Jin Hong
CausalDisenSeg: A Causality-Guided Disentanglement Framework with Counterfactual Reasoning for Robust Brain Tumor Segmentation Under Missing Modalities
Apr 15, 2026Abstract:In clinical practice, the robustness of deep learning models for multimodal brain tumor segmentation is severely compromised by incomplete MRI data. This vulnerability stems primarily from modality bias, where models exploit spurious correlations as shortcuts rather than learning true anatomical structures. Existing feature fusion methods fail to fundamentally eliminate this dependency. To address this, we propose CausalDisenSeg, a novel Structural Causal Model (SCM)-grounded framework that achieves robust segmentation via causality-guided disentanglement and counterfactual reasoning. We reframe the problem as isolating the anatomical Causal Factor from the stylistic Bias Factor. Our framework implements a three-stage causal intervention: (1) Explicit Causal Disentanglement: A Conditional Variational Autoencoder (CVAE) coupled with an HSIC constraint mathematically enforces statistical orthogonality between anatomical and style features. (2) Causal Representation Reinforcement: A Region Causality Module (RCM) explicitly grounds causal features in physical tumor regions. (3) Counterfactual Reasoning: A dual-adversarial strategy actively suppresses the residual Natural Direct Effect (NDE) of the bias, forcing its spatial attention to be mutually exclusive from the causal path. Extensive experiments on the BraTS 2020 dataset demonstrate that CausalDisenSeg significantly outperforms state-of-the-art methods in accuracy and consistency across severe missing-modality scenarios. Furthermore, cross-dataset evaluation on BraTS 2023 under the same protocol yields a state-of-the-art macro-average DSC of 84.49.
MGML: A Plug-and-Play Meta-Guided Multi-Modal Learning Framework for Incomplete Multimodal Brain Tumor Segmentation
Dec 30, 2025Abstract:Leveraging multimodal information from Magnetic Resonance Imaging (MRI) plays a vital role in lesion segmentation, especially for brain tumors. However, in clinical practice, multimodal MRI data are often incomplete, making it challenging to fully utilize the available information. Therefore, maximizing the utilization of this incomplete multimodal information presents a crucial research challenge. We present a novel meta-guided multi-modal learning (MGML) framework that comprises two components: meta-parameterized adaptive modality fusion and consistency regularization module. The meta-parameterized adaptive modality fusion (Meta-AMF) enables the model to effectively integrate information from multiple modalities under varying input conditions. By generating adaptive soft-label supervision signals based on the available modalities, Meta-AMF explicitly promotes more coherent multimodal fusion. In addition, the consistency regularization module enhances segmentation performance and implicitly reinforces the robustness and generalization of the overall framework. Notably, our approach does not alter the original model architecture and can be conveniently integrated into the training pipeline for end-to-end model optimization. We conducted extensive experiments on the public BraTS2020 and BraTS2023 datasets. Compared to multiple state-of-the-art methods from previous years, our method achieved superior performance. On BraTS2020, for the average Dice scores across fifteen missing modality combinations, building upon the baseline, our method obtained scores of 87.55, 79.36, and 62.67 for the whole tumor (WT), the tumor core (TC), and the enhancing tumor (ET), respectively. We have made our source code publicly available at https://github.com/worldlikerr/MGML.
ERANet: Edge Replacement Augmentation for Semi-Supervised Meniscus Segmentation with Prototype Consistency Alignment and Conditional Self-Training
Feb 11, 2025Abstract:Manual segmentation is labor-intensive, and automatic segmentation remains challenging due to the inherent variability in meniscal morphology, partial volume effects, and low contrast between the meniscus and surrounding tissues. To address these challenges, we propose ERANet, an innovative semi-supervised framework for meniscus segmentation that effectively leverages both labeled and unlabeled images through advanced augmentation and learning strategies. ERANet integrates three key components: edge replacement augmentation (ERA), prototype consistency alignment (PCA), and a conditional self-training (CST) strategy within a mean teacher architecture. ERA introduces anatomically relevant perturbations by simulating meniscal variations, ensuring that augmentations align with the structural context. PCA enhances segmentation performance by aligning intra-class features and promoting compact, discriminative feature representations, particularly in scenarios with limited labeled data. CST improves segmentation robustness by iteratively refining pseudo-labels and mitigating the impact of label noise during training. Together, these innovations establish ERANet as a robust and scalable solution for meniscus segmentation, effectively addressing key barriers to practical implementation. We validated ERANet comprehensively on 3D Double Echo Steady State (DESS) and 3D Fast/Turbo Spin Echo (FSE/TSE) MRI sequences. The results demonstrate the superior performance of ERANet compared to state-of-the-art methods. The proposed framework achieves reliable and accurate segmentation of meniscus structures, even when trained on minimal labeled data. Extensive ablation studies further highlight the synergistic contributions of ERA, PCA, and CST, solidifying ERANet as a transformative solution for semi-supervised meniscus segmentation in medical imaging.
WaveNet-SF: A Hybrid Network for Retinal Disease Detection Based on Wavelet Transform in the Spatial-Frequency Domain
Jan 21, 2025



Abstract:Retinal diseases are a leading cause of vision impairment and blindness, with timely diagnosis being critical for effective treatment. Optical Coherence Tomography (OCT) has become a standard imaging modality for retinal disease diagnosis, but OCT images often suffer from issues such as speckle noise, complex lesion shapes, and varying lesion sizes, making interpretation challenging. In this paper, we propose a novel framework, WaveNet-SF, to enhance retinal disease detection by integrating spatial-domain and frequency-domain learning. The framework utilizes wavelet transforms to decompose OCT images into low- and high-frequency components, enabling the model to extract both global structural features and fine-grained details. To improve lesion detection, we introduce a multi-scale wavelet spatial attention (MSW-SA) module, which enhances the model's focus on regions of interest at multiple scales. Additionally, a high-frequency feature compensation block (HFFC) is incorporated to recover edge information lost during wavelet decomposition, suppress noise, and preserve fine details crucial for lesion detection. Our approach achieves state-of-the-art (SOTA) classification accuracies of 97.82% and 99. 58% on the OCT-C8 and OCT2017 datasets, respectively, surpassing existing methods. These results demonstrate the efficacy of WaveNet-SF in addressing the challenges of OCT image analysis and its potential as a powerful tool for retinal disease diagnosis.
Rethinking domain generalization in medical image segmentation: One image as one domain
Jan 08, 2025Abstract:Domain shifts in medical image segmentation, particularly when data comes from different centers, pose significant challenges. Intra-center variability, such as differences in scanner models or imaging protocols, can cause domain shifts as large as, or even larger than, those between centers. To address this, we propose the "one image as one domain" (OIOD) hypothesis, which treats each image as a unique domain, enabling flexible and robust domain generalization. Based on this hypothesis, we develop a unified disentanglement-based domain generalization (UniDDG) framework, which simultaneously handles both multi-source and single-source domain generalization without requiring explicit domain labels. This approach simplifies training with a fixed architecture, independent of the number of source domains, reducing complexity and enhancing scalability. We decouple each input image into content representation and style code, then exchange and combine these within the batch for segmentation, reconstruction, and further disentanglement. By maintaining distinct style codes for each image, our model ensures thorough decoupling of content representations and style codes, improving domain invariance of the content representations. Additionally, we enhance generalization with expansion mask attention (EMA) for boundary preservation and style augmentation (SA) to simulate diverse image styles, improving robustness to domain shifts. Extensive experiments show that our method achieves Dice scores of 84.43% and 88.91% for multi-source to single-center and single-center generalization in optic disc and optic cup segmentation, respectively, and 86.96% and 88.56% for prostate segmentation, outperforming current state-of-the-art domain generalization methods, offering superior performance and adaptability across clinical settings.
DGSSA: Domain generalization with structural and stylistic augmentation for retinal vessel segmentation
Jan 07, 2025



Abstract:Retinal vascular morphology is crucial for diagnosing diseases such as diabetes, glaucoma, and hypertension, making accurate segmentation of retinal vessels essential for early intervention. Traditional segmentation methods assume that training and testing data share similar distributions, which can lead to poor performance on unseen domains due to domain shifts caused by variations in imaging devices and patient demographics. This paper presents a novel approach, DGSSA, for retinal vessel image segmentation that enhances model generalization by combining structural and style augmentation strategies. We utilize a space colonization algorithm to generate diverse vascular-like structures that closely mimic actual retinal vessels, which are then used to generate pseudo-retinal images with an improved Pix2Pix model, allowing the segmentation model to learn a broader range of structure distributions. Additionally, we utilize PixMix to implement random photometric augmentations and introduce uncertainty perturbations, thereby enriching stylistic diversity and significantly enhancing the model's adaptability to varying imaging conditions. Our framework has been rigorously evaluated on four challenging datasets-DRIVE, CHASEDB, HRF, and STARE-demonstrating state-of-the-art performance that surpasses existing methods. This validates the effectiveness of our proposed approach, highlighting its potential for clinical application in automated retinal vessel analysis.
Embodied AI in Mobile Robots: Coverage Path Planning with Large Language Models
Jul 02, 2024



Abstract:In recent years, Large Language Models (LLMs) have demonstrated remarkable capabilities in understanding and solving mathematical problems, leading to advancements in various fields. We propose an LLM-embodied path planning framework for mobile agents, focusing on solving high-level coverage path planning issues and low-level control. Our proposed multi-layer architecture uses prompted LLMs in the path planning phase and integrates them with the mobile agents' low-level actuators. To evaluate the performance of various LLMs, we propose a coverage-weighted path planning metric to assess the performance of the embodied models. Our experiments show that the proposed framework improves LLMs' spatial inference abilities. We demonstrate that the proposed multi-layer framework significantly enhances the efficiency and accuracy of these tasks by leveraging the natural language understanding and generative capabilities of LLMs. Our experiments show that this framework can improve LLMs' 2D plane reasoning abilities and complete coverage path planning tasks. We also tested three LLM kernels: gpt-4o, gemini-1.5-flash, and claude-3.5-sonnet. The experimental results show that claude-3.5 can complete the coverage planning task in different scenarios, and its indicators are better than those of the other models.
Quantum-Inspired Machine Learning: a Survey
Sep 08, 2023



Abstract:Quantum-inspired Machine Learning (QiML) is a burgeoning field, receiving global attention from researchers for its potential to leverage principles of quantum mechanics within classical computational frameworks. However, current review literature often presents a superficial exploration of QiML, focusing instead on the broader Quantum Machine Learning (QML) field. In response to this gap, this survey provides an integrated and comprehensive examination of QiML, exploring QiML's diverse research domains including tensor network simulations, dequantized algorithms, and others, showcasing recent advancements, practical applications, and illuminating potential future research avenues. Further, a concrete definition of QiML is established by analyzing various prior interpretations of the term and their inherent ambiguities. As QiML continues to evolve, we anticipate a wealth of future developments drawing from quantum mechanics, quantum computing, and classical machine learning, enriching the field further. This survey serves as a guide for researchers and practitioners alike, providing a holistic understanding of QiML's current landscape and future directions.
Multi-resolution Spatiotemporal Enhanced Transformer Denoising with Functional Diffusive GANs for Constructing Brain Effective Connectivity in MCI analysis
May 18, 2023



Abstract:Effective connectivity can describe the causal patterns among brain regions. These patterns have the potential to reveal the pathological mechanism and promote early diagnosis and effective drug development for cognitive disease. However, the current studies mainly focus on using empirical functional time series to calculate effective connections, which may not comprehensively capture the complex causal relationships between brain regions. In this paper, a novel Multi-resolution Spatiotemporal Enhanced Transformer Denoising (MSETD) network with an adversarially functional diffusion model is proposed to map functional magnetic resonance imaging (fMRI) into effective connectivity for mild cognitive impairment (MCI) analysis. To be specific, the denoising framework leverages a conditional diffusion process that progressively translates the noise and conditioning fMRI to effective connectivity in an end-to-end manner. To ensure reverse diffusion quality and diversity, the multi-resolution enhanced transformer generator is designed to extract local and global spatiotemporal features. Furthermore, a multi-scale diffusive transformer discriminator is devised to capture the temporal patterns at different scales for generation stability. Evaluations of the ADNI datasets demonstrate the feasibility and efficacy of the proposed model. The proposed model not only achieves superior prediction performance compared with other competing methods but also identifies MCI-related causal connections that are consistent with clinical studies.
Source-free unsupervised domain adaptation for cross-modality abdominal multi-organ segmentation
Nov 30, 2021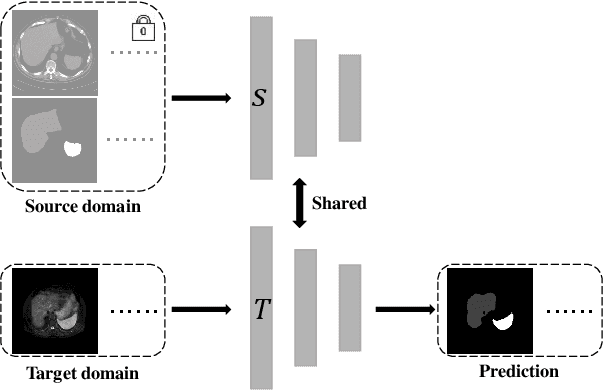
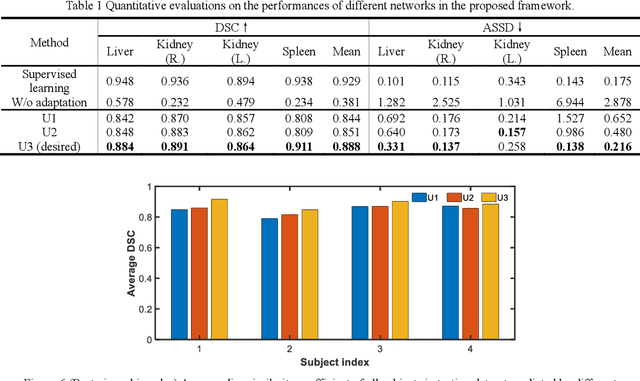
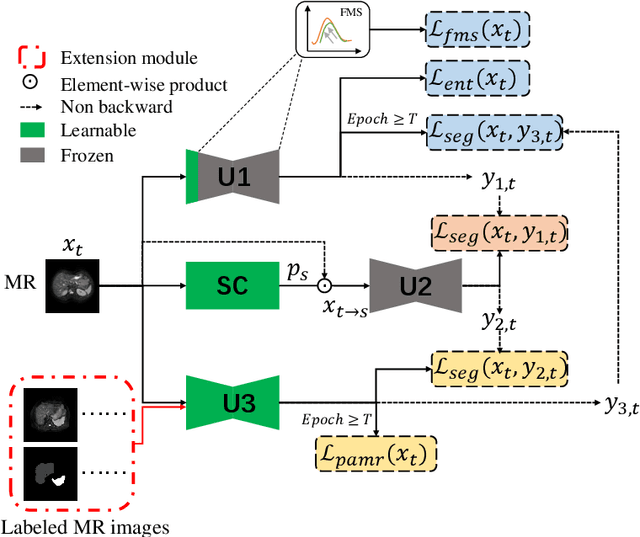
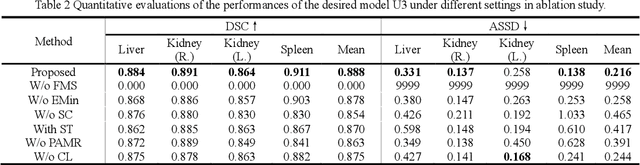
Abstract:It is valuable to achieve domain adaptation to transfer the learned knowledge from the source labeled CT dataset to the target unlabeled MR dataset for abdominal multi-organ segmentation. Meanwhile, it is highly desirable to avoid high annotation cost of target dataset and protect privacy of source dataset. Therefore, we propose an effective source-free unsupervised domain adaptation method for cross-modality abdominal multi-organ segmentation without accessing the source dataset. The process of the proposed framework includes two stages. At the first stage, the feature map statistics loss is used to align the distributions of the source and target features in the top segmentation network, and entropy minimization loss is used to encourage high confidence segmentations. The pseudo-labels outputted from the top segmentation network is used to guide the style compensation network to generate source-like images. The pseudo-labels outputted from the middle segmentation network is used to supervise the learning of the desired model (the bottom segmentation network). At the second stage, the circular learning and the pixel-adaptive mask refinement are used to further improve the performance of the desired model. With this approach, we achieve satisfactory performances on the segmentations of liver, right kidney, left kidney, and spleen with the dice similarity coefficients of 0.884, 0.891, 0.864, and 0.911, respectively. In addition, the proposed approach can be easily extended to the situation when there exists target annotation data. The performance improves from 0.888 to 0.922 in average dice similarity coefficient, close to the supervised learning (0.929), with only one labeled MR volume.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge