Katarzyna Bozek
Understanding Cell Fate Decisions with Temporal Attention
Mar 17, 2026Abstract:Understanding non-genetic determinants of cell fate is critical for developing and improving cancer therapies, as genetically identical cells can exhibit divergent outcomes under the same treatment conditions. In this work, we present a deep learning approach for cell fate prediction from raw long-term live-cell recordings of cancer cell populations under chemotherapeutic treatment. Our Transformer model is trained to predict cell fate directly from raw image sequences, without relying on predefined morphological or molecular features. Beyond classification, we introduce a comprehensive explainability framework for interpreting the temporal and morphological cues guiding the model's predictions. We demonstrate that prediction of cell outcomes is possible based on the video only, our model achieves balanced accuracy of 0.94 and an F1-score of 0.93. Attention and masking experiments further indicate that the signal predictive of the cell fate is not uniquely located in the final frames of a cell trajectory, as reliable predictions are possible up to 10 h before the event. Our analysis reveals distinct temporal distribution of predictive information in the mitotic and apoptotic sequences, as well as the role of cell morphology and p53 signaling in determining cell outcomes. Together, these findings demonstrate that attention-based temporal models enable accurate cell fate prediction while providing biologically interpretable insights into non-genetic determinants of cellular decision-making. The code is available at https://github.com/bozeklab/Cell-Fate-Prediction.
Self-Supervised ImageNet Representations for In Vivo Confocal Microscopy: Tortuosity Grading without Segmentation Maps
Mar 16, 2026Abstract:The tortuosity of corneal nerve fibers are used as indication for different diseases. Current state-of-the-art methods for grading the tortuosity heavily rely on expensive segmentation maps of these nerve fibers. In this paper, we demonstrate that self-supervised pretrained features from ImageNet are transferable to the domain of in vivo confocal microscopy. We show that DINO should not be disregarded as a deep learning model for medical imaging, although it was superseded by two later versions. After careful fine-tuning, DINO improves upon the state-of-the-art in terms of accuracy (84,25%) and sensitivity (77,97%). Our fine-tuned model focuses on the key morphological elements in grading without the use of segmentation maps.
Context-aware Skin Cancer Epithelial Cell Classification with Scalable Graph Transformers
Feb 17, 2026Abstract:Whole-slide images (WSIs) from cancer patients contain rich information that can be used for medical diagnosis or to follow treatment progress. To automate their analysis, numerous deep learning methods based on convolutional neural networks and Vision Transformers have been developed and have achieved strong performance in segmentation and classification tasks. However, due to the large size and complex cellular organization of WSIs, these models rely on patch-based representations, losing vital tissue-level context. We propose using scalable Graph Transformers on a full-WSI cell graph for classification. We evaluate this methodology on a challenging task: the classification of healthy versus tumor epithelial cells in cutaneous squamous cell carcinoma (cSCC), where both cell types exhibit very similar morphologies and are therefore difficult to differentiate for image-based approaches. We first compared image-based and graph-based methods on a single WSI. Graph Transformer models SGFormer and DIFFormer achieved balanced accuracies of $85.2 \pm 1.5$ ($\pm$ standard error) and $85.1 \pm 2.5$ in 3-fold cross-validation, respectively, whereas the best image-based method reached $81.2 \pm 3.0$. By evaluating several node feature configurations, we found that the most informative representation combined morphological and texture features as well as the cell classes of non-epithelial cells, highlighting the importance of the surrounding cellular context. We then extended our work to train on several WSIs from several patients. To address the computational constraints of image-based models, we extracted four $2560 \times 2560$ pixel patches from each image and converted them into graphs. In this setting, DIFFormer achieved a balanced accuracy of $83.6 \pm 1.9$ (3-fold cross-validation), while the state-of-the-art image-based model CellViT256 reached $78.1 \pm 0.5$.
AMAP-APP: Efficient Segmentation and Morphometry Quantification of Fluorescent Microscopy Images of Podocytes
Feb 16, 2026Abstract:Background: Automated podocyte foot process quantification is vital for kidney research, but the established "Automatic Morphological Analysis of Podocytes" (AMAP) method is hindered by high computational demands, a lack of a user interface, and Linux dependency. We developed AMAP-APP, a cross-platform desktop application designed to overcome these barriers. Methods: AMAP-APP optimizes efficiency by replacing intensive instance segmentation with classic image processing while retaining the original semantic segmentation model. It introduces a refined Region of Interest (ROI) algorithm to improve precision. Validation involved 365 mouse and human images (STED and confocal), benchmarking performance against the original AMAP via Pearson correlation and Two One-Sided T-tests (TOST). Results: AMAP-APP achieved a 147-fold increase in processing speed on consumer hardware. Morphometric outputs (area, perimeter, circularity, and slit diaphragm density) showed high correlation (r>0.90) and statistical equivalence (TOST P<0.05) to the original method. Additionally, the new ROI algorithm demonstrated superior accuracy compared to the original, showing reduced deviation from manual delineations. Conclusion: AMAP-APP democratizes deep learning-based podocyte morphometry. By eliminating the need for high-performance computing clusters and providing a user-friendly interface for Windows, macOS, and Linux, it enables widespread adoption in nephrology research and potential clinical diagnostics.
ActiTect: A Generalizable Machine Learning Pipeline for REM Sleep Behavior Disorder Screening through Standardized Actigraphy
Nov 12, 2025Abstract:Isolated rapid eye movement sleep behavior disorder (iRBD) is a major prodromal marker of $α$-synucleinopathies, often preceding the clinical onset of Parkinson's disease, dementia with Lewy bodies, or multiple system atrophy. While wrist-worn actimeters hold significant potential for detecting RBD in large-scale screening efforts by capturing abnormal nocturnal movements, they become inoperable without a reliable and efficient analysis pipeline. This study presents ActiTect, a fully automated, open-source machine learning tool to identify RBD from actigraphy recordings. To ensure generalizability across heterogeneous acquisition settings, our pipeline includes robust preprocessing and automated sleep-wake detection to harmonize multi-device data and extract physiologically interpretable motion features characterizing activity patterns. Model development was conducted on a cohort of 78 individuals, yielding strong discrimination under nested cross-validation (AUROC = 0.95). Generalization was confirmed on a blinded local test set (n = 31, AUROC = 0.86) and on two independent external cohorts (n = 113, AUROC = 0.84; n = 57, AUROC = 0.94). To assess real-world robustness, leave-one-dataset-out cross-validation across the internal and external cohorts demonstrated consistent performance (AUROC range = 0.84-0.89). A complementary stability analysis showed that key predictive features remained reproducible across datasets, supporting the final pooled multi-center model as a robust pre-trained resource for broader deployment. By being open-source and easy to use, our tool promotes widespread adoption and facilitates independent validation and collaborative improvements, thereby advancing the field toward a unified and generalizable RBD detection model using wearable devices.
Histo-Miner: Deep Learning based Tissue Features Extraction Pipeline from H&E Whole Slide Images of Cutaneous Squamous Cell Carcinoma
May 07, 2025Abstract:Recent advancements in digital pathology have enabled comprehensive analysis of Whole-Slide Images (WSI) from tissue samples, leveraging high-resolution microscopy and computational capabilities. Despite this progress, there is a lack of labeled datasets and open source pipelines specifically tailored for analysis of skin tissue. Here we propose Histo-Miner, a deep learning-based pipeline for analysis of skin WSIs and generate two datasets with labeled nuclei and tumor regions. We develop our pipeline for the analysis of patient samples of cutaneous squamous cell carcinoma (cSCC), a frequent non-melanoma skin cancer. Utilizing the two datasets, comprising 47,392 annotated cell nuclei and 144 tumor-segmented WSIs respectively, both from cSCC patients, Histo-Miner employs convolutional neural networks and vision transformers for nucleus segmentation and classification as well as tumor region segmentation. Performance of trained models positively compares to state of the art with multi-class Panoptic Quality (mPQ) of 0.569 for nucleus segmentation, macro-averaged F1 of 0.832 for nucleus classification and mean Intersection over Union (mIoU) of 0.884 for tumor region segmentation. From these predictions we generate a compact feature vector summarizing tissue morphology and cellular interactions, which can be used for various downstream tasks. Here, we use Histo-Miner to predict cSCC patient response to immunotherapy based on pre-treatment WSIs from 45 patients. Histo-Miner identifies percentages of lymphocytes, the granulocyte to lymphocyte ratio in tumor vicinity and the distances between granulocytes and plasma cells in tumors as predictive features for therapy response. This highlights the applicability of Histo-Miner to clinically relevant scenarios, providing direct interpretation of the classification and insights into the underlying biology.
KPIs 2024 Challenge: Advancing Glomerular Segmentation from Patch- to Slide-Level
Feb 11, 2025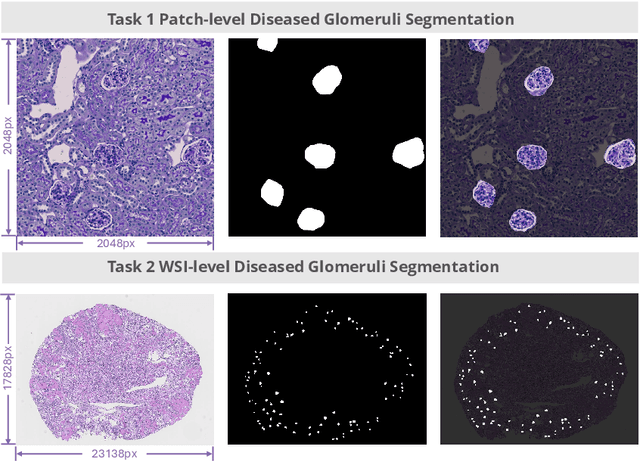
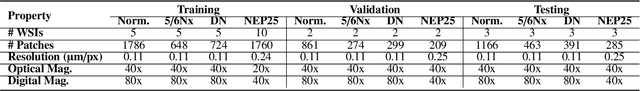
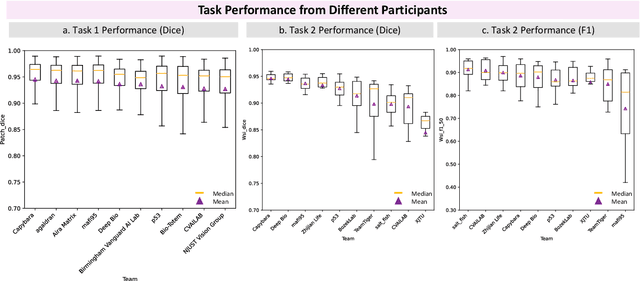

Abstract:Chronic kidney disease (CKD) is a major global health issue, affecting over 10% of the population and causing significant mortality. While kidney biopsy remains the gold standard for CKD diagnosis and treatment, the lack of comprehensive benchmarks for kidney pathology segmentation hinders progress in the field. To address this, we organized the Kidney Pathology Image Segmentation (KPIs) Challenge, introducing a dataset that incorporates preclinical rodent models of CKD with over 10,000 annotated glomeruli from 60+ Periodic Acid Schiff (PAS)-stained whole slide images. The challenge includes two tasks, patch-level segmentation and whole slide image segmentation and detection, evaluated using the Dice Similarity Coefficient (DSC) and F1-score. By encouraging innovative segmentation methods that adapt to diverse CKD models and tissue conditions, the KPIs Challenge aims to advance kidney pathology analysis, establish new benchmarks, and enable precise, large-scale quantification for disease research and diagnosis.
Fine-tuning a Multiple Instance Learning Feature Extractor with Masked Context Modelling and Knowledge Distillation
Mar 08, 2024



Abstract:The first step in Multiple Instance Learning (MIL) algorithms for Whole Slide Image (WSI) classification consists of tiling the input image into smaller patches and computing their feature vectors produced by a pre-trained feature extractor model. Feature extractor models that were pre-trained with supervision on ImageNet have proven to transfer well to this domain, however, this pre-training task does not take into account that visual information in neighboring patches is highly correlated. Based on this observation, we propose to increase downstream MIL classification by fine-tuning the feature extractor model using \textit{Masked Context Modelling with Knowledge Distillation}. In this task, the feature extractor model is fine-tuned by predicting masked patches in a bigger context window. Since reconstructing the input image would require a powerful image generation model, and our goal is not to generate realistically looking image patches, we predict instead the feature vectors produced by a larger teacher network. A single epoch of the proposed task suffices to increase the downstream performance of the feature-extractor model when used in a MIL scenario, even capable of outperforming the downstream performance of the teacher model, while being considerably smaller and requiring a fraction of its compute.
Language models are good pathologists: using attention-based sequence reduction and text-pretrained transformers for efficient WSI classification
Nov 14, 2022



Abstract:In digital pathology, Whole Slide Image (WSI) analysis is usually formulated as a Multiple Instance Learning (MIL) problem. Although transformer-based architectures have been used for WSI classification, these methods require modifications to adapt them to specific challenges of this type of image data. Despite their power across domains, reference transformer models in classical Computer Vision (CV) and Natural Language Processing (NLP) tasks are not used for pathology slide analysis. In this work we demonstrate the use of standard, frozen, text-pretrained, transformer language models in application to WSI classification. We propose SeqShort, a multi-head attention-based sequence reduction input layer to summarize each WSI in a fixed and short size sequence of instances. This allows us to reduce the computational costs of self-attention on long sequences, and to include positional information that is unavailable in other MIL approaches. We demonstrate the effectiveness of our methods in the task of cancer subtype classification, without the need of designing a WSI-specific transformer or performing in-domain self-supervised pretraining, while keeping a reduced compute budget and number of trainable parameters.
Pixel personality for dense object tracking in a 2D honeybee hive
Dec 31, 2018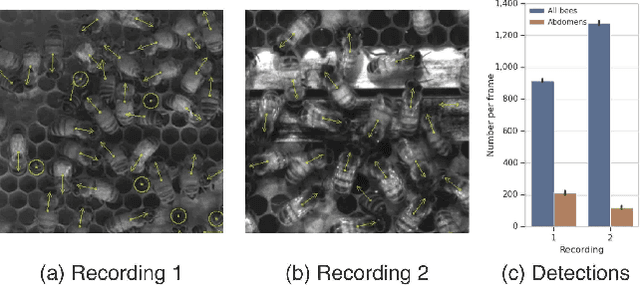
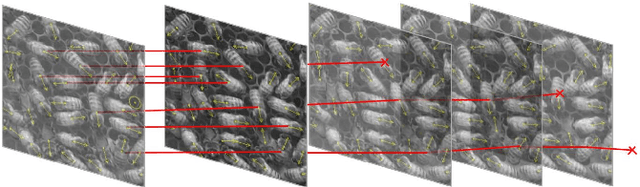
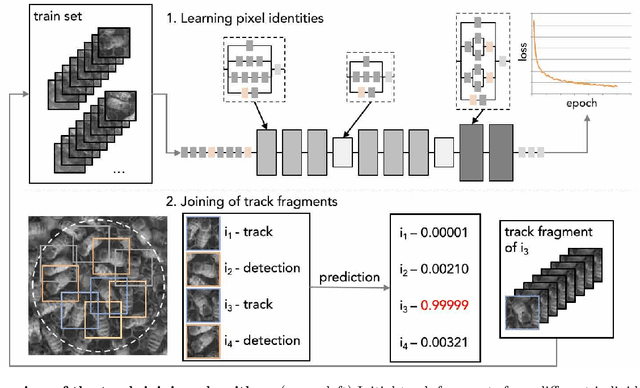
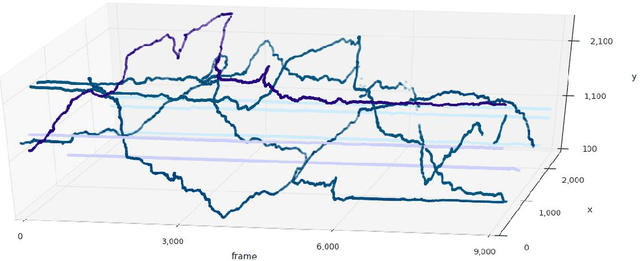
Abstract:Tracking large numbers of densely-arranged, interacting objects is challenging due to occlusions and the resulting complexity of possible trajectory combinations, as well as the sparsity of relevant, labeled datasets. Here we describe a novel technique of collective tracking in the model environment of a 2D honeybee hive in which sample colonies consist of $N\sim10^3$ highly similar individuals, tightly packed, and in rapid, irregular motion. Such a system offers universal challenges for multi-object tracking, while being conveniently accessible for image recording. We first apply an accurate, segmentation-based object detection method to build initial short trajectory segments by matching object configurations based on class, position and orientation. We then join these tracks into full single object trajectories by creating an object recognition model which is adaptively trained to recognize honeybee individuals through their visual appearance across multiple frames, an attribute we denote as pixel personality. Overall, we reconstruct ~46% of the trajectories in 5 min recordings from two different hives and over 71% of the tracks for at least 2 min. We provide validated trajectories spanning 3000 video frames of 876 unmarked moving bees in two distinct colonies in different locations and filmed with different pixel resolutions, which we expect to be useful in the further development of general-purpose tracking solutions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge