Karthikeyan Natesan Ramamurthy
ICML 2023 Topological Deep Learning Challenge : Design and Results
Oct 02, 2023

Abstract:This paper presents the computational challenge on topological deep learning that was hosted within the ICML 2023 Workshop on Topology and Geometry in Machine Learning. The competition asked participants to provide open-source implementations of topological neural networks from the literature by contributing to the python packages TopoNetX (data processing) and TopoModelX (deep learning). The challenge attracted twenty-eight qualifying submissions in its two-month duration. This paper describes the design of the challenge and summarizes its main findings.
The Impact of Positional Encoding on Length Generalization in Transformers
May 31, 2023



Abstract:Length generalization, the ability to generalize from small training context sizes to larger ones, is a critical challenge in the development of Transformer-based language models. Positional encoding (PE) has been identified as a major factor influencing length generalization, but the exact impact of different PE schemes on extrapolation in downstream tasks remains unclear. In this paper, we conduct a systematic empirical study comparing the length generalization performance of decoder-only Transformers with five different position encoding approaches including Absolute Position Embedding (APE), T5's Relative PE, ALiBi, and Rotary, in addition to Transformers without positional encoding (NoPE). Our evaluation encompasses a battery of reasoning and mathematical tasks. Our findings reveal that the most commonly used positional encoding methods, such as ALiBi, Rotary, and APE, are not well suited for length generalization in downstream tasks. More importantly, NoPE outperforms other explicit positional encoding methods while requiring no additional computation. We theoretically demonstrate that NoPE can represent both absolute and relative PEs, but when trained with SGD, it mostly resembles T5's relative PE attention patterns. Finally, we find that scratchpad is not always helpful to solve length generalization and its format highly impacts the model's performance. Overall, our work suggests that explicit position embeddings are not essential for decoder-only Transformers to generalize well to longer sequences.
Function Composition in Trustworthy Machine Learning: Implementation Choices, Insights, and Questions
Feb 17, 2023



Abstract:Ensuring trustworthiness in machine learning (ML) models is a multi-dimensional task. In addition to the traditional notion of predictive performance, other notions such as privacy, fairness, robustness to distribution shift, adversarial robustness, interpretability, explainability, and uncertainty quantification are important considerations to evaluate and improve (if deficient). However, these sub-disciplines or 'pillars' of trustworthiness have largely developed independently, which has limited us from understanding their interactions in real-world ML pipelines. In this paper, focusing specifically on compositions of functions arising from the different pillars, we aim to reduce this gap, develop new insights for trustworthy ML, and answer questions such as the following. Does the composition of multiple fairness interventions result in a fairer model compared to a single intervention? How do bias mitigation algorithms for fairness affect local post-hoc explanations? Does a defense algorithm for untargeted adversarial attacks continue to be effective when composed with a privacy transformation? Toward this end, we report initial empirical results and new insights from 9 different compositions of functions (or pipelines) on 7 real-world datasets along two trustworthy dimensions - fairness and explainability. We also report progress, and implementation choices, on an extensible composer tool to encourage the combination of functionalities from multiple pillars. To-date, the tool supports bias mitigation algorithms for fairness and post-hoc explainability methods. We hope this line of work encourages the thoughtful consideration of multiple pillars when attempting to formulate and resolve a trustworthiness problem.
Active Sequential Two-Sample Testing
Feb 02, 2023



Abstract:Two-sample testing tests whether the distributions generating two samples are identical. We pose the two-sample testing problem in a new scenario where the sample measurements (or sample features) are inexpensive to access, but their group memberships (or labels) are costly. We devise the first \emph{active sequential two-sample testing framework} that not only sequentially but also \emph{actively queries} sample labels to address the problem. Our test statistic is a likelihood ratio where one likelihood is found by maximization over all class priors, and the other is given by a classification model. The classification model is adaptively updated and then used to guide an active query scheme called bimodal query to label sample features in the regions with high dependency between the feature variables and the label variables. The theoretical contributions in the paper include proof that our framework produces an \emph{anytime-valid} $p$-value; and, under reachable conditions and a mild assumption, the framework asymptotically generates a minimum normalized log-likelihood ratio statistic that a passive query scheme can only achieve when the feature variable and the label variable have the highest dependence. Lastly, we provide a \emph{query-switching (QS)} algorithm to decide when to switch from passive query to active query and adapt bimodal query to increase the testing power of our test. Extensive experiments justify our theoretical contributions and the effectiveness of QS.
Equi-Tuning: Group Equivariant Fine-Tuning of Pretrained Models
Oct 13, 2022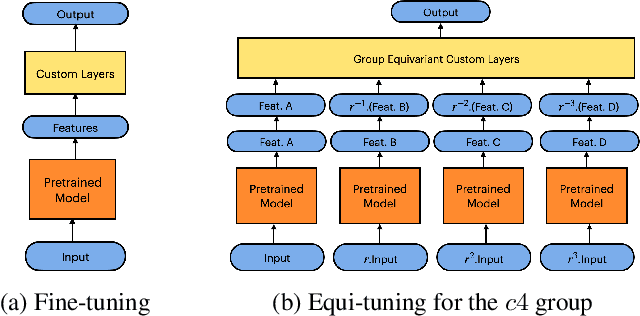
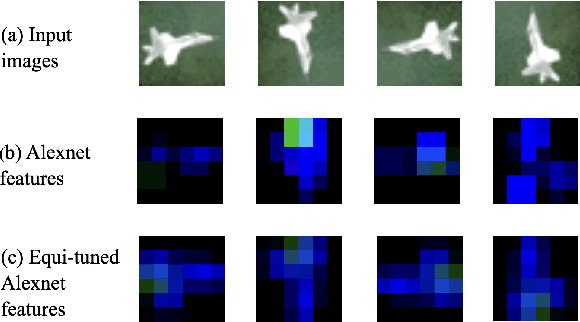
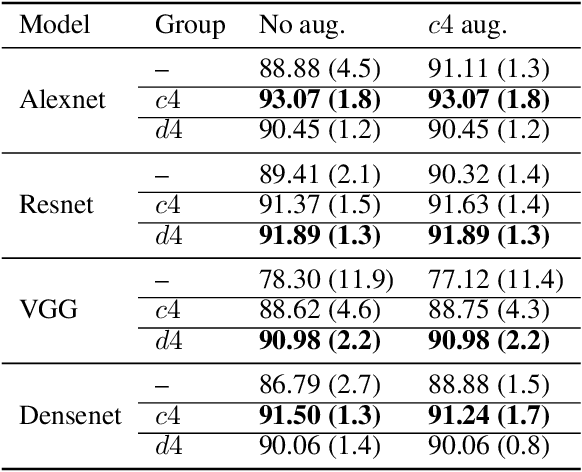

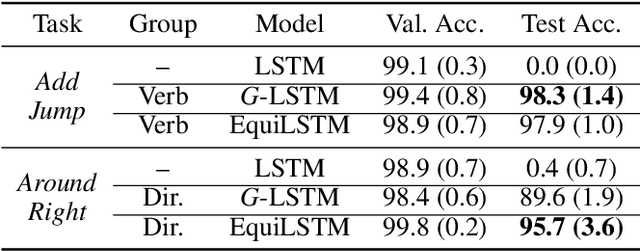
Abstract:We introduce equi-tuning, a novel fine-tuning method that transforms (potentially non-equivariant) pretrained models into group equivariant models while incurring minimum $L_2$ loss between the feature representations of the pretrained and the equivariant models. Large pretrained models can be equi-tuned for different groups to satisfy the needs of various downstream tasks. Equi-tuned models benefit from both group equivariance as an inductive bias and semantic priors from pretrained models. We provide applications of equi-tuning on three different tasks: image classification, compositional generalization in language, and fairness in natural language generation (NLG). We also provide a novel group-theoretic definition for fairness in NLG. The effectiveness of this definition is shown by testing it against a standard empirical method of fairness in NLG. We provide experimental results for equi-tuning using a variety of pretrained models: Alexnet, Resnet, VGG, and Densenet for image classification; RNNs, GRUs, and LSTMs for compositional generalization; and GPT2 for fairness in NLG. We test these models on benchmark datasets across all considered tasks to show the generality and effectiveness of the proposed method.
Higher-Order Attention Networks
Jun 01, 2022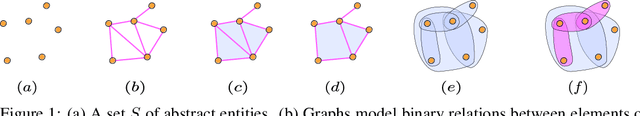
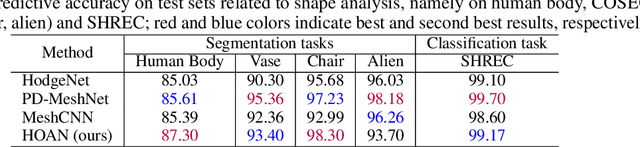
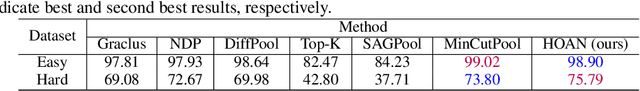

Abstract:This paper introduces higher-order attention networks (HOANs), a novel class of attention-based neural networks defined on a generalized higher-order domain called a combinatorial complex (CC). Similar to hypergraphs, CCs admit arbitrary set-like relations between a collection of abstract entities. Simultaneously, CCs permit the construction of hierarchical higher-order relations analogous to those supported by cell complexes. Thus, CCs effectively generalize both hypergraphs and cell complexes and combine their desirable characteristics. By exploiting the rich combinatorial nature of CCs, HOANs define a new class of message-passing attention-based networks that unifies higher-order neural networks. Our evaluation on tasks related to mesh shape analysis and graph learning demonstrates that HOANs attain competitive, and in some examples superior, predictive performance in comparison to state-of-the-art neural networks.
Analogies and Feature Attributions for Model Agnostic Explanation of Similarity Learners
Feb 02, 2022
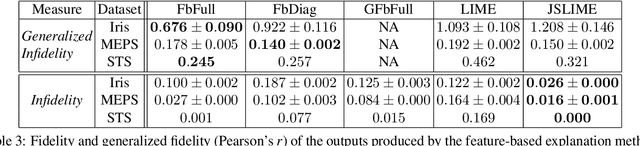
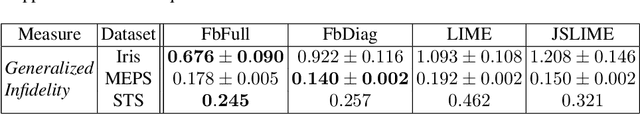
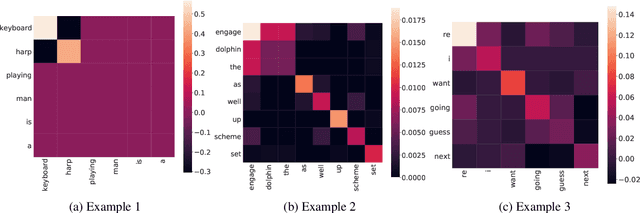

Abstract:Post-hoc explanations for black box models have been studied extensively in classification and regression settings. However, explanations for models that output similarity between two inputs have received comparatively lesser attention. In this paper, we provide model agnostic local explanations for similarity learners applicable to tabular and text data. We first propose a method that provides feature attributions to explain the similarity between a pair of inputs as determined by a black box similarity learner. We then propose analogies as a new form of explanation in machine learning. Here the goal is to identify diverse analogous pairs of examples that share the same level of similarity as the input pair and provide insight into (latent) factors underlying the model's prediction. The selection of analogies can optionally leverage feature attributions, thus connecting the two forms of explanation while still maintaining complementarity. We prove that our analogy objective function is submodular, making the search for good-quality analogies efficient. We apply the proposed approaches to explain similarities between sentences as predicted by a state-of-the-art sentence encoder, and between patients in a healthcare utilization application. Efficacy is measured through quantitative evaluations, a careful user study, and examples of explanations.
Ground-Truth, Whose Truth? -- Examining the Challenges with Annotating Toxic Text Datasets
Dec 07, 2021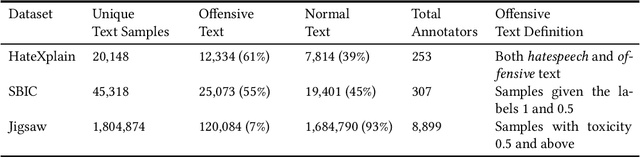
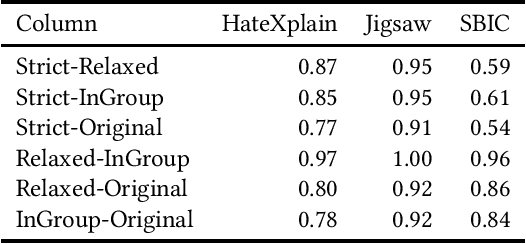
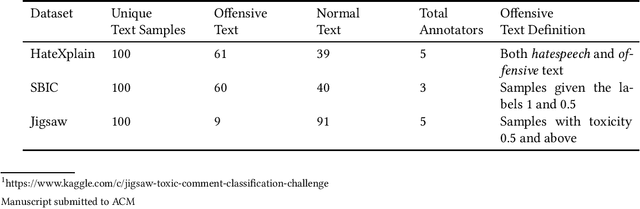
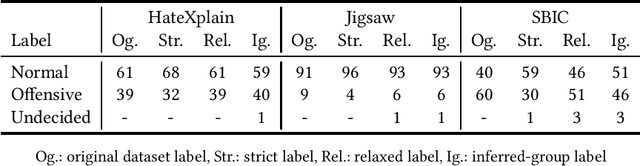

Abstract:The use of machine learning (ML)-based language models (LMs) to monitor content online is on the rise. For toxic text identification, task-specific fine-tuning of these models are performed using datasets labeled by annotators who provide ground-truth labels in an effort to distinguish between offensive and normal content. These projects have led to the development, improvement, and expansion of large datasets over time, and have contributed immensely to research on natural language. Despite the achievements, existing evidence suggests that ML models built on these datasets do not always result in desirable outcomes. Therefore, using a design science research (DSR) approach, this study examines selected toxic text datasets with the goal of shedding light on some of the inherent issues and contributing to discussions on navigating these challenges for existing and future projects. To achieve the goal of the study, we re-annotate samples from three toxic text datasets and find that a multi-label approach to annotating toxic text samples can help to improve dataset quality. While this approach may not improve the traditional metric of inter-annotator agreement, it may better capture dependence on context and diversity in annotators. We discuss the implications of these results for both theory and practice.
A label efficient two-sample test
Nov 29, 2021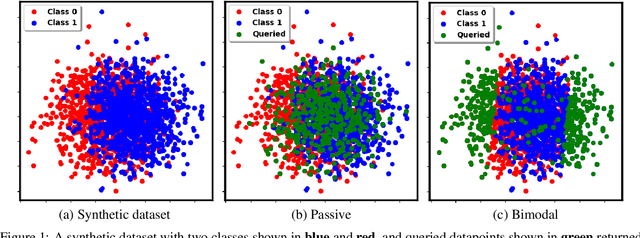
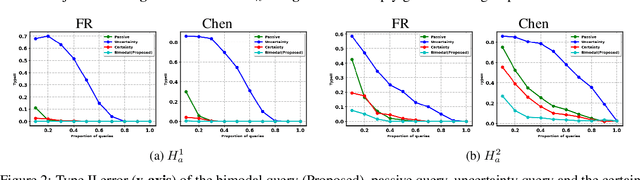
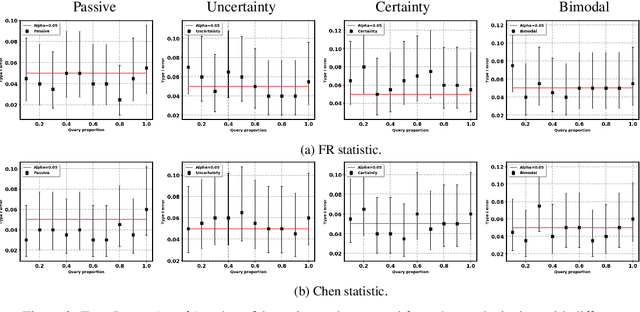
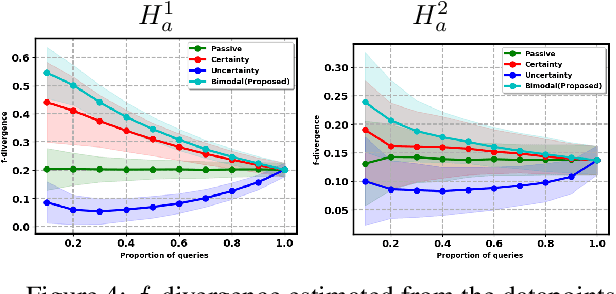
Abstract:Two-sample tests evaluate whether two samples are realizations of the same distribution (the null hypothesis) or two different distributions (the alternative hypothesis). In the traditional formulation of this problem, the statistician has access to both the measurements (feature variables) and the group variable (label variable). However, in several important applications, feature variables can be easily measured but the binary label variable is unknown and costly to obtain. In this paper, we consider this important variation on the classical two-sample test problem and pose it as a problem of obtaining the labels of only a small number of samples in service of performing a two-sample test. We devise a label efficient three-stage framework: firstly, a classifier is trained with samples uniformly labeled to model the posterior probabilities of the labels; secondly, a novel query scheme dubbed \emph{bimodal query} is used to query labels of samples from both classes with maximum posterior probabilities, and lastly, the classical Friedman-Rafsky (FR) two-sample test is performed on the queried samples. Our theoretical analysis shows that bimodal query is optimal for two-sample testing using the FR statistic under reasonable conditions and that the three-stage framework controls the Type I error. Extensive experiments performed on synthetic, benchmark, and application-specific datasets demonstrate that the three-stage framework has decreased Type II error over uniform querying and certainty-based querying with same number of labels while controlling the Type I error. Source code for our algorithms and experimental results is available at https://github.com/wayne0908/Label-Efficient-Two-Sample.
Data-Centric AI Requires Rethinking Data Notion
Oct 13, 2021Abstract:The transition towards data-centric AI requires revisiting data notions from mathematical and implementational standpoints to obtain unified data-centric machine learning packages. Towards this end, this work proposes unifying principles offered by categorical and cochain notions of data, and discusses the importance of these principles in data-centric AI transition. In the categorical notion, data is viewed as a mathematical structure that we act upon via morphisms to preserve this structure. As for cochain notion, data can be viewed as a function defined in a discrete domain of interest and acted upon via operators. While these notions are almost orthogonal, they provide a unifying definition to view data, ultimately impacting the way machine learning packages are developed, implemented, and utilized by practitioners.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge