Joao Ramalhinho
HyKey: Hyperspectral Keypoint Detection and Matching in Minimally Invasive Surgery
Apr 19, 2026Abstract:Purpose: 3D reconstruction in minimally invasive surgery (MIS) enables enhanced surgical guidance through improved visualisation, tool tracking, and augmented reality. However, traditional RGB-based keypoint detection and matching pipelines struggle with surgical challenges, such as poor texture and complex illumination. We investigate whether using snapshot hyperspectral imaging (HSI) can provide improved results on keypoint detection and matching surgical scenes. Methods: We developed HyKey, a HYperspectral KEYpoint detection and description model made up of a hybrid 3D-2D convolutional neural network that jointly extracts spatial-spectral features from HSI. The model was trained using synthetic homographic augmentation and epipolar geometry constraints on a robotically-acquired dual-camera RGB-HSI laparoscopic dataset of ex-vivo organs with calibrated camera poses. We benchmarked performance against established RGB-based methods, including SuperPoint and ALIKE. Results: Our HSI-based model outperformed RGB baselines on registered RGB frames, achieving 96.62% mean matching accuracy and 67.18% mean average accuracy at 10 degree on pose estimation, demonstrating consistent improvements across multiple evaluation metrics. Conclusion: Integrating spectral information from an HSI cube offers a promising approach for robust monocular 3D reconstruction in MIS, addressing limitations of texture-poor surgical environments through enhanced spectral-spatial feature discrimination. Our model and dataset are available at https://github.com/alexsaikia/HyKey-Hyperspectral-Keypoint-Detection
Intraoperative Liver Surface Completion with Graph Convolutional VAE
Sep 08, 2020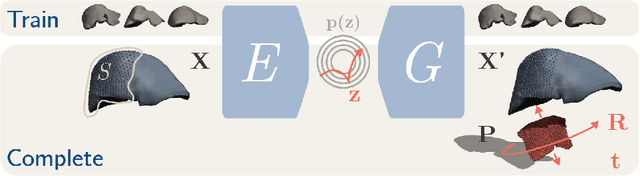
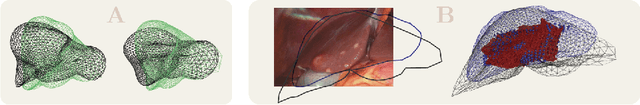
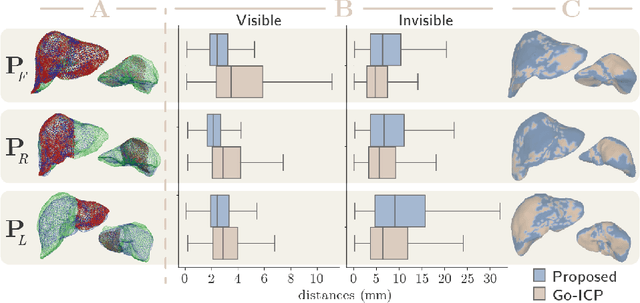
Abstract:In this work we propose a method based on geometric deep learning to predict the complete surface of the liver, given a partial point cloud of the organ obtained during the surgical laparoscopic procedure. We introduce a new data augmentation technique that randomly perturbs shapes in their frequency domain to compensate the limited size of our dataset. The core of our method is a variational autoencoder (VAE) that is trained to learn a latent space for complete shapes of the liver. At inference time, the generative part of the model is embedded in an optimisation procedure where the latent representation is iteratively updated to generate a model that matches the intraoperative partial point cloud. The effect of this optimisation is a progressive non-rigid deformation of the initially generated shape. Our method is qualitatively evaluated on real data and quantitatively evaluated on synthetic data. We compared with a state-of-the-art rigid registration algorithm, that our method outperformed in visible areas.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge