John P. Cunningham
A Probabilistic Model of Cardiac Physiology and Electrocardiograms
Dec 01, 2018
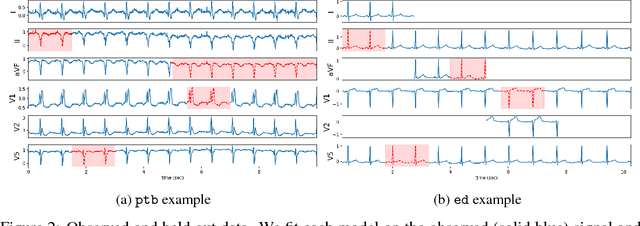
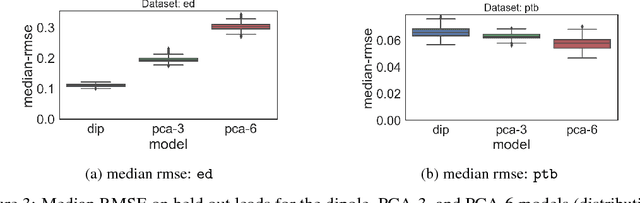
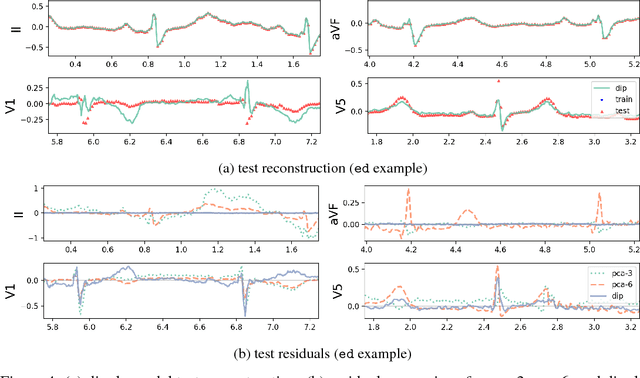
Abstract:An electrocardiogram (EKG) is a common, non-invasive test that measures the electrical activity of a patient's heart. EKGs contain useful diagnostic information about patient health that may be absent from other electronic health record (EHR) data. As multi-dimensional waveforms, they could be modeled using generic machine learning tools, such as a linear factor model or a variational autoencoder. We take a different approach:~we specify a model that directly represents the underlying electrophysiology of the heart and the EKG measurement process. We apply our model to two datasets, including a sample of emergency department EKG reports with missing data. We show that our model can more accurately reconstruct missing data (measured by test reconstruction error) than a standard baseline when there is significant missing data. More broadly, this physiological representation of heart function may be useful in a variety of settings, including prediction, causal analysis, and discovery.
Calibrating Deep Convolutional Gaussian Processes
May 26, 2018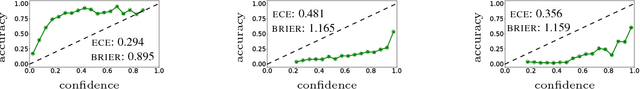
Abstract:The wide adoption of Convolutional Neural Networks (CNNs) in applications where decision-making under uncertainty is fundamental, has brought a great deal of attention to the ability of these models to accurately quantify the uncertainty in their predictions. Previous work on combining CNNs with Gaussian processes (GPs) has been developed under the assumption that the predictive probabilities of these models are well-calibrated. In this paper we show that, in fact, current combinations of CNNs and GPs are miscalibrated. We proposes a novel combination that considerably outperforms previous approaches on this aspect, while achieving state-of-the-art performance on image classification tasks.
Bayesian estimation for large scale multivariate Ornstein-Uhlenbeck model of brain connectivity
May 25, 2018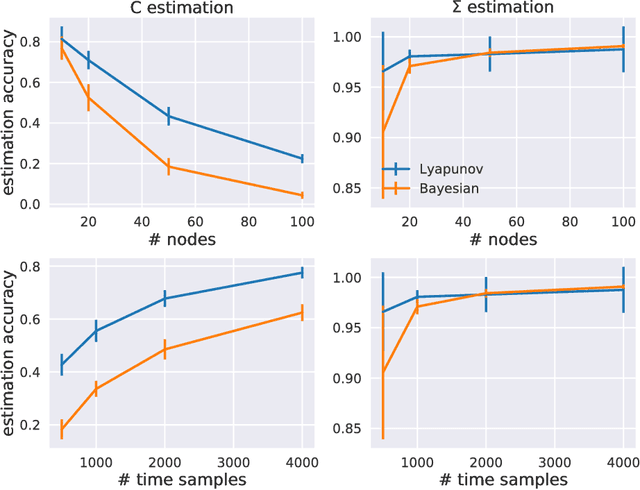
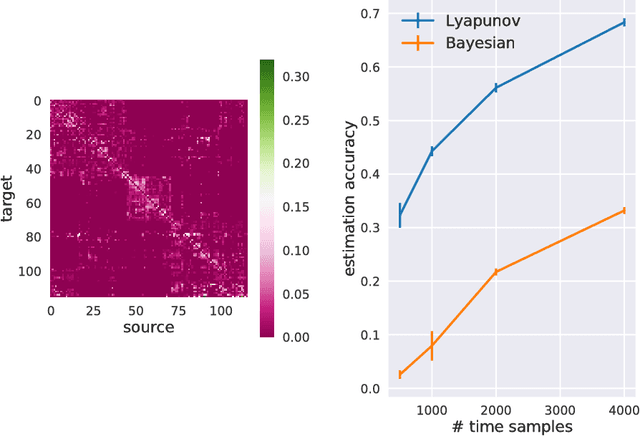
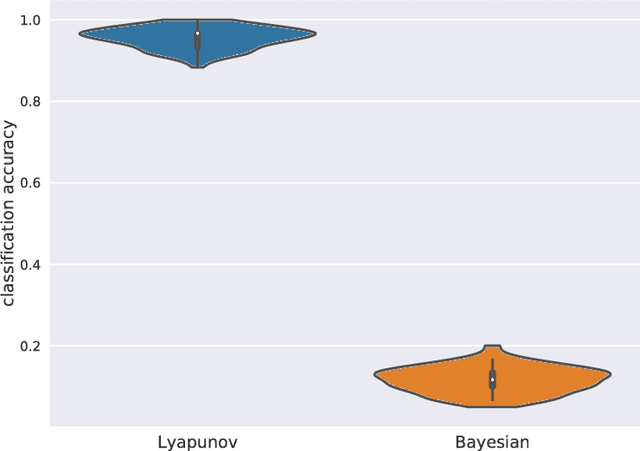
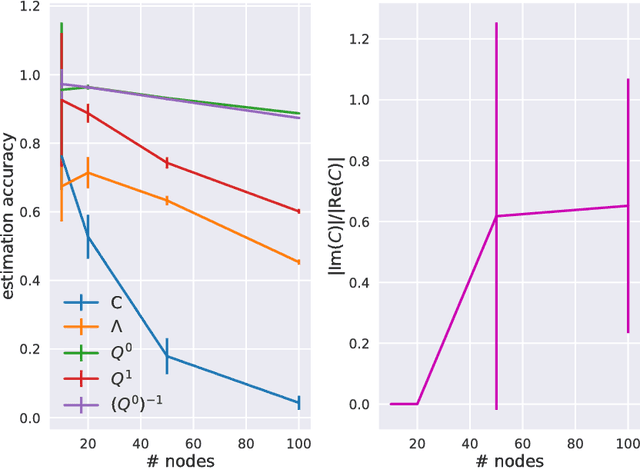
Abstract:Estimation of reliable whole-brain connectivity is a crucial step towards the use of connectivity information in quantitative approaches to the study of neuropsychiatric disorders. When estimating brain connectivity a challenge is imposed by the paucity of time samples and the large dimensionality of the measurements. Bayesian estimation methods for network models offer a number of advantages in this context but are not commonly employed. Here we compare three different estimation methods for the multivariate Ornstein-Uhlenbeck model, that has recently gained some popularity for characterizing whole-brain connectivity. We first show that a Bayesian estimation of model parameters assuming uniform priors is equivalent to an application of the method of moments. Then, using synthetic data, we show that the Bayesian estimate scales poorly with number of nodes in the network as compared to an iterative Lyapunov optimization. In particular when the network size is in the order of that used for whole-brain studies (about 100 nodes) the Bayesian method needs about eight times more time samples than Lyapunov method in order to achieve similar estimation accuracy. We also show that the higher estimation accuracy of Lyapunov method is reflected in a much better classification of individuals based on the estimated connectivity from a real dataset of BOLD fMRI. Finally we show that the poor accuracy of Bayesian method is due to numerical errors, when the imaginary part of the connectivity estimate gets large compared to its real part.
Expectation propagation as a way of life: A framework for Bayesian inference on partitioned data
Mar 10, 2018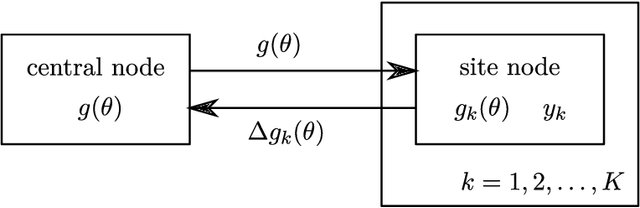
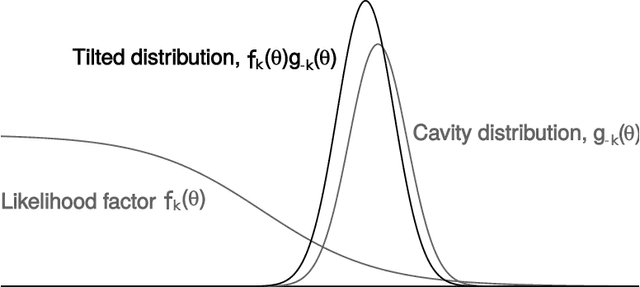
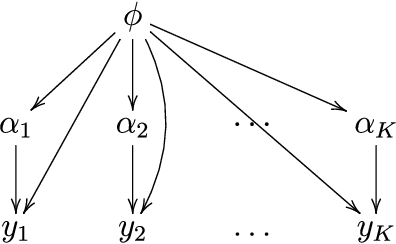
Abstract:A common approach for Bayesian computation with big data is to partition the data into smaller pieces, perform local inference for each piece separately, and finally combine the results to obtain an approximation to the global posterior. Looking at this from the bottom up, one can perform separate analyses on individual sources of data and then combine these in a larger Bayesian model. In either case, the idea of distributed modeling and inference has both conceptual and computational appeal, but from the Bayesian perspective there is no general way of handling the prior distribution: if the prior is included in each separate inference, it will be multiply-counted when the inferences are combined; but if the prior is itself divided into pieces, it may not provide enough regularization for each separate computation, thus eliminating one of the key advantages of Bayesian methods. To resolve this dilemma, we propose expectation propagation (EP) as a general prototype for distributed Bayesian inference. The central idea is to factor the likelihood according to the data partitions, and to iteratively combine each factor with an approximate model of the prior and all other parts of the data, thus producing an overall approximation to the global posterior at convergence. In this paper, we give an introduction to EP and an overview of some recent developments of the method, with particular emphasis on its use in combining inferences from partitioned data. In addition to distributed modeling of large datasets, our unified treatment also includes hierarchical modeling of data with a naturally partitioned structure. The paper describes a general algorithmic framework, rather than a specific algorithm, and presents an example implementation for it.
Reparameterizing the Birkhoff Polytope for Variational Permutation Inference
Oct 26, 2017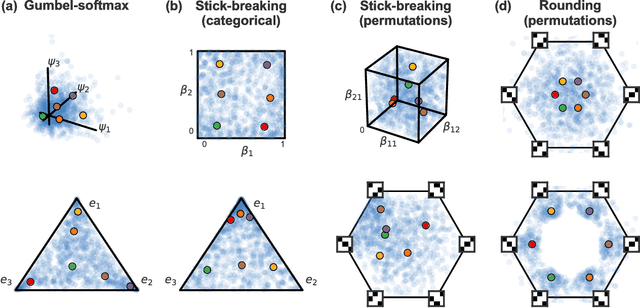
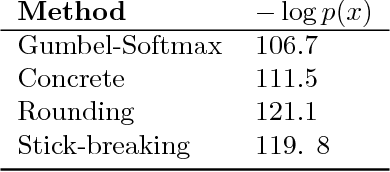
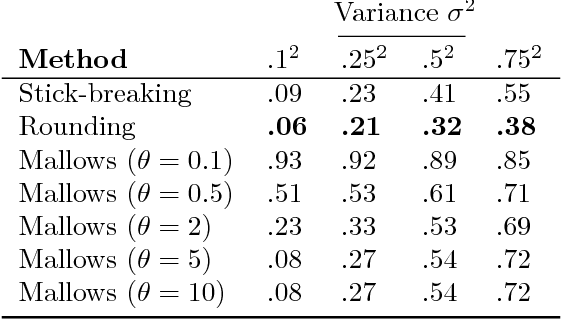
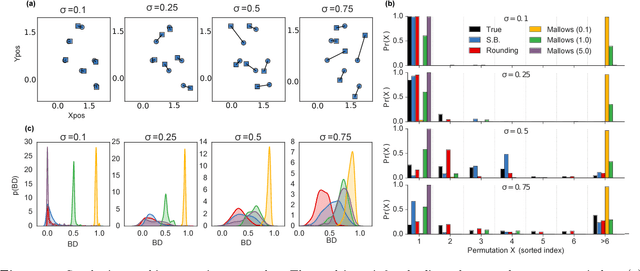

Abstract:Many matching, tracking, sorting, and ranking problems require probabilistic reasoning about possible permutations, a set that grows factorially with dimension. Combinatorial optimization algorithms may enable efficient point estimation, but fully Bayesian inference poses a severe challenge in this high-dimensional, discrete space. To surmount this challenge, we start with the usual step of relaxing a discrete set (here, of permutation matrices) to its convex hull, which here is the Birkhoff polytope: the set of all doubly-stochastic matrices. We then introduce two novel transformations: first, an invertible and differentiable stick-breaking procedure that maps unconstrained space to the Birkhoff polytope; second, a map that rounds points toward the vertices of the polytope. Both transformations include a temperature parameter that, in the limit, concentrates the densities on permutation matrices. We then exploit these transformations and reparameterization gradients to introduce variational inference over permutation matrices, and we demonstrate its utility in a series of experiments.
Sparse Probit Linear Mixed Model
Jul 17, 2017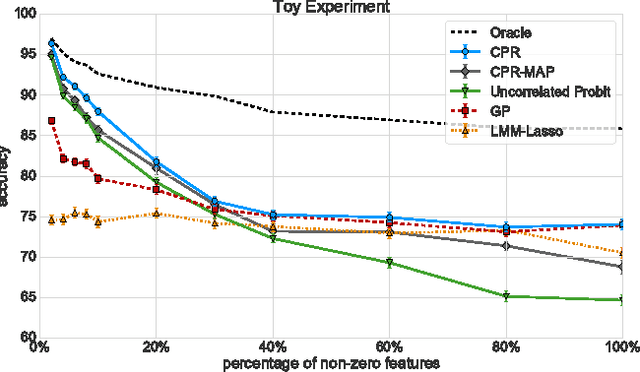
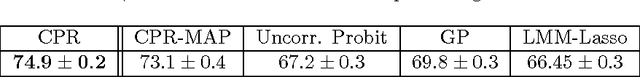
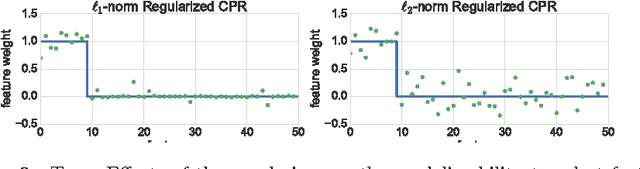
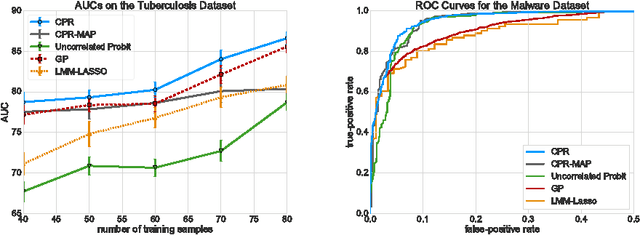
Abstract:Linear Mixed Models (LMMs) are important tools in statistical genetics. When used for feature selection, they allow to find a sparse set of genetic traits that best predict a continuous phenotype of interest, while simultaneously correcting for various confounding factors such as age, ethnicity and population structure. Formulated as models for linear regression, LMMs have been restricted to continuous phenotypes. We introduce the Sparse Probit Linear Mixed Model (Probit-LMM), where we generalize the LMM modeling paradigm to binary phenotypes. As a technical challenge, the model no longer possesses a closed-form likelihood function. In this paper, we present a scalable approximate inference algorithm that lets us fit the model to high-dimensional data sets. We show on three real-world examples from different domains that in the setup of binary labels, our algorithm leads to better prediction accuracies and also selects features which show less correlation with the confounding factors.
* Published version, 21 pages, 6 figures
Maximum Entropy Flow Networks
Apr 28, 2017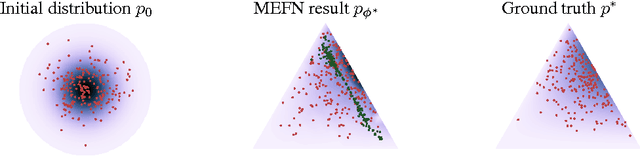
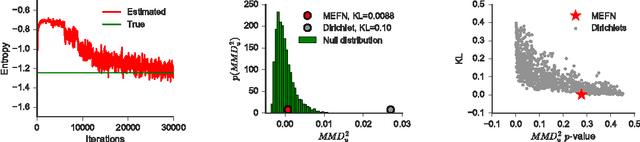
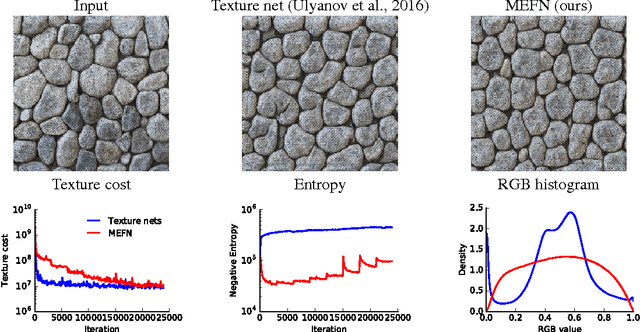
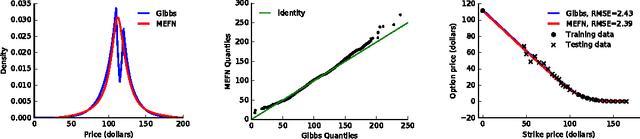
Abstract:Maximum entropy modeling is a flexible and popular framework for formulating statistical models given partial knowledge. In this paper, rather than the traditional method of optimizing over the continuous density directly, we learn a smooth and invertible transformation that maps a simple distribution to the desired maximum entropy distribution. Doing so is nontrivial in that the objective being maximized (entropy) is a function of the density itself. By exploiting recent developments in normalizing flow networks, we cast the maximum entropy problem into a finite-dimensional constrained optimization, and solve the problem by combining stochastic optimization with the augmented Lagrangian method. Simulation results demonstrate the effectiveness of our method, and applications to finance and computer vision show the flexibility and accuracy of using maximum entropy flow networks.
Linear dynamical neural population models through nonlinear embeddings
Oct 25, 2016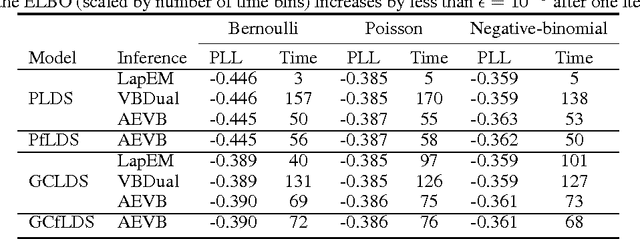
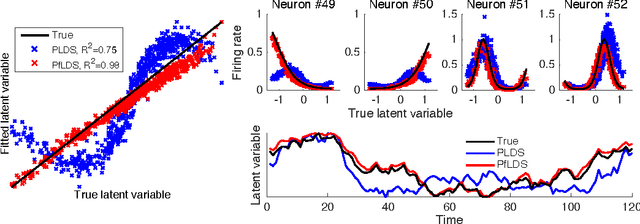
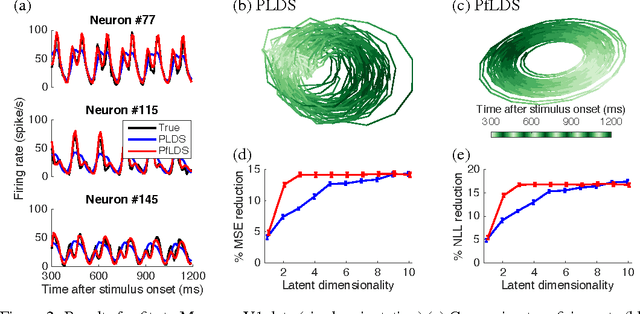
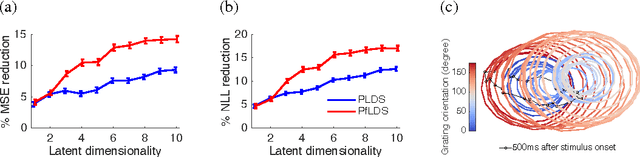
Abstract:A body of recent work in modeling neural activity focuses on recovering low-dimensional latent features that capture the statistical structure of large-scale neural populations. Most such approaches have focused on linear generative models, where inference is computationally tractable. Here, we propose fLDS, a general class of nonlinear generative models that permits the firing rate of each neuron to vary as an arbitrary smooth function of a latent, linear dynamical state. This extra flexibility allows the model to capture a richer set of neural variability than a purely linear model, but retains an easily visualizable low-dimensional latent space. To fit this class of non-conjugate models we propose a variational inference scheme, along with a novel approximate posterior capable of capturing rich temporal correlations across time. We show that our techniques permit inference in a wide class of generative models.We also show in application to two neural datasets that, compared to state-of-the-art neural population models, fLDS captures a much larger proportion of neural variability with a small number of latent dimensions, providing superior predictive performance and interpretability.
Bayesian Learning of Kernel Embeddings
Jun 02, 2016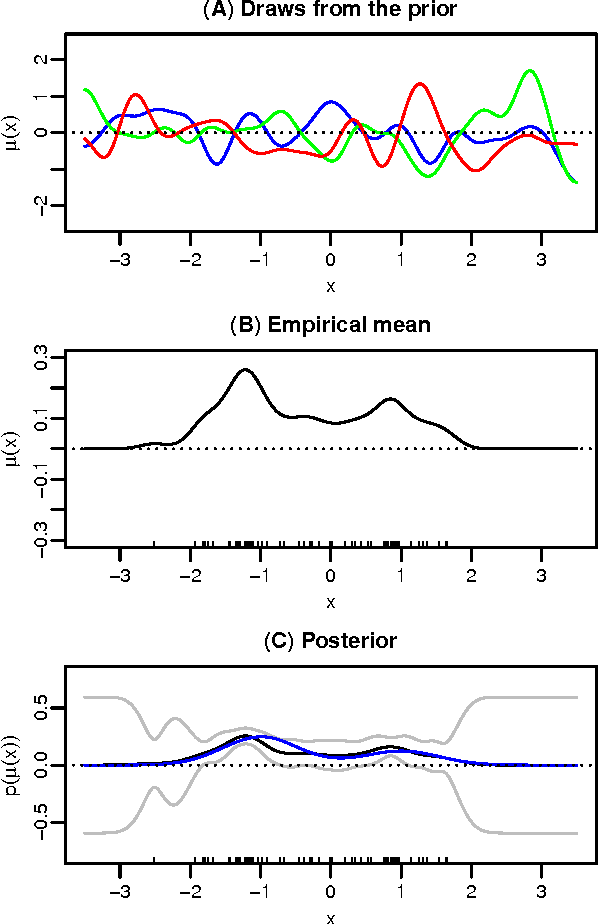
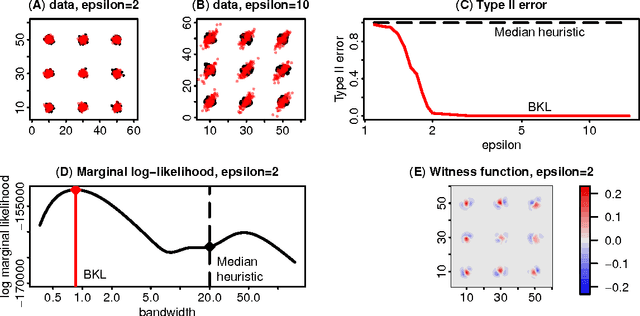
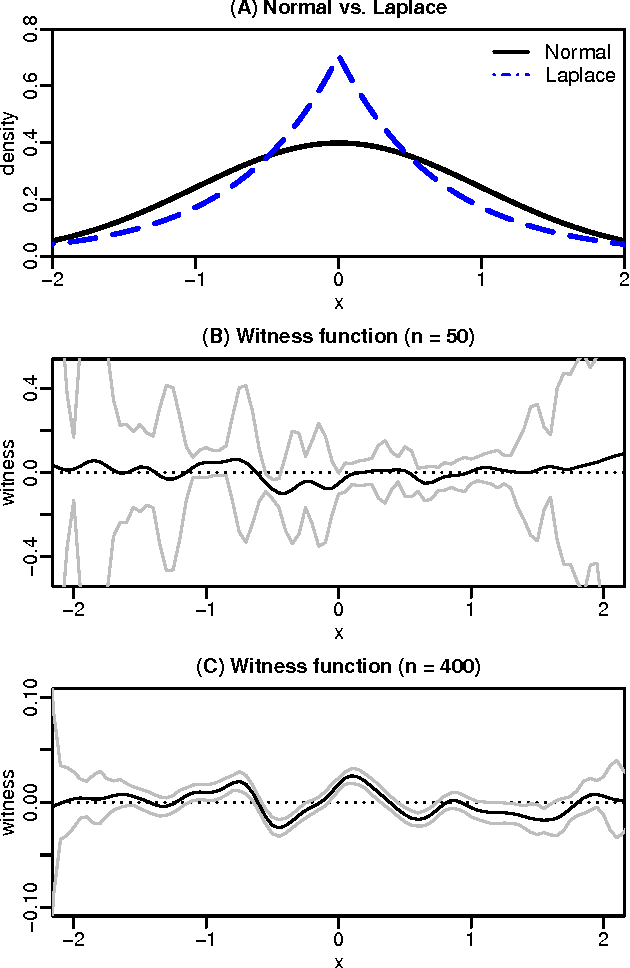
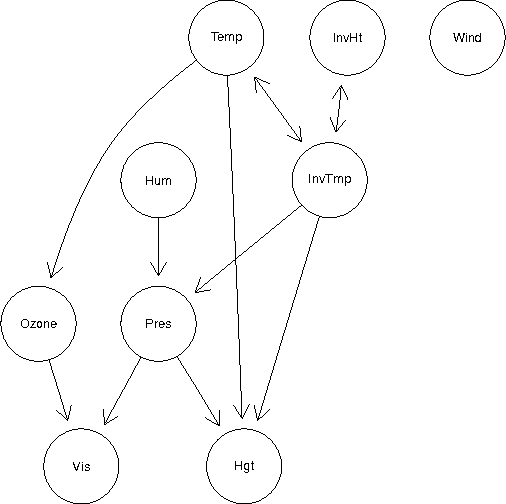
Abstract:Kernel methods are one of the mainstays of machine learning, but the problem of kernel learning remains challenging, with only a few heuristics and very little theory. This is of particular importance in methods based on estimation of kernel mean embeddings of probability measures. For characteristic kernels, which include most commonly used ones, the kernel mean embedding uniquely determines its probability measure, so it can be used to design a powerful statistical testing framework, which includes nonparametric two-sample and independence tests. In practice, however, the performance of these tests can be very sensitive to the choice of kernel and its lengthscale parameters. To address this central issue, we propose a new probabilistic model for kernel mean embeddings, the Bayesian Kernel Embedding model, combining a Gaussian process prior over the Reproducing Kernel Hilbert Space containing the mean embedding with a conjugate likelihood function, thus yielding a closed form posterior over the mean embedding. The posterior mean of our model is closely related to recently proposed shrinkage estimators for kernel mean embeddings, while the posterior uncertainty is a new, interesting feature with various possible applications. Critically for the purposes of kernel learning, our model gives a simple, closed form marginal pseudolikelihood of the observed data given the kernel hyperparameters. This marginal pseudolikelihood can either be optimized to inform the hyperparameter choice or fully Bayesian inference can be used.
Preconditioning Kernel Matrices
May 25, 2016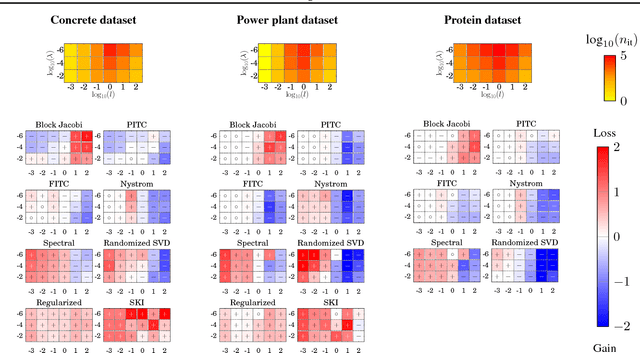
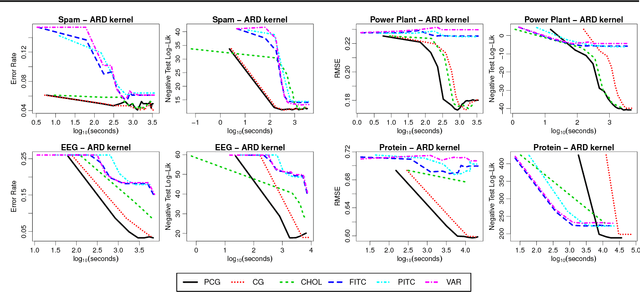
Abstract:The computational and storage complexity of kernel machines presents the primary barrier to their scaling to large, modern, datasets. A common way to tackle the scalability issue is to use the conjugate gradient algorithm, which relieves the constraints on both storage (the kernel matrix need not be stored) and computation (both stochastic gradients and parallelization can be used). Even so, conjugate gradient is not without its own issues: the conditioning of kernel matrices is often such that conjugate gradients will have poor convergence in practice. Preconditioning is a common approach to alleviating this issue. Here we propose preconditioned conjugate gradients for kernel machines, and develop a broad range of preconditioners particularly useful for kernel matrices. We describe a scalable approach to both solving kernel machines and learning their hyperparameters. We show this approach is exact in the limit of iterations and outperforms state-of-the-art approximations for a given computational budget.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge