José Miguel Hernández-Lobato
RECOMBINER: Robust and Enhanced Compression with Bayesian Implicit Neural Representations
Sep 29, 2023



Abstract:COMpression with Bayesian Implicit NEural Representations (COMBINER) is a recent data compression method that addresses a key inefficiency of previous Implicit Neural Representation (INR)-based approaches: it avoids quantization and enables direct optimization of the rate-distortion performance. However, COMBINER still has significant limitations: 1) it uses factorized priors and posterior approximations that lack flexibility; 2) it cannot effectively adapt to local deviations from global patterns in the data; and 3) its performance can be susceptible to modeling choices and the variational parameters' initializations. Our proposed method, Robust and Enhanced COMBINER (RECOMBINER), addresses these issues by 1) enriching the variational approximation while maintaining its computational cost via a linear reparameterization of the INR weights, 2) augmenting our INRs with learnable positional encodings that enable them to adapt to local details and 3) splitting high-resolution data into patches to increase robustness and utilizing expressive hierarchical priors to capture dependency across patches. We conduct extensive experiments across several data modalities, showcasing that RECOMBINER achieves competitive results with the best INR-based methods and even outperforms autoencoder-based codecs on low-resolution images at low bitrates.
SE(3) Equivariant Augmented Coupling Flows
Aug 20, 2023



Abstract:Coupling normalizing flows allow for fast sampling and density evaluation, making them the tool of choice for probabilistic modeling of physical systems. However, the standard coupling architecture precludes endowing flows that operate on the Cartesian coordinates of atoms with the SE(3) and permutation invariances of physical systems. This work proposes a coupling flow that preserves SE(3) and permutation equivariance by performing coordinate splits along additional augmented dimensions. At each layer, the flow maps atoms' positions into learned SE(3) invariant bases, where we apply standard flow transformations, such as monotonic rational-quadratic splines, before returning to the original basis. Crucially, our flow preserves fast sampling and density evaluation, and may be used to produce unbiased estimates of expectations with respect to the target distribution via importance sampling. When trained on the DW4, LJ13 and QM9-positional datasets, our flow is competitive with equivariant continuous normalizing flows, while allowing sampling two orders of magnitude faster. Moreover, to the best of our knowledge, we are the first to learn the full Boltzmann distribution of alanine dipeptide by only modeling the Cartesian positions of its atoms. Lastly, we demonstrate that our flow can be trained to approximately sample from the Boltzmann distribution of the DW4 and LJ13 particle systems using only their energy functions.
Minimal Random Code Learning with Mean-KL Parameterization
Jul 15, 2023


Abstract:This paper studies the qualitative behavior and robustness of two variants of Minimal Random Code Learning (MIRACLE) used to compress variational Bayesian neural networks. MIRACLE implements a powerful, conditionally Gaussian variational approximation for the weight posterior $Q_{\mathbf{w}}$ and uses relative entropy coding to compress a weight sample from the posterior using a Gaussian coding distribution $P_{\mathbf{w}}$. To achieve the desired compression rate, $D_{\mathrm{KL}}[Q_{\mathbf{w}} \Vert P_{\mathbf{w}}]$ must be constrained, which requires a computationally expensive annealing procedure under the conventional mean-variance (Mean-Var) parameterization for $Q_{\mathbf{w}}$. Instead, we parameterize $Q_{\mathbf{w}}$ by its mean and KL divergence from $P_{\mathbf{w}}$ to constrain the compression cost to the desired value by construction. We demonstrate that variational training with Mean-KL parameterization converges twice as fast and maintains predictive performance after compression. Furthermore, we show that Mean-KL leads to more meaningful variational distributions with heavier tails and compressed weight samples which are more robust to pruning.
Online Laplace Model Selection Revisited
Jul 12, 2023



Abstract:The Laplace approximation provides a closed-form model selection objective for neural networks (NN). Online variants, which optimise NN parameters jointly with hyperparameters, like weight decay strength, have seen renewed interest in the Bayesian deep learning community. However, these methods violate Laplace's method's critical assumption that the approximation is performed around a mode of the loss, calling into question their soundness. This work re-derives online Laplace methods, showing them to target a variational bound on a mode-corrected variant of the Laplace evidence which does not make stationarity assumptions. Online Laplace and its mode-corrected counterpart share stationary points where 1. the NN parameters are a maximum a posteriori, satisfying the Laplace method's assumption, and 2. the hyperparameters maximise the Laplace evidence, motivating online methods. We demonstrate that these optima are roughly attained in practise by online algorithms using full-batch gradient descent on UCI regression datasets. The optimised hyperparameters prevent overfitting and outperform validation-based early stopping.
Leveraging Task Structures for Improved Identifiability in Neural Network Representations
Jun 26, 2023Abstract:This work extends the theory of identifiability in supervised learning by considering the consequences of having access to a distribution of tasks. In such cases, we show that identifiability is achievable even in the case of regression, extending prior work restricted to the single-task classification case. Furthermore, we show that the existence of a task distribution which defines a conditional prior over latent variables reduces the equivalence class for identifiability to permutations and scaling, a much stronger and more useful result. When we further assume a causal structure over these tasks, our approach enables simple maximum marginal likelihood optimization together with downstream applicability to causal representation learning. Empirically, we validate that our model outperforms more general unsupervised models in recovering canonical representations for synthetic and real-world data.
Tanimoto Random Features for Scalable Molecular Machine Learning
Jun 26, 2023Abstract:The Tanimoto coefficient is commonly used to measure the similarity between molecules represented as discrete fingerprints, either as a distance metric or a positive definite kernel. While many kernel methods can be accelerated using random feature approximations, at present there is a lack of such approximations for the Tanimoto kernel. In this paper we propose two kinds of novel random features to allow this kernel to scale to large datasets, and in the process discover a novel extension of the kernel to real vectors. We theoretically characterize these random features, and provide error bounds on the spectral norm of the Gram matrix. Experimentally, we show that the random features proposed in this work are effective at approximating the Tanimoto coefficient in real-world datasets and that the kernels explored in this work are useful for molecular property prediction and optimization tasks.
Sampling from Gaussian Process Posteriors using Stochastic Gradient Descent
Jun 20, 2023Abstract:Gaussian processes are a powerful framework for quantifying uncertainty and for sequential decision-making but are limited by the requirement of solving linear systems. In general, this has a cubic cost in dataset size and is sensitive to conditioning. We explore stochastic gradient algorithms as a computationally efficient method of approximately solving these linear systems: we develop low-variance optimization objectives for sampling from the posterior and extend these to inducing points. Counterintuitively, stochastic gradient descent often produces accurate predictions, even in cases where it does not converge quickly to the optimum. We explain this through a spectral characterization of the implicit bias from non-convergence. We show that stochastic gradient descent produces predictive distributions close to the true posterior both in regions with sufficient data coverage, and in regions sufficiently far away from the data. Experimentally, stochastic gradient descent achieves state-of-the-art performance on sufficiently large-scale or ill-conditioned regression tasks. Its uncertainty estimates match the performance of significantly more expensive baselines on a large-scale Bayesian~optimization~task.
Compression with Bayesian Implicit Neural Representations
May 30, 2023



Abstract:Many common types of data can be represented as functions that map coordinates to signal values, such as pixel locations to RGB values in the case of an image. Based on this view, data can be compressed by overfitting a compact neural network to its functional representation and then encoding the network weights. However, most current solutions for this are inefficient, as quantization to low-bit precision substantially degrades the reconstruction quality. To address this issue, we propose overfitting variational Bayesian neural networks to the data and compressing an approximate posterior weight sample using relative entropy coding instead of quantizing and entropy coding it. This strategy enables direct optimization of the rate-distortion performance by minimizing the $\beta$-ELBO, and target different rate-distortion trade-offs for a given network architecture by adjusting $\beta$. Moreover, we introduce an iterative algorithm for learning prior weight distributions and employ a progressive refinement process for the variational posterior that significantly enhances performance. Experiments show that our method achieves strong performance on image and audio compression while retaining simplicity.
Fast and Painless Image Reconstruction in Deep Image Prior Subspaces
Feb 20, 2023



Abstract:The deep image prior (DIP) is a state-of-the-art unsupervised approach for solving linear inverse problems in imaging. We address two key issues that have held back practical deployment of the DIP: the long computing time needed to train a separate deep network per reconstruction, and the susceptibility to overfitting due to a lack of robust early stopping strategies in the unsupervised setting. To this end, we restrict DIP optimisation to a sparse linear subspace of the full parameter space. We construct the subspace from the principal eigenspace of a set of parameter vectors sampled at equally spaced intervals during DIP pre-training on synthetic task-agnostic data. The low-dimensionality of the resulting subspace reduces DIP's capacity to fit noise and allows the use of fast second order optimisation methods, e.g., natural gradient descent or L-BFGS. Experiments across tomographic tasks of different geometry, ill-posedness and stopping criteria consistently show that second order optimisation in a subspace is Pareto-optimal in terms of optimisation time to reconstruction fidelity trade-off.
Sampling-based inference for large linear models, with application to linearised Laplace
Oct 10, 2022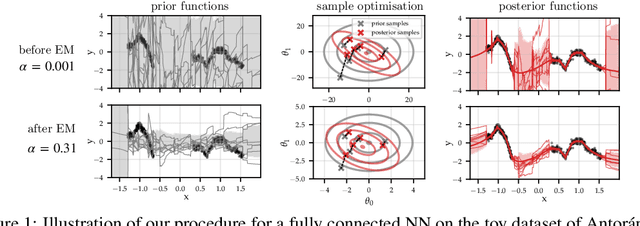
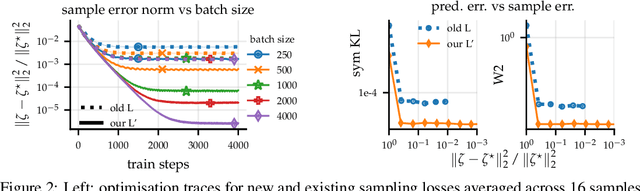
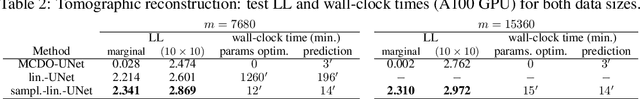
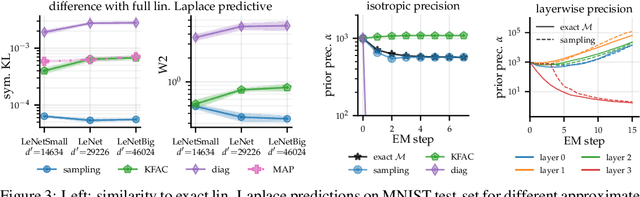
Abstract:Large-scale linear models are ubiquitous throughout machine learning, with contemporary application as surrogate models for neural network uncertainty quantification; that is, the linearised Laplace method. Alas, the computational cost associated with Bayesian linear models constrains this method's application to small networks, small output spaces and small datasets. We address this limitation by introducing a scalable sample-based Bayesian inference method for conjugate Gaussian multi-output linear models, together with a matching method for hyperparameter (regularisation) selection. Furthermore, we use a classic feature normalisation method (the g-prior) to resolve a previously highlighted pathology of the linearised Laplace method. Together, these contributions allow us to perform linearised neural network inference with ResNet-18 on CIFAR100 (11M parameters, 100 output dimensions x 50k datapoints) and with a U-Net on a high-resolution tomographic reconstruction task (2M parameters, 251k output dimensions).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge