Konrad P. Kording
Teresa
An adversarial algorithm for variational inference with a new role for acetylcholine
Jun 18, 2020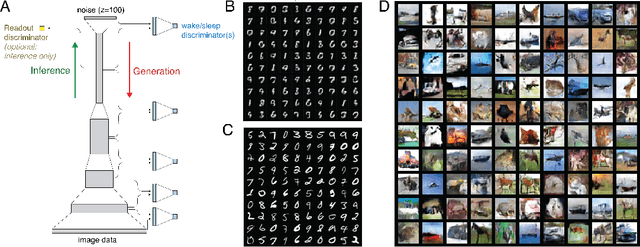
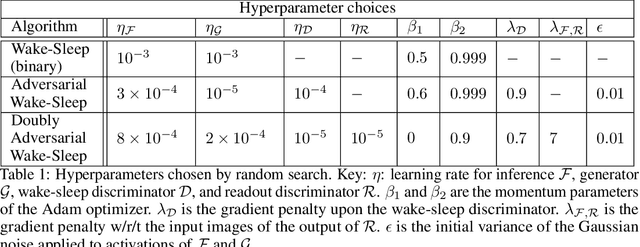

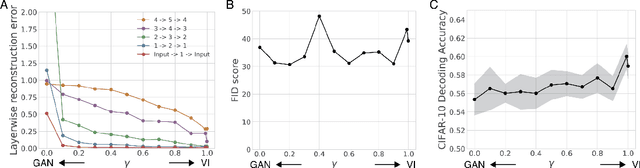

Abstract:Sensory learning in the mammalian cortex has long been hypothesized to involve the objective of variational inference (VI). Likely the most well-known algorithm for cortical VI is the Wake-Sleep algorithm (Hinton et al. 1995). However Wake-Sleep problematically assumes that neural activities are independent given lower-layers during generation. Here, we construct a VI system that is both compatible with neurobiology and avoids this assumption. The core of the system is a wake-sleep discriminator that classifies network states as inferred or self-generated. Inference connections learn by opposing this discriminator. This adversarial dynamic solves a core problem within VI, which is to match the distribution of stimulus-evoked (inference) activity to that of self-generated activity. Meanwhile, generative connections learn to predict lower-level activity as in standard VI. We implement this algorithm and show that it can successfully train the approximate inference network for generative models. Our proposed algorithm makes several biological predictions that can be tested. Most importantly, it predicts a teaching signal that is remarkably similar to known properties of the cholinergic system.
Spike-based causal inference for weight alignment
Oct 03, 2019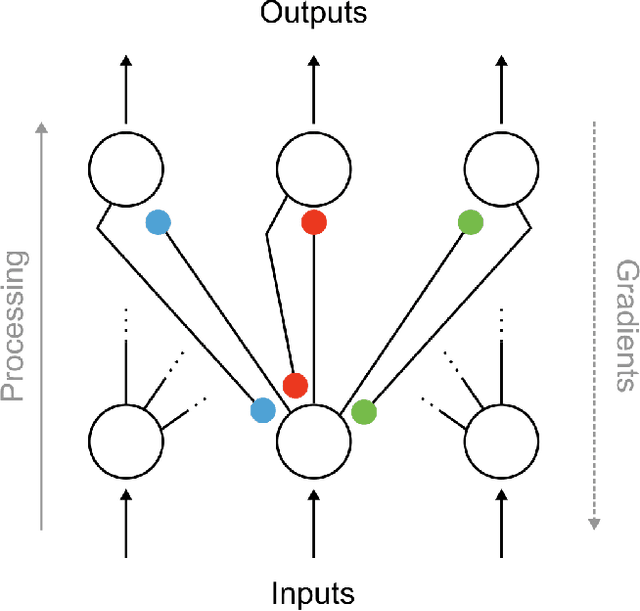
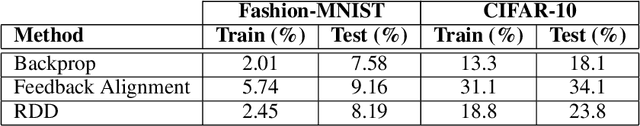
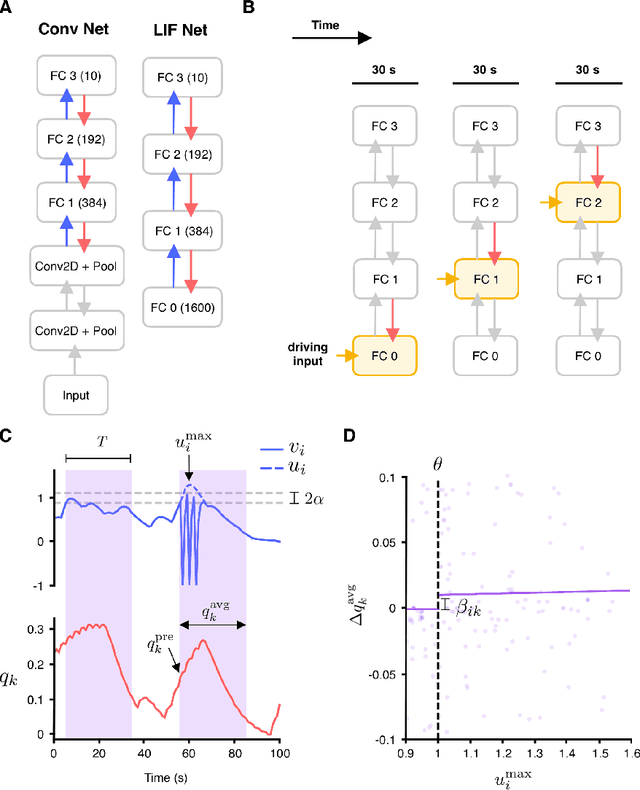
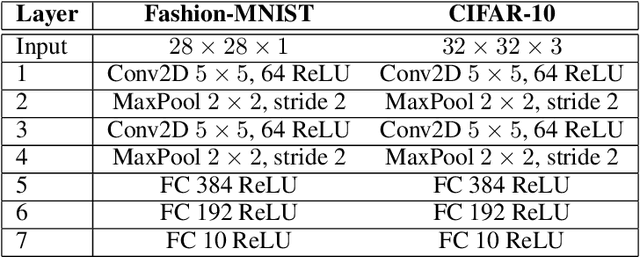
Abstract:In artificial neural networks trained with gradient descent, the weights used for processing stimuli are also used during backward passes to calculate gradients. For the real brain to approximate gradients, gradient information would have to be propagated separately, such that one set of synaptic weights is used for processing and another set is used for backward passes. This produces the so-called "weight transport problem" for biological models of learning, where the backward weights used to calculate gradients need to mirror the forward weights used to process stimuli. This weight transport problem has been considered so hard that popular proposals for biological learning assume that the backward weights are simply random, as in the feedback alignment algorithm. However, such random weights do not appear to work well for large networks. Here we show how the discontinuity introduced in a spiking system can lead to a solution to this problem. The resulting algorithm is a special case of an estimator used for causal inference in econometrics, regression discontinuity design. We show empirically that this algorithm rapidly makes the backward weights approximate the forward weights. As the backward weights become correct, this improves learning performance over feedback alignment on tasks such as Fashion-MNIST and CIFAR-10. Our results demonstrate that a simple learning rule in a spiking network can allow neurons to produce the right backward connections and thus solve the weight transport problem.
Identifying Weights and Architectures of Unknown ReLU Networks
Oct 02, 2019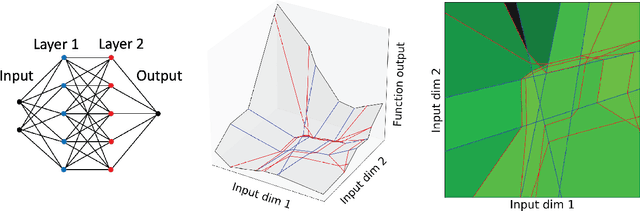
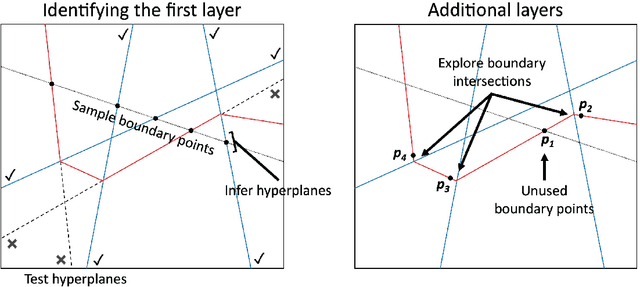
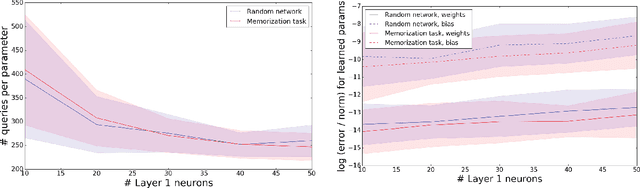
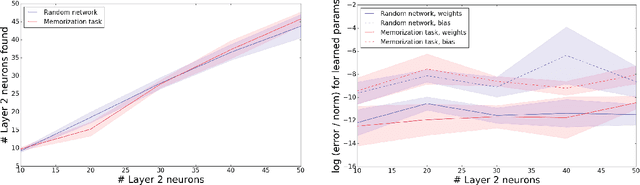
Abstract:The output of a neural network depends on its parameters in a highly nonlinear way, and it is widely assumed that a network's parameters cannot be identified from its outputs. Here, we show that in many cases it is possible to reconstruct the architecture, weights, and biases of a deep ReLU network given the ability to query the network. ReLU networks are piecewise linear and the boundaries between pieces correspond to inputs for which one of the ReLUs switches between inactive and active states. Thus, first-layer ReLUs can be identified (up to sign and scaling) based on the orientation of their associated hyperplanes. Later-layer ReLU boundaries bend when they cross earlier-layer boundaries and the extent of bending reveals the weights between them. Our algorithm uses this to identify the units in the network and weights connecting them (up to isomorphism). The fact that considerable parts of deep networks can be identified from their outputs has implications for security, neuroscience, and our understanding of neural networks.
Movement science needs different pose tracking algorithms
Jul 24, 2019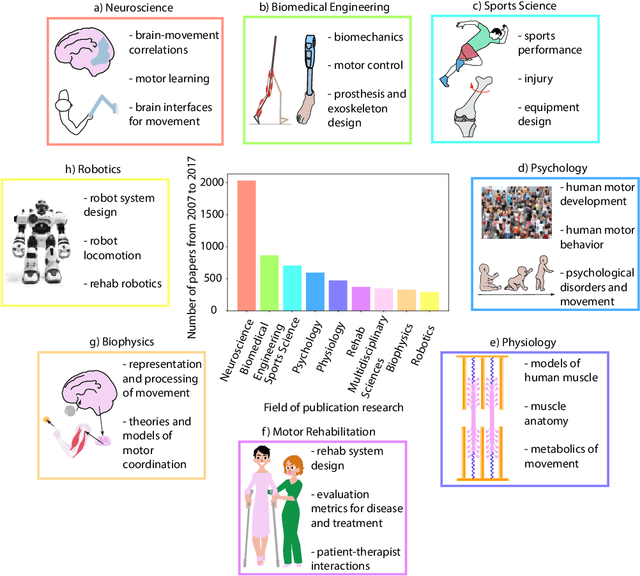
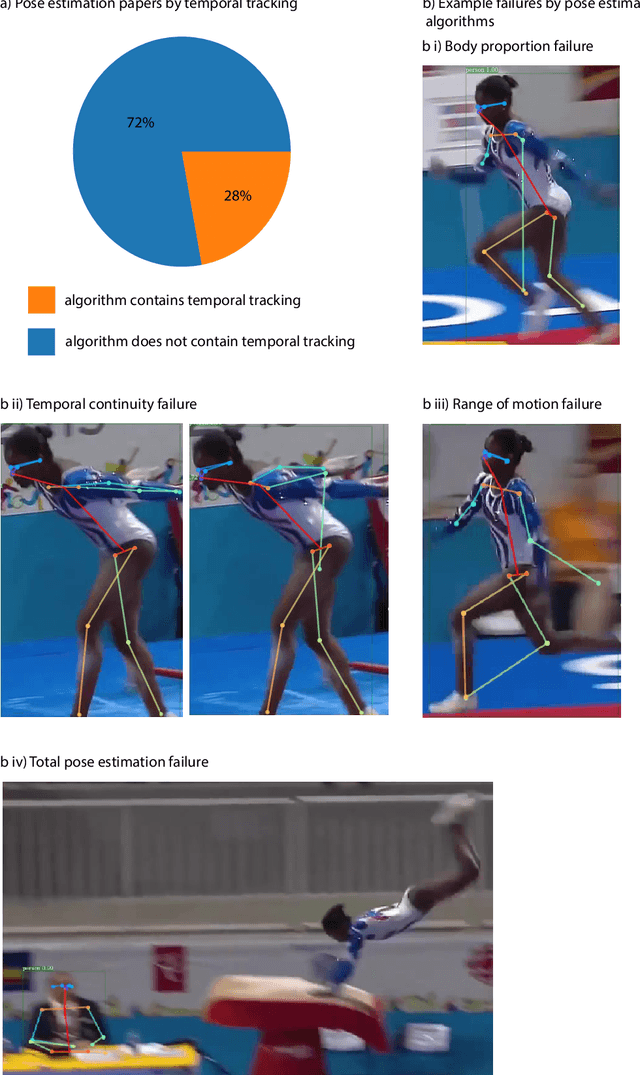

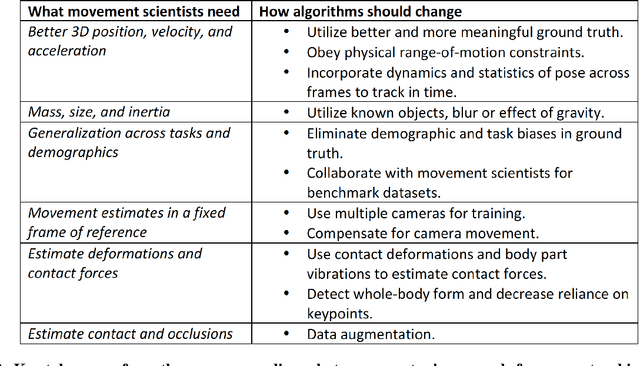
Abstract:Over the last decade, computer science has made progress towards extracting body pose from single camera photographs or videos. This promises to enable movement science to detect disease, quantify movement performance, and take the science out of the lab into the real world. However, current pose tracking algorithms fall short of the needs of movement science; the types of movement data that matter are poorly estimated. For instance, the metrics currently used for evaluating pose tracking algorithms use noisy hand-labeled ground truth data and do not prioritize precision of relevant variables like three-dimensional position, velocity, acceleration, and forces which are crucial for movement science. Here, we introduce the scientific disciplines that use movement data, the types of data they need, and discuss the changes needed to make pose tracking truly transformative for movement science.
What does it mean to understand a neural network?
Jul 15, 2019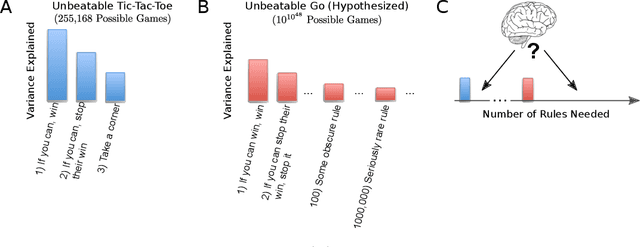
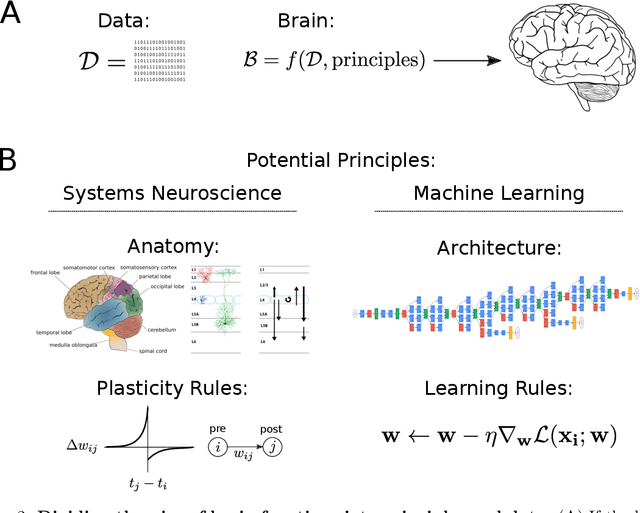
Abstract:We can define a neural network that can learn to recognize objects in less than 100 lines of code. However, after training, it is characterized by millions of weights that contain the knowledge about many object types across visual scenes. Such networks are thus dramatically easier to understand in terms of the code that makes them than the resulting properties, such as tuning or connections. In analogy, we conjecture that rules for development and learning in brains may be far easier to understand than their resulting properties. The analogy suggests that neuroscience would benefit from a focus on learning and development.
Tackling Climate Change with Machine Learning
Jun 10, 2019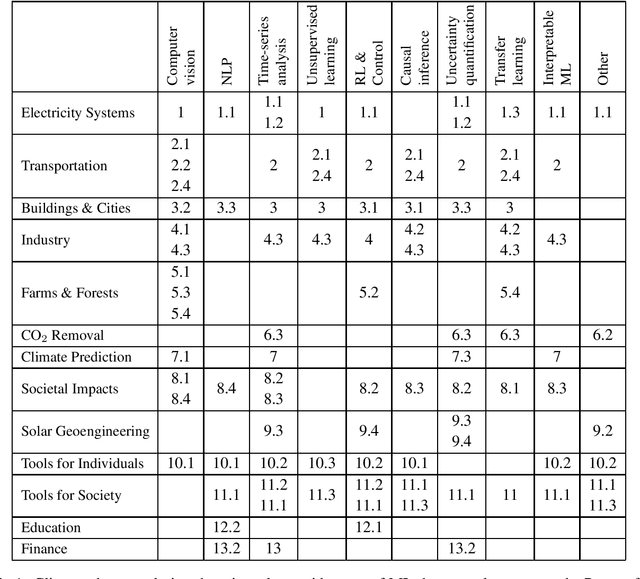
Abstract:Climate change is one of the greatest challenges facing humanity, and we, as machine learning experts, may wonder how we can help. Here we describe how machine learning can be a powerful tool in reducing greenhouse gas emissions and helping society adapt to a changing climate. From smart grids to disaster management, we identify high impact problems where existing gaps can be filled by machine learning, in collaboration with other fields. Our recommendations encompass exciting research questions as well as promising business opportunities. We call on the machine learning community to join the global effort against climate change.
The Roles of Supervised Machine Learning in Systems Neuroscience
May 21, 2018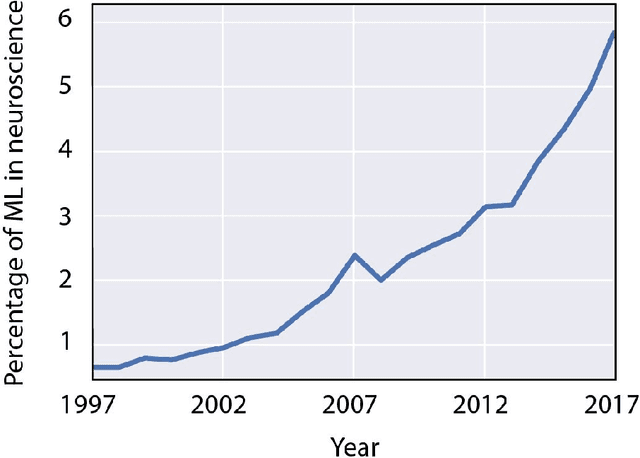
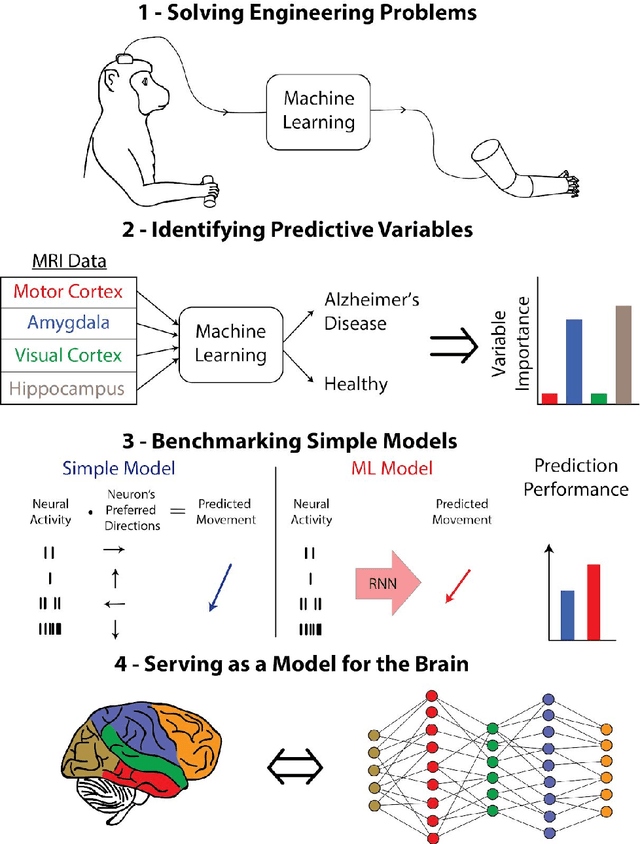
Abstract:Over the last several years, the use of machine learning (ML) in neuroscience has been increasing exponentially. Here, we review ML's contributions, both realized and potential, across several areas of systems neuroscience. We describe four primary roles of ML within neuroscience: 1) creating solutions to engineering problems, 2) identifying predictive variables, 3) setting benchmarks for simple models of the brain, and 4) serving itself as a model for the brain. The breadth and ease of its applicability suggests that machine learning should be in the toolbox of most systems neuroscientists.
Machine learning for neural decoding
May 04, 2018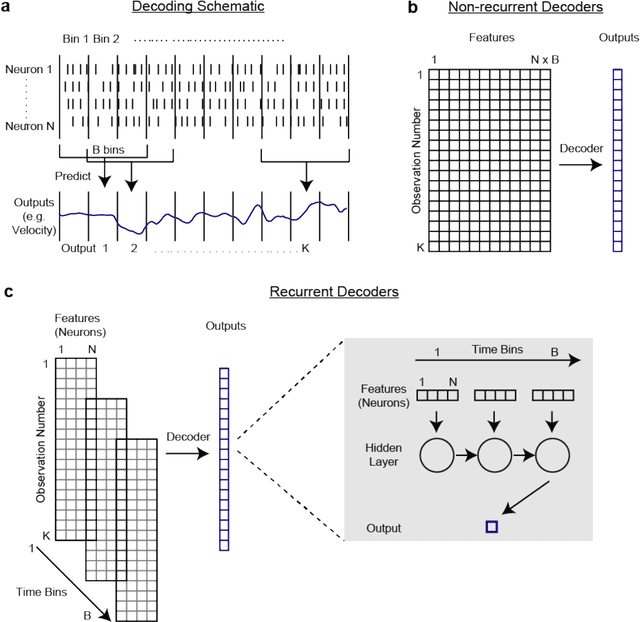

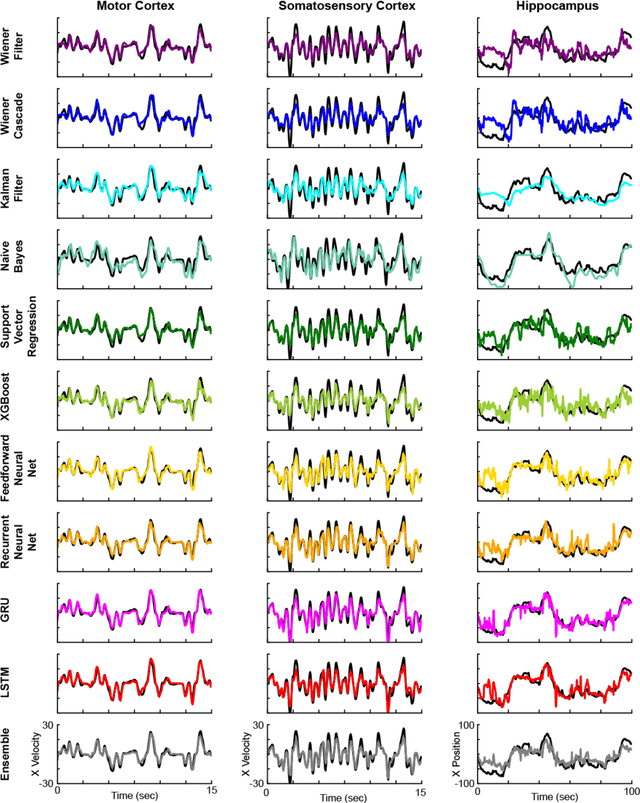
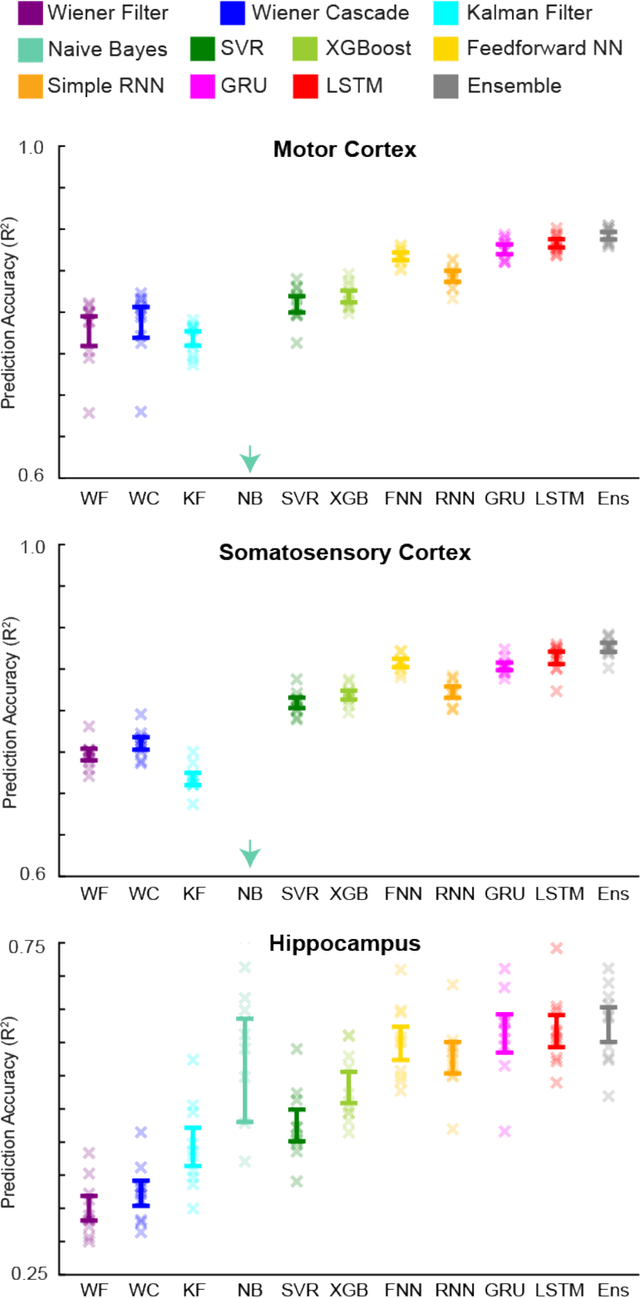
Abstract:Despite rapid advances in machine learning tools, the majority of neural decoding approaches still use traditional methods. Improving the performance of neural decoding algorithms allows us to better understand the information contained in a neural population, and can help advance engineering applications such as brain machine interfaces. Here, we apply modern machine learning techniques, including neural networks and gradient boosting, to decode from spiking activity in 1) motor cortex, 2) somatosensory cortex, and 3) hippocampus. We compare the predictive ability of these modern methods with traditional decoding methods such as Wiener and Kalman filters. Modern methods, in particular neural networks and ensembles, significantly outperformed the traditional approaches. For instance, for all of the three brain areas, an LSTM decoder explained over 40% of the unexplained variance from a Wiener filter. These results suggest that modern machine learning techniques should become the standard methodology for neural decoding. We provide a tutorial and code to facilitate wider implementation of these methods.
Puzzle Imaging: Using Large-scale Dimensionality Reduction Algorithms for Localization
Jun 21, 2015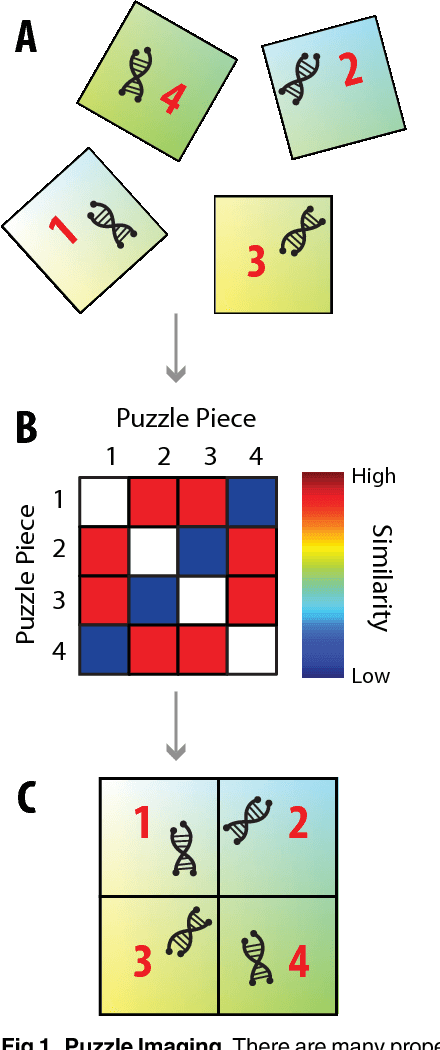
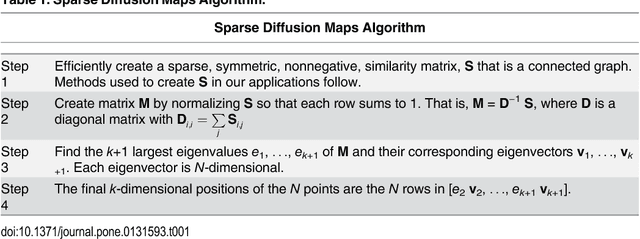

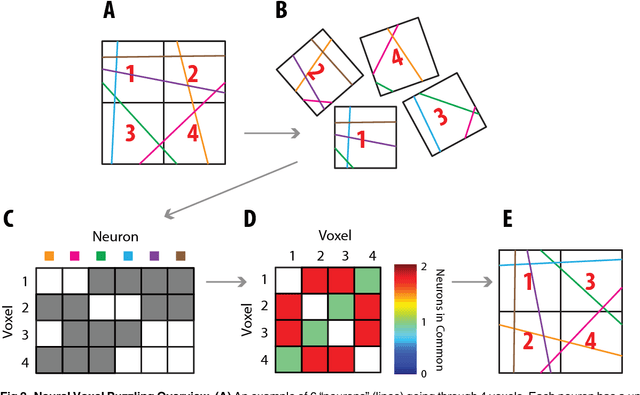
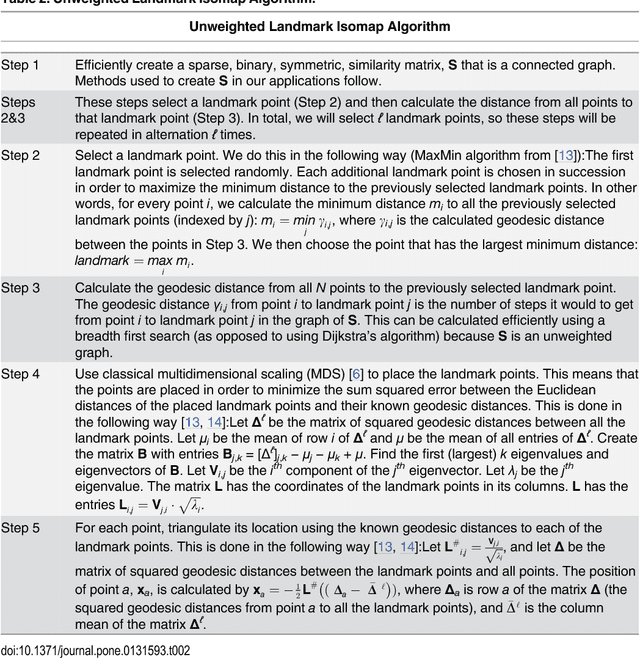
Abstract:Current high-resolution imaging techniques require an intact sample that preserves spatial relationships. We here present a novel approach, "puzzle imaging," that allows imaging a spatially scrambled sample. This technique takes many spatially disordered samples, and then pieces them back together using local properties embedded within the sample. We show that puzzle imaging can efficiently produce high-resolution images using dimensionality reduction algorithms. We demonstrate the theoretical capabilities of puzzle imaging in three biological scenarios, showing that (1) relatively precise 3-dimensional brain imaging is possible; (2) the physical structure of a neural network can often be recovered based only on the neural connectivity matrix; and (3) a chemical map could be reproduced using bacteria with chemosensitive DNA and conjugative transfer. The ability to reconstruct scrambled images promises to enable imaging based on DNA sequencing of homogenized tissue samples.
Self-Expressive Decompositions for Matrix Approximation and Clustering
May 04, 2015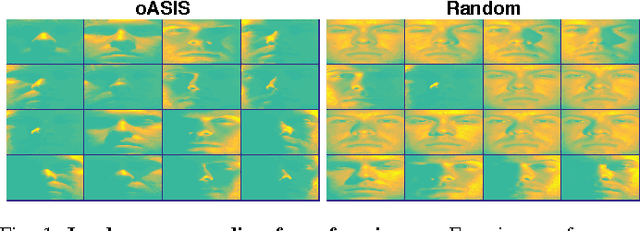
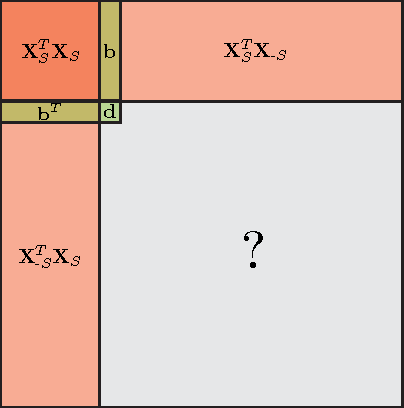
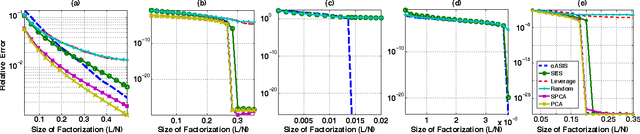
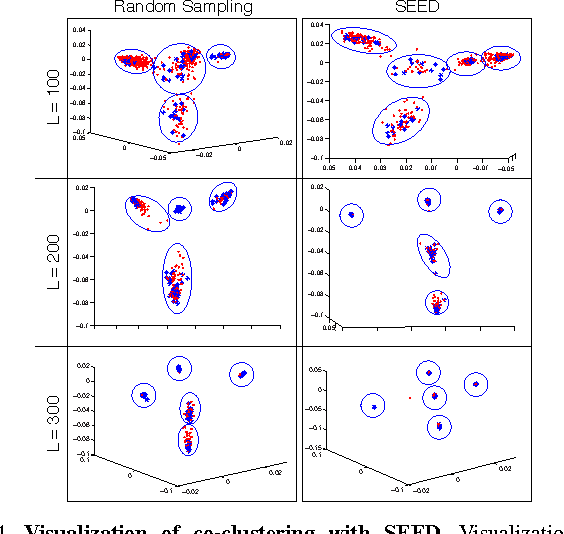
Abstract:Data-aware methods for dimensionality reduction and matrix decomposition aim to find low-dimensional structure in a collection of data. Classical approaches discover such structure by learning a basis that can efficiently express the collection. Recently, "self expression", the idea of using a small subset of data vectors to represent the full collection, has been developed as an alternative to learning. Here, we introduce a scalable method for computing sparse SElf-Expressive Decompositions (SEED). SEED is a greedy method that constructs a basis by sequentially selecting incoherent vectors from the dataset. After forming a basis from a subset of vectors in the dataset, SEED then computes a sparse representation of the dataset with respect to this basis. We develop sufficient conditions under which SEED exactly represents low rank matrices and vectors sampled from a unions of independent subspaces. We show how SEED can be used in applications ranging from matrix approximation and denoising to clustering, and apply it to numerous real-world datasets. Our results demonstrate that SEED is an attractive low-complexity alternative to other sparse matrix factorization approaches such as sparse PCA and self-expressive methods for clustering.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge