Kensaku Mori
Depth Estimation from Single-shot Monocular Endoscope Image Using Image Domain Adaptation And Edge-Aware Depth Estimation
Jan 12, 2022
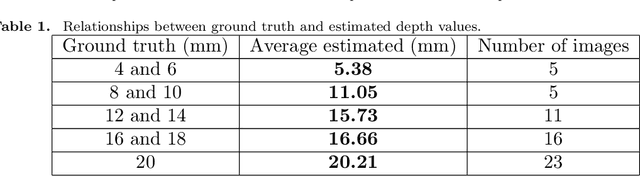
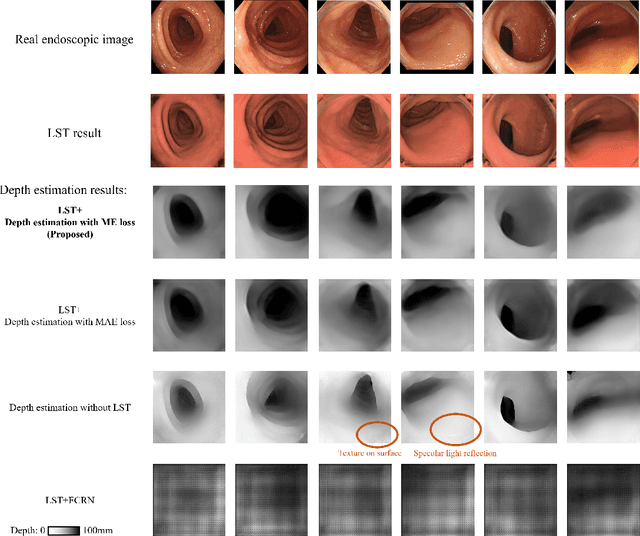
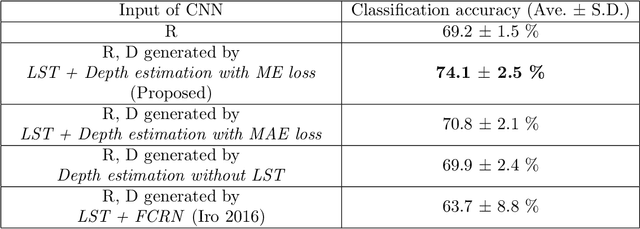
Abstract:We propose a depth estimation method from a single-shot monocular endoscopic image using Lambertian surface translation by domain adaptation and depth estimation using multi-scale edge loss. We employ a two-step estimation process including Lambertian surface translation from unpaired data and depth estimation. The texture and specular reflection on the surface of an organ reduce the accuracy of depth estimations. We apply Lambertian surface translation to an endoscopic image to remove these texture and reflections. Then, we estimate the depth by using a fully convolutional network (FCN). During the training of the FCN, improvement of the object edge similarity between an estimated image and a ground truth depth image is important for getting better results. We introduced a muti-scale edge loss function to improve the accuracy of depth estimation. We quantitatively evaluated the proposed method using real colonoscopic images. The estimated depth values were proportional to the real depth values. Furthermore, we applied the estimated depth images to automated anatomical location identification of colonoscopic images using a convolutional neural network. The identification accuracy of the network improved from 69.2% to 74.1% by using the estimated depth images.
* Accepted paper as an oral presentation at Joint MICCAI workshop 2021, AE-CAI/CARE/OR2.0
COVID-19 Infection Segmentation from Chest CT Images Based on Scale Uncertainty
Jan 09, 2022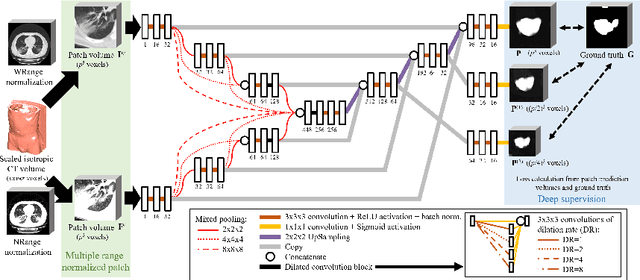
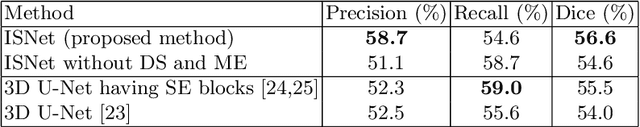
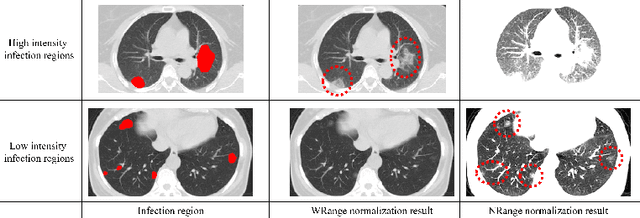

Abstract:This paper proposes a segmentation method of infection regions in the lung from CT volumes of COVID-19 patients. COVID-19 spread worldwide, causing many infected patients and deaths. CT image-based diagnosis of COVID-19 can provide quick and accurate diagnosis results. An automated segmentation method of infection regions in the lung provides a quantitative criterion for diagnosis. Previous methods employ whole 2D image or 3D volume-based processes. Infection regions have a considerable variation in their sizes. Such processes easily miss small infection regions. Patch-based process is effective for segmenting small targets. However, selecting the appropriate patch size is difficult in infection region segmentation. We utilize the scale uncertainty among various receptive field sizes of a segmentation FCN to obtain infection regions. The receptive field sizes can be defined as the patch size and the resolution of volumes where patches are clipped from. This paper proposes an infection segmentation network (ISNet) that performs patch-based segmentation and a scale uncertainty-aware prediction aggregation method that refines the segmentation result. We design ISNet to segment infection regions that have various intensity values. ISNet has multiple encoding paths to process patch volumes normalized by multiple intensity ranges. We collect prediction results generated by ISNets having various receptive field sizes. Scale uncertainty among the prediction results is extracted by the prediction aggregation method. We use an aggregation FCN to generate a refined segmentation result considering scale uncertainty among the predictions. In our experiments using 199 chest CT volumes of COVID-19 cases, the prediction aggregation method improved the dice similarity score from 47.6% to 62.1%.
* Accepted paper as a oral presentation at CILP2021, 10th MICCAI CLIP Workshop
Lung infection and normal region segmentation from CT volumes of COVID-19 cases
Jan 09, 2022Abstract:This paper proposes an automated segmentation method of infection and normal regions in the lung from CT volumes of COVID-19 patients. From December 2019, novel coronavirus disease 2019 (COVID-19) spreads over the world and giving significant impacts to our economic activities and daily lives. To diagnose the large number of infected patients, diagnosis assistance by computers is needed. Chest CT is effective for diagnosis of viral pneumonia including COVID-19. A quantitative analysis method of condition of the lung from CT volumes by computers is required for diagnosis assistance of COVID-19. This paper proposes an automated segmentation method of infection and normal regions in the lung from CT volumes using a COVID-19 segmentation fully convolutional network (FCN). In diagnosis of lung diseases including COVID-19, analysis of conditions of normal and infection regions in the lung is important. Our method recognizes and segments lung normal and infection regions in CT volumes. To segment infection regions that have various shapes and sizes, we introduced dense pooling connections and dilated convolutions in our FCN. We applied the proposed method to CT volumes of COVID-19 cases. From mild to severe cases of COVID-19, the proposed method correctly segmented normal and infection regions in the lung. Dice scores of normal and infection regions were 0.911 and 0.753, respectively.
* Accepted paper as a poster presentation at SPIE Medical Imaging 2021
Multi-task Federated Learning for Heterogeneous Pancreas Segmentation
Aug 19, 2021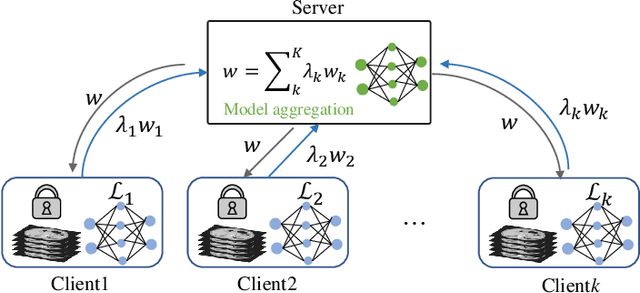
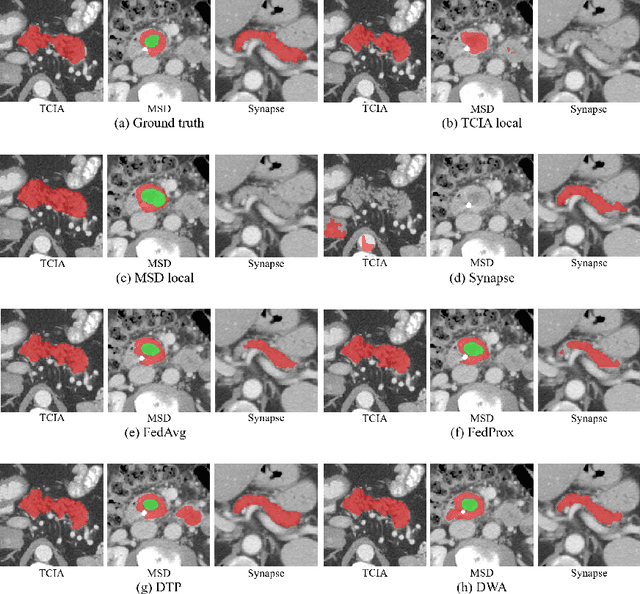
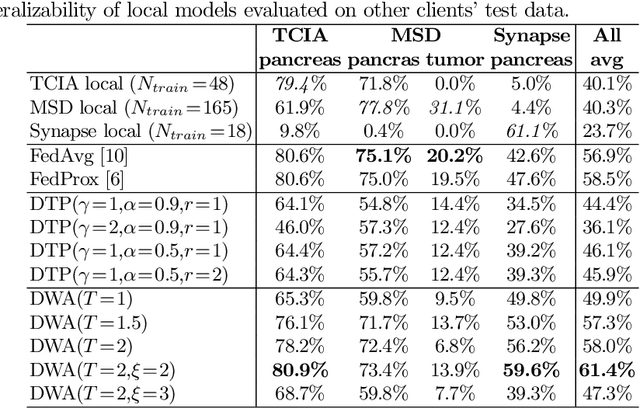
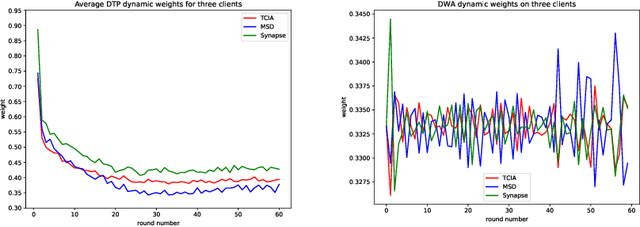

Abstract:Federated learning (FL) for medical image segmentation becomes more challenging in multi-task settings where clients might have different categories of labels represented in their data. For example, one client might have patient data with "healthy'' pancreases only while datasets from other clients may contain cases with pancreatic tumors. The vanilla federated averaging algorithm makes it possible to obtain more generalizable deep learning-based segmentation models representing the training data from multiple institutions without centralizing datasets. However, it might be sub-optimal for the aforementioned multi-task scenarios. In this paper, we investigate heterogeneous optimization methods that show improvements for the automated segmentation of pancreas and pancreatic tumors in abdominal CT images with FL settings.
Micro CT Image-Assisted Cross Modality Super-Resolution of Clinical CT Images Utilizing Synthesized Training Dataset
Oct 20, 2020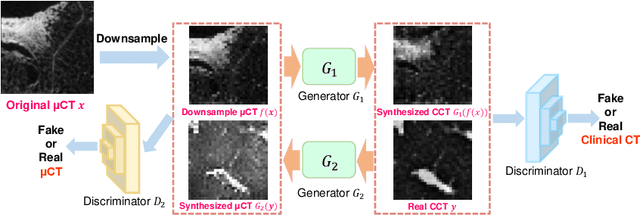
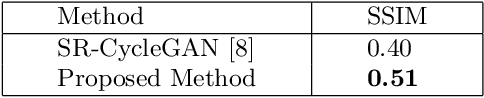
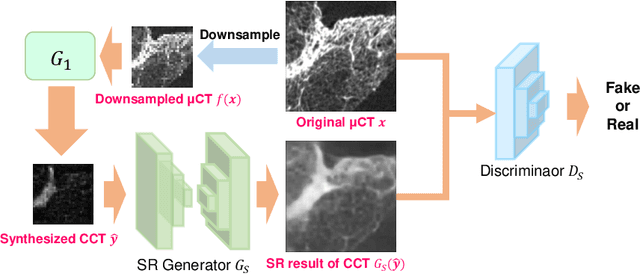

Abstract:This paper proposes a novel, unsupervised super-resolution (SR) approach for performing the SR of a clinical CT into the resolution level of a micro CT ($\mu$CT). The precise non-invasive diagnosis of lung cancer typically utilizes clinical CT data. Due to the resolution limitations of clinical CT (about $0.5 \times 0.5 \times 0.5$ mm$^3$), it is difficult to obtain enough pathological information such as the invasion area at alveoli level. On the other hand, $\mu$CT scanning allows the acquisition of volumes of lung specimens with much higher resolution ($50 \times 50 \times 50 \mu {\rm m}^3$ or higher). Thus, super-resolution of clinical CT volume may be helpful for diagnosis of lung cancer. Typical SR methods require aligned pairs of low-resolution (LR) and high-resolution (HR) images for training. Unfortunately, obtaining paired clinical CT and $\mu$CT volumes of human lung tissues is infeasible. Unsupervised SR methods are required that do not need paired LR and HR images. In this paper, we create corresponding clinical CT-$\mu$CT pairs by simulating clinical CT images from $\mu$CT images by modified CycleGAN. After this, we use simulated clinical CT-$\mu$CT image pairs to train an SR network based on SRGAN. Finally, we use the trained SR network to perform SR of the clinical CT images. We compare our proposed method with another unsupervised SR method for clinical CT images named SR-CycleGAN. Experimental results demonstrate that the proposed method can successfully perform SR of clinical CT images of lung cancer patients with $\mu$CT level resolution, and quantitatively and qualitatively outperformed conventional method (SR-CycleGAN), improving the SSIM (structure similarity) form 0.40 to 0.51.
Automated Pancreas Segmentation Using Multi-institutional Collaborative Deep Learning
Sep 28, 2020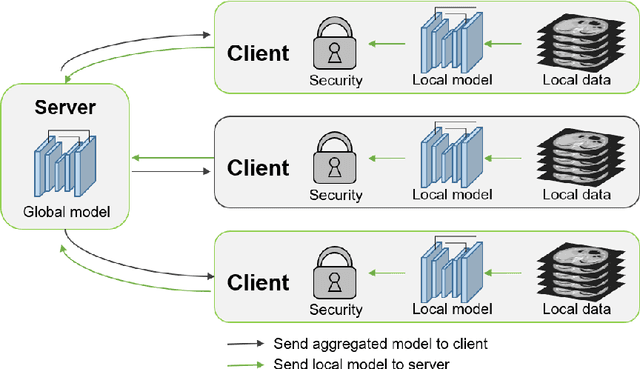
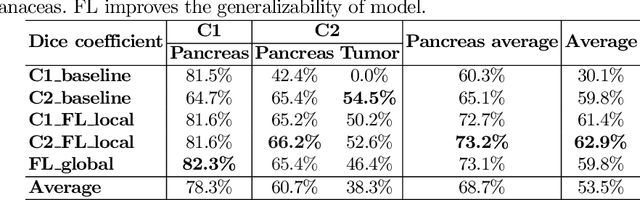
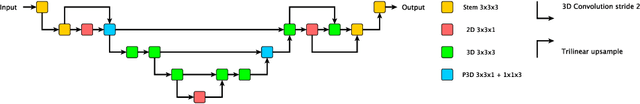
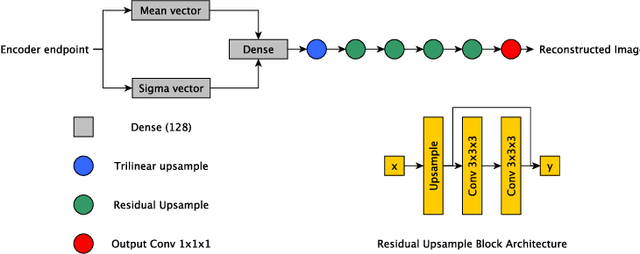
Abstract:The performance of deep learning-based methods strongly relies on the number of datasets used for training. Many efforts have been made to increase the data in the medical image analysis field. However, unlike photography images, it is hard to generate centralized databases to collect medical images because of numerous technical, legal, and privacy issues. In this work, we study the use of federated learning between two institutions in a real-world setting to collaboratively train a model without sharing the raw data across national boundaries. We quantitatively compare the segmentation models obtained with federated learning and local training alone. Our experimental results show that federated learning models have higher generalizability than standalone training.
Regression Forest-Based Atlas Localization and Direction Specific Atlas Generation for Pancreas Segmentation
May 07, 2020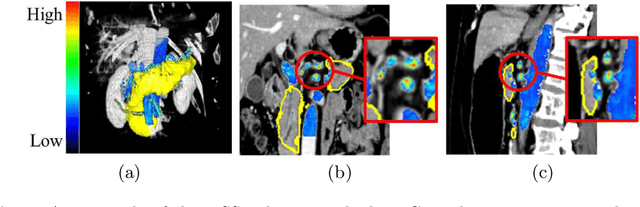
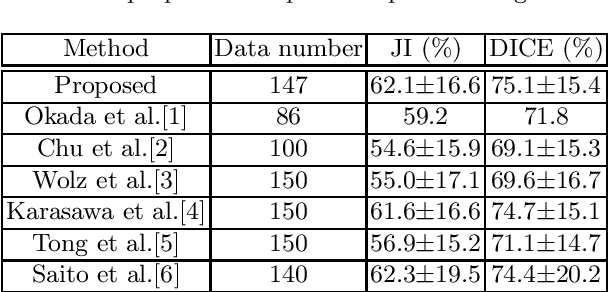
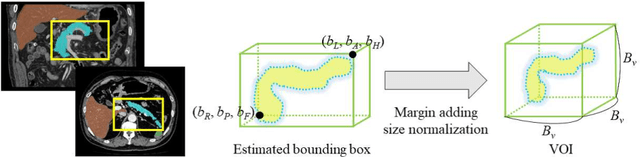
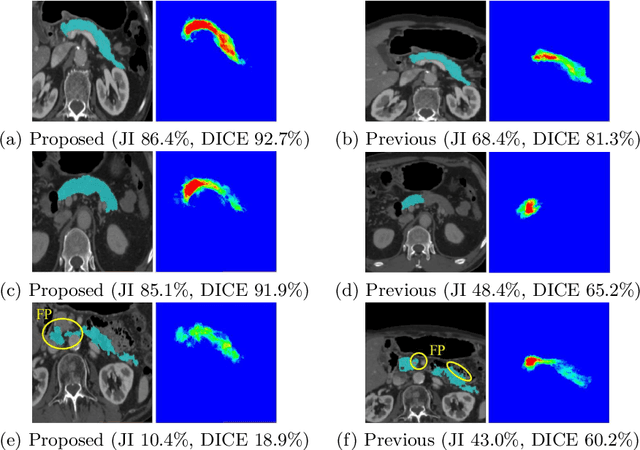
Abstract:This paper proposes a fully automated atlas-based pancreas segmentation method from CT volumes utilizing atlas localization by regression forest and atlas generation using blood vessel information. Previous probabilistic atlas-based pancreas segmentation methods cannot deal with spatial variations that are commonly found in the pancreas well. Also, shape variations are not represented by an averaged atlas. We propose a fully automated pancreas segmentation method that deals with two types of variations mentioned above. The position and size of the pancreas is estimated using a regression forest technique. After localization, a patient-specific probabilistic atlas is generated based on a new image similarity that reflects the blood vessel position and direction information around the pancreas. We segment it using the EM algorithm with the atlas as prior followed by the graph-cut. In evaluation results using 147 CT volumes, the Jaccard index and the Dice overlap of the proposed method were 62.1% and 75.1%, respectively. Although we automated all of the segmentation processes, segmentation results were superior to the other state-of-the-art methods in the Dice overlap.
* Accepted paper as a poster presentation at MICCAI 2016 (International Conference on Medical Image Computing and Computer-Assisted Intervention), Athens, Greece
Automated eye disease classification method from anterior eye image using anatomical structure focused image classification technique
May 04, 2020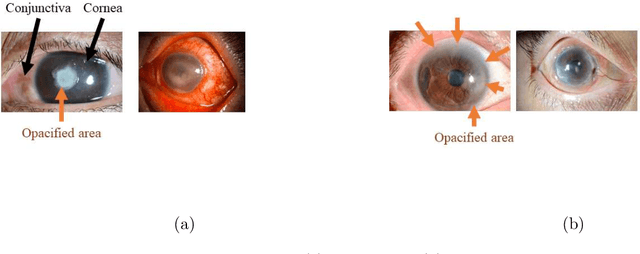
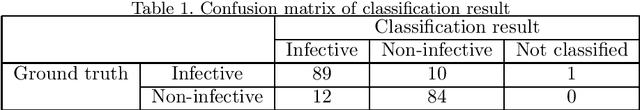
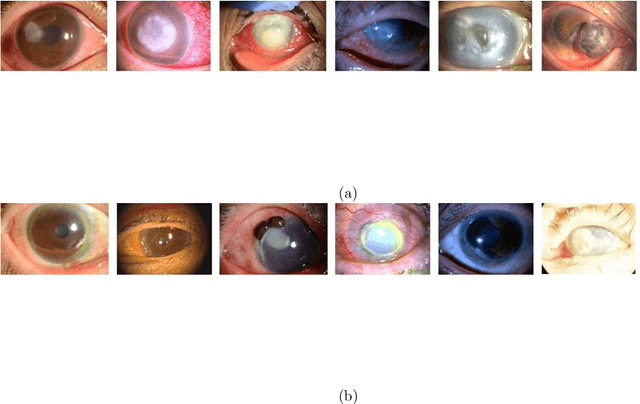
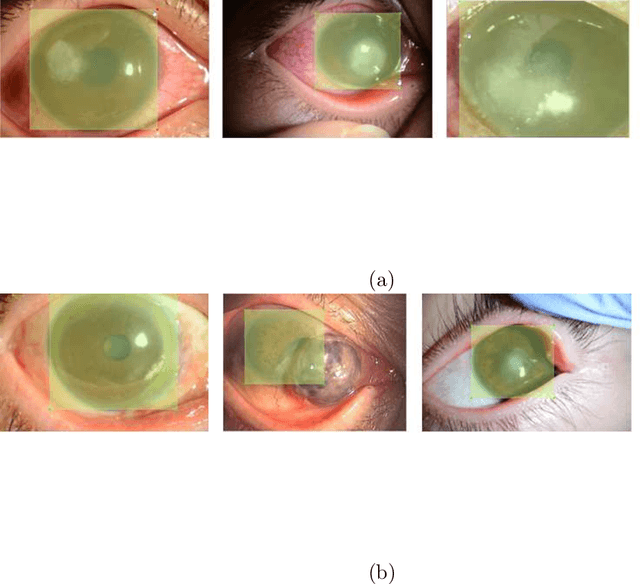
Abstract:This paper presents an automated classification method of infective and non-infective diseases from anterior eye images. Treatments for cases of infective and non-infective diseases are different. Distinguishing them from anterior eye images is important to decide a treatment plan. Ophthalmologists distinguish them empirically. Quantitative classification of them based on computer assistance is necessary. We propose an automated classification method of anterior eye images into cases of infective or non-infective disease. Anterior eye images have large variations of the eye position and brightness of illumination. This makes the classification difficult. If we focus on the cornea, positions of opacified areas in the corneas are different between cases of the infective and non-infective diseases. Therefore, we solve the anterior eye image classification task by using an object detection approach targeting the cornea. This approach can be said as "anatomical structure focused image classification". We use the YOLOv3 object detection method to detect corneas of infective disease and corneas of non-infective disease. The detection result is used to define a classification result of a image. In our experiments using anterior eye images, 88.3% of images were correctly classified by the proposed method.
* Accepted paper as a poster presentation at SPIE Medical Imaging 2020, Houston, TX, USA
Colonoscope tracking method based on shape estimation network
Apr 20, 2020Abstract:This paper presents a colonoscope tracking method utilizing a colon shape estimation method. CT colonography is used as a less-invasive colon diagnosis method. If colonic polyps or early-stage cancers are found, they are removed in a colonoscopic examination. In the colonoscopic examination, understanding where the colonoscope running in the colon is difficult. A colonoscope navigation system is necessary to reduce overlooking of polyps. We propose a colonoscope tracking method for navigation systems. Previous colonoscope tracking methods caused large tracking errors because they do not consider deformations of the colon during colonoscope insertions. We utilize the shape estimation network (SEN), which estimates deformed colon shape during colonoscope insertions. The SEN is a neural network containing long short-term memory (LSTM) layer. To perform colon shape estimation suitable to the real clinical situation, we trained the SEN using data obtained during colonoscope operations of physicians. The proposed tracking method performs mapping of the colonoscope tip position to a position in the colon using estimation results of the SEN. We evaluated the proposed method in a phantom study. We confirmed that tracking errors of the proposed method was enough small to perform navigation in the ascending, transverse, and descending colons.
* Accepted paper as an oral presentation at SPIE Medical Imaging 2019, San Diego, CA, USA
Colon Shape Estimation Method for Colonoscope Tracking using Recurrent Neural Networks
Apr 20, 2020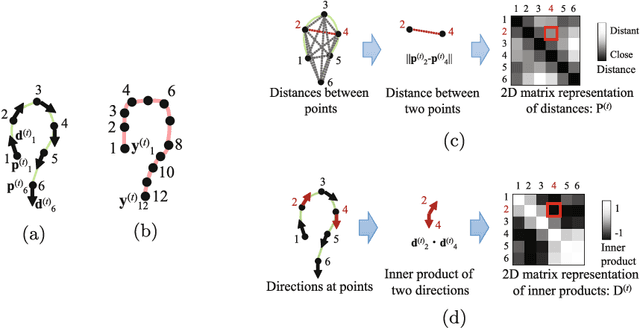
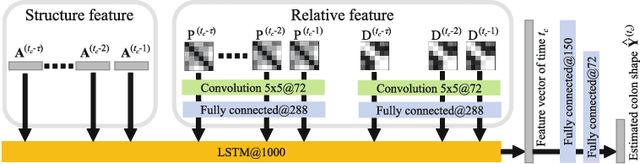
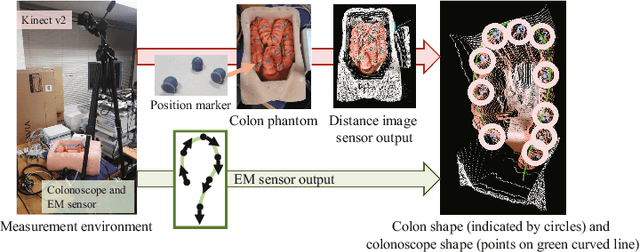
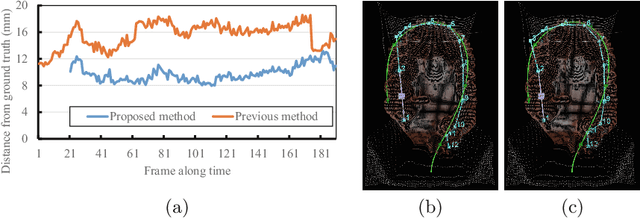
Abstract:We propose an estimation method using a recurrent neural network (RNN) of the colon's shape where deformation was occurred by a colonoscope insertion. Colonoscope tracking or a navigation system that navigates physician to polyp positions is needed to reduce such complications as colon perforation. Previous tracking methods caused large tracking errors at the transverse and sigmoid colons because these areas largely deform during colonoscope insertion. Colon deformation should be taken into account in tracking processes. We propose a colon deformation estimation method using RNN and obtain the colonoscope shape from electromagnetic sensors during its insertion into the colon. This method obtains positional, directional, and an insertion length from the colonoscope shape. From its shape, we also calculate the relative features that represent the positional and directional relationships between two points on a colonoscope. Long short-term memory is used to estimate the current colon shape from the past transition of the features of the colonoscope shape. We performed colon shape estimation in a phantom study and correctly estimated the colon shapes during colonoscope insertion with 12.39 (mm) estimation error.
* Accepted paper as a poster presentation at MICCAI 2018 (International Conference on Medical Image Computing and Computer-Assisted Intervention), Granada, Spain
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge