently estimated via deep generative models. This class is sufficiently general to contain various structured distributions such as product distributions, classically smooth distributions and distributions supported on a low-dimensional manifold. Our analysis provides some insights on how deep generative models can avoid the curse of dimensionality for nonparametric distribution estimation. We conduct thorough simulation study and real data analysis to empirically demonstrate that the proposed data perturbation technique improves the estimation performance significantly.
Accelerated Algorithms for Convex and Non-Convex Optimization on Manifolds
Oct 18, 2020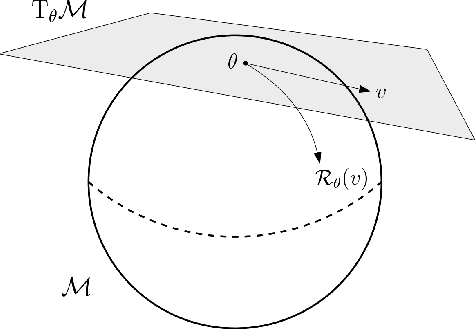
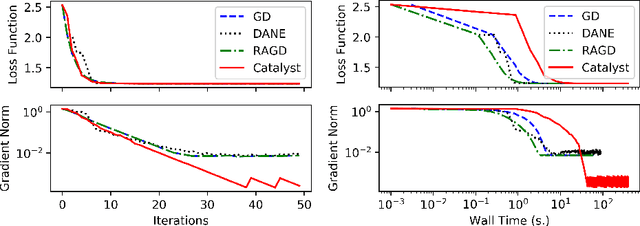
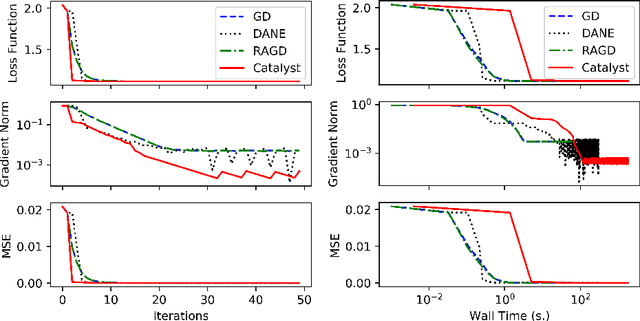
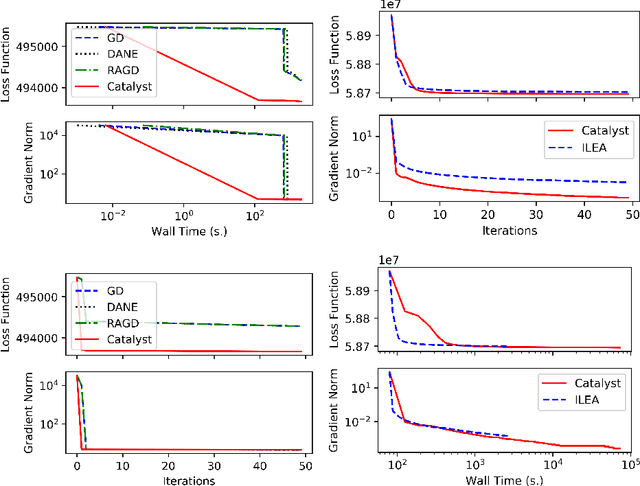
Abstract:We propose a general scheme for solving convex and non-convex optimization problems on manifolds. The central idea is that, by adding a multiple of the squared retraction distance to the objective function in question, we "convexify" the objective function and solve a series of convex sub-problems in the optimization procedure. One of the key challenges for optimization on manifolds is the difficulty of verifying the complexity of the objective function, e.g., whether the objective function is convex or non-convex, and the degree of non-convexity. Our proposed algorithm adapts to the level of complexity in the objective function. We show that when the objective function is convex, the algorithm provably converges to the optimum and leads to accelerated convergence. When the objective function is non-convex, the algorithm will converge to a stationary point. Our proposed method unifies insights from Nesterov's original idea for accelerating gradient descent algorithms with recent developments in optimization algorithms in Euclidean space. We demonstrate the utility of our algorithms on several manifold optimization tasks such as estimating intrinsic and extrinsic Fr\'echet means on spheres and low-rank matrix factorization with Grassmann manifolds applied to the Netflix rating data set.
Community detection, pattern recognition, and hypergraph-based learning: approaches using metric geometry and persistent homology
Sep 29, 2020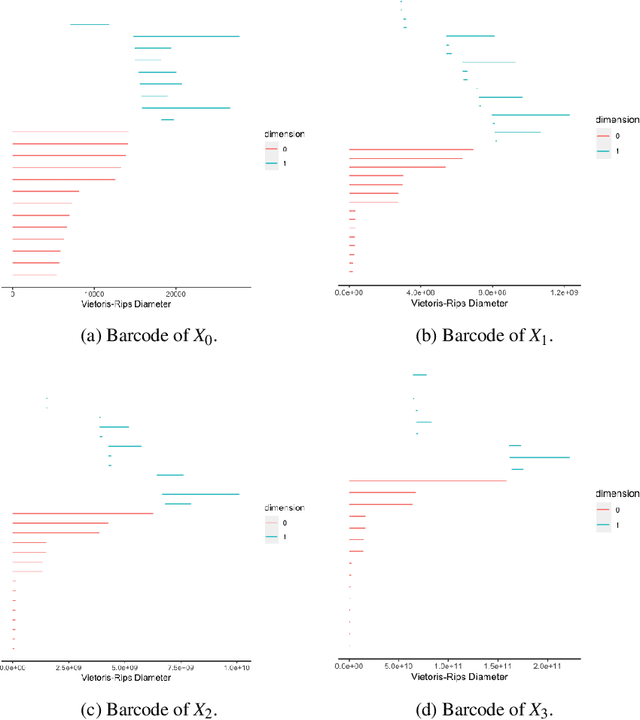
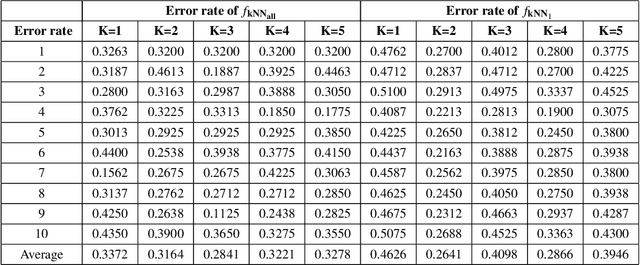
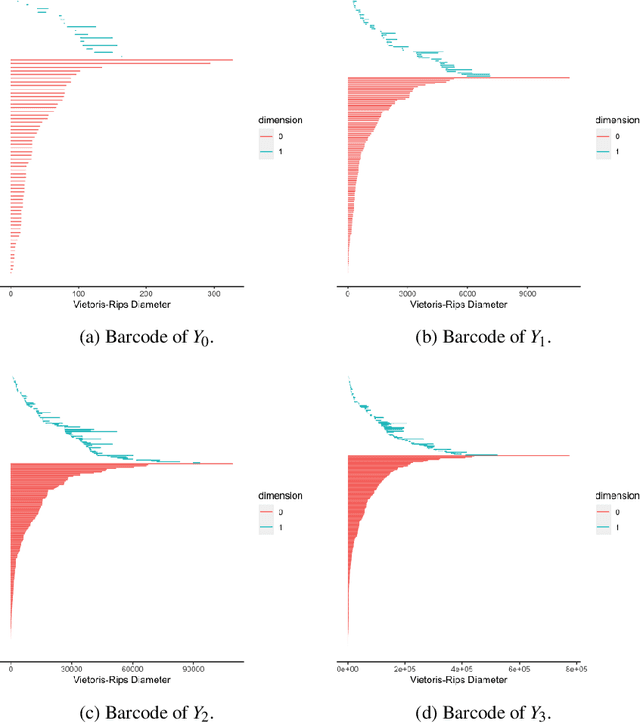
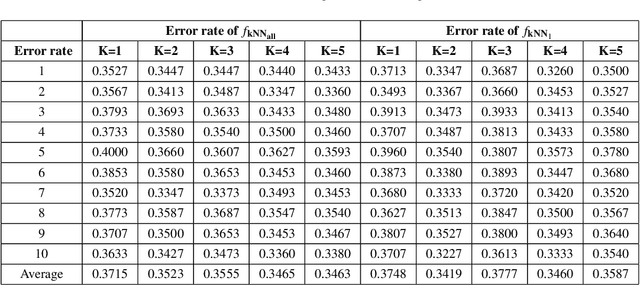
Abstract:Hypergraph data appear and are hidden in many places in the modern age. They are data structure that can be used to model many real data examples since their structures contain information about higher order relations among data points. One of the main contributions of our paper is to introduce a new topological structure to hypergraph data which bears a resemblance to a usual metric space structure. Using this new topological space structure of hypergraph data, we propose several approaches to study community detection problem, detecting persistent features arising from homological structure of hypergraph data. Also based on the topological space structure of hypergraph data introduced in our paper, we introduce a modified nearest neighbors methods which is a generalization of the classical nearest neighbors methods from machine learning. Our modified nearest neighbors methods have an advantage of being very flexible and applicable even for discrete structures as in hypergraphs. We then apply our modified nearest neighbors methods to study sign prediction problem in hypegraph data constructed using our method.
Weight Prediction for Variants of Weighted Directed Networks
Sep 29, 2020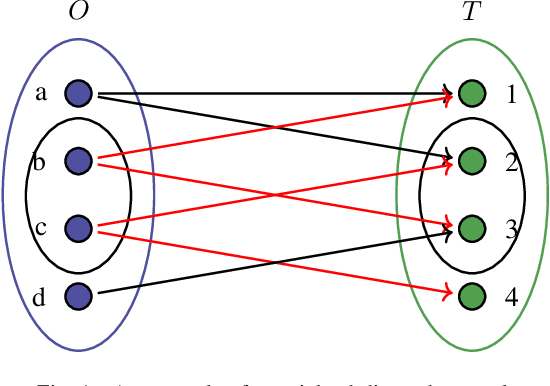

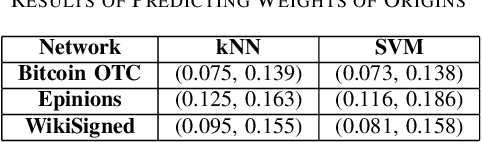
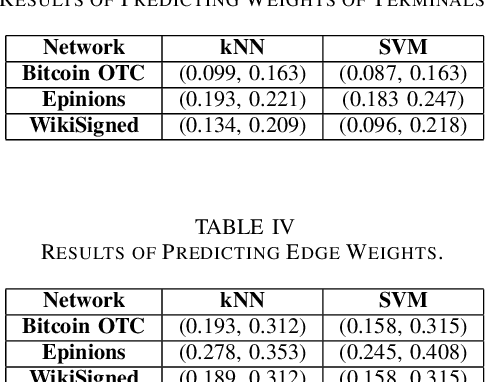
Abstract:A weighted directed network (WDN) is a directed graph in which each edge is associated to a unique value called weight. These networks are very suitable for modeling real-world social networks in which there is an assessment of one vertex toward other vertices. One of the main problems studied in this paper is prediction of edge weights in such networks. We introduce, for the first time, a metric geometry approach to studying edge weight prediction in WDNs. We modify a usual notion of WDNs, and introduce a new type of WDNs which we coin the term \textit{almost-weighted directed networks} (AWDNs). AWDNs can capture the weight information of a network from a given training set. We then construct a class of metrics (or distances) for AWDNs which equips such networks with a metric space structure. Using the metric geometry structure of AWDNs, we propose modified $k$ nearest neighbors (kNN) methods and modified support-vector machine (SVM) methods which will then be used to predict edge weights in AWDNs. In many real-world datasets, in addition to edge weights, one can also associate weights to vertices which capture information of vertices; association of weights to vertices especially plays an important role in graph embedding problems. Adopting a similar approach, we introduce two new types of directed networks in which weights are associated to either a subset of origin vertices or a subset of terminal vertices . We, for the first time, construct novel classes of metrics on such networks, and based on these new metrics propose modified $k$NN and SVM methods for predicting weights of origins and terminals in these networks. We provide experimental results on several real-world datasets, using our geometric methodologies.
Neural Time-Dependent Partial Differential Equation
Sep 08, 2020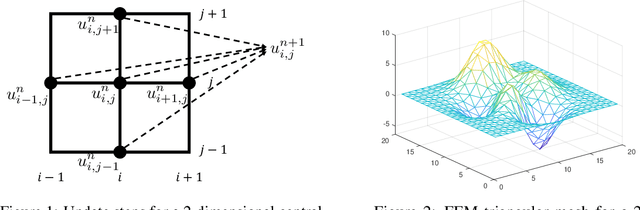
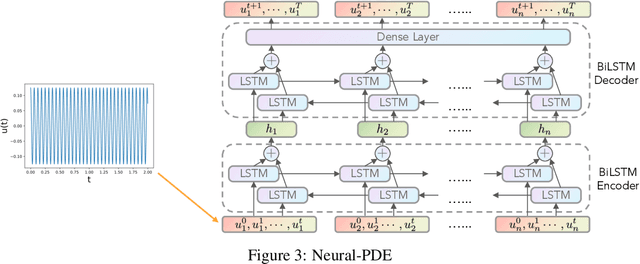
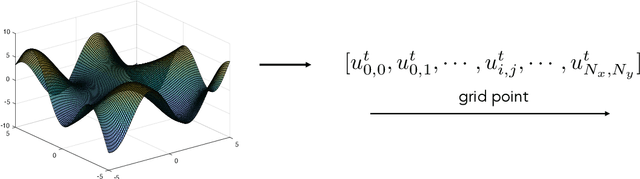
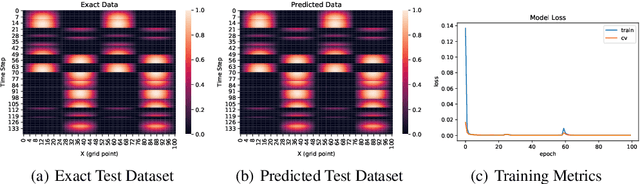
Abstract:Partial differential equations (PDEs) play a crucial role in studying a vast number of problems in science and engineering. Numerically solving nonlinear and/or high-dimensional PDEs is often a challenging task. Inspired by the traditional finite difference and finite elements methods and emerging advancements in machine learning, we propose a sequence deep learning framework called Neural-PDE, which allows to automatically learn governing rules of any time-dependent PDE system from existing data by using a bidirectional LSTM encoder, and predict the next n time steps data. One critical feature of our proposed framework is that the Neural-PDE is able to simultaneously learn and simulate the multiscale variables.We test the Neural-PDE by a range of examples from one-dimensional PDEs to a high-dimensional and nonlinear complex fluids model. The results show that the Neural-PDE is capable of learning the initial conditions, boundary conditions and differential operators without the knowledge of the specific form of a PDE system.In our experiments the Neural-PDE can efficiently extract the dynamics within 20 epochs training, and produces accurate predictions. Furthermore, unlike the traditional machine learning approaches in learning PDE such as CNN and MLP which require vast parameters for model precision, Neural-PDE shares parameters across all time steps, thus considerably reduces the computational complexity and leads to a fast learning algorithm.
Optimization of Graph Neural Networks with Natural Gradient Descent
Aug 21, 2020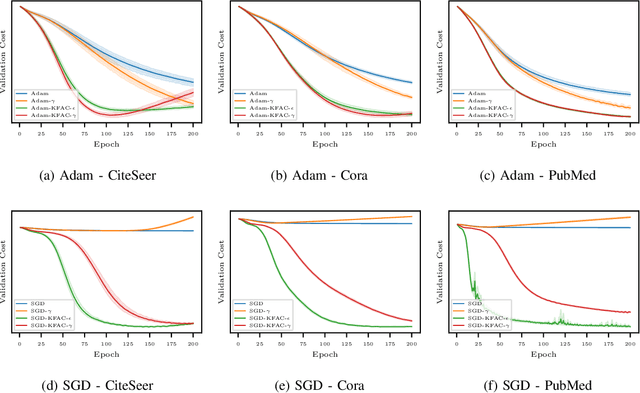
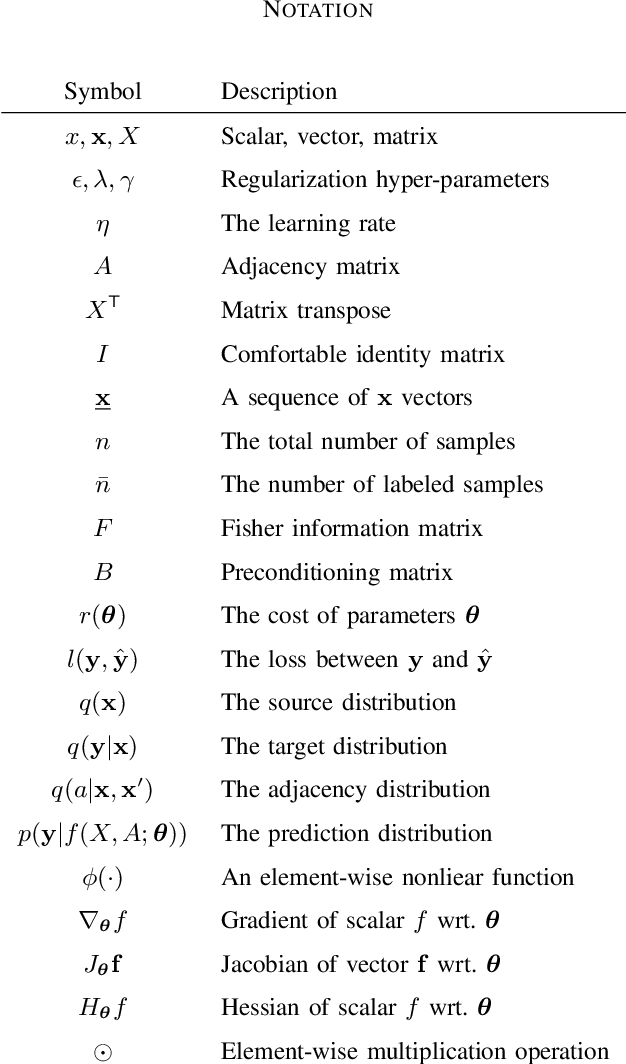
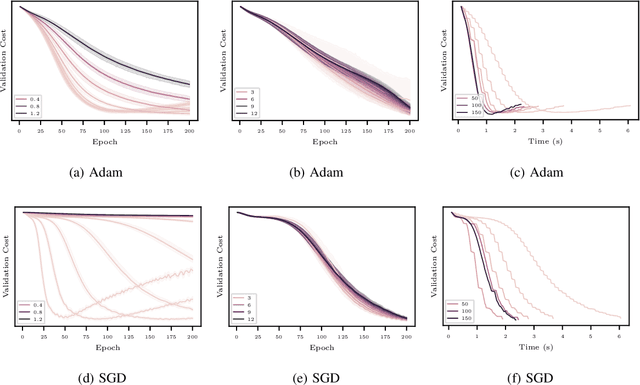
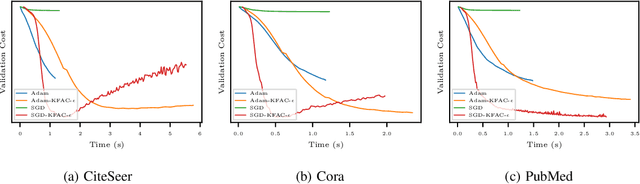

Abstract:In this work, we propose to employ information-geometric tools to optimize a graph neural network architecture such as the graph convolutional networks. More specifically, we develop optimization algorithms for the graph-based semi-supervised learning by employing the natural gradient information in the optimization process. This allows us to efficiently exploit the geometry of the underlying statistical model or parameter space for optimization and inference. To the best of our knowledge, this is the first work that has utilized the natural gradient for the optimization of graph neural networks that can be extended to other semi-supervised problems. Efficient computations algorithms are developed and extensive numerical studies are conducted to demonstrate the superior performance of our algorithms over existing algorithms such as ADAM and SGD.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge