Jens Petersen
RootPainter3D: Interactive-machine-learning enables rapid and accurate contouring for radiotherapy
Jun 22, 2021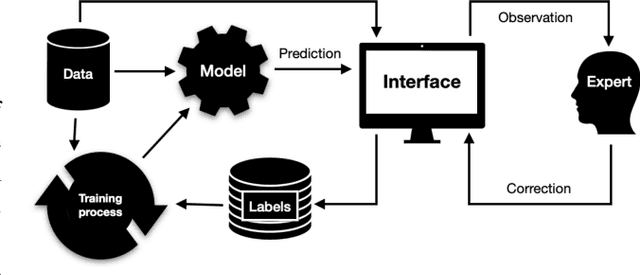
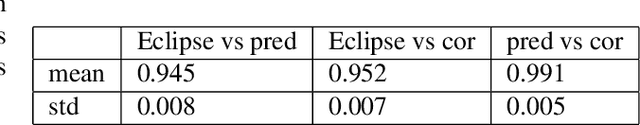
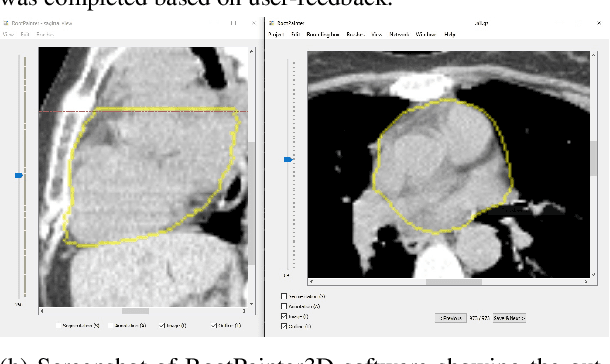
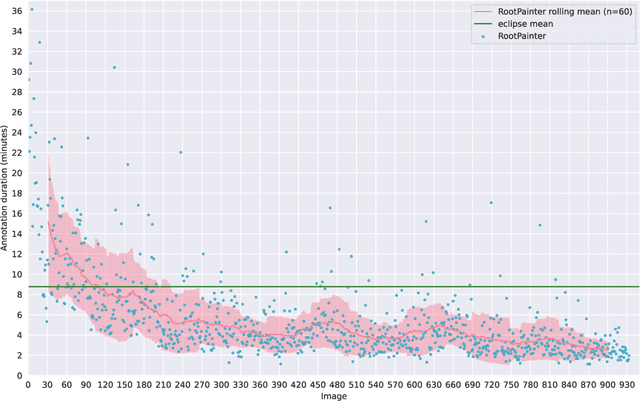
Abstract:Organ-at-risk contouring is still a bottleneck in radiotherapy, with many deep learning methods falling short of promised results when evaluated on clinical data. We investigate the accuracy and time-savings resulting from the use of an interactive-machine-learning method for an organ-at-risk contouring task. We compare the method to the Eclipse contouring software and find strong agreement with manual delineations, with a dice score of 0.95. The annotations created using corrective-annotation also take less time to create as more images are annotated, resulting in substantial time savings compared to manual methods, with hearts that take 2 minutes and 2 seconds to delineate on average, after 923 images have been delineated, compared to 7 minutes and 1 seconds when delineating manually. Our experiment demonstrates that interactive-machine-learning with corrective-annotation provides a fast and accessible way for non computer-scientists to train deep-learning models to segment their own structures of interest as part of routine clinical workflows. Source code is available at \href{https://github.com/Abe404/RootPainter3D}{this HTTPS URL}.
GP-ConvCNP: Better Generalization for Convolutional Conditional Neural Processes on Time Series Data
Jun 11, 2021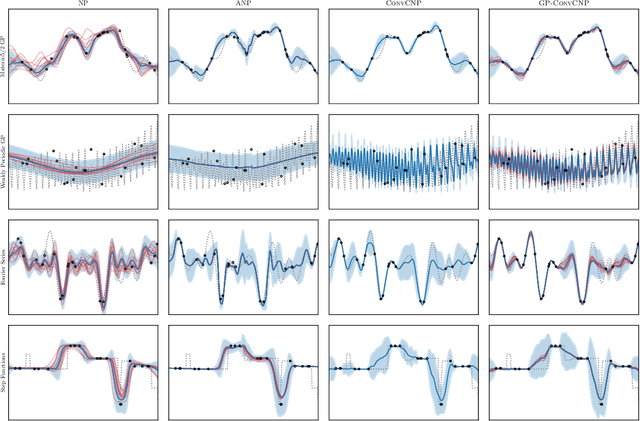
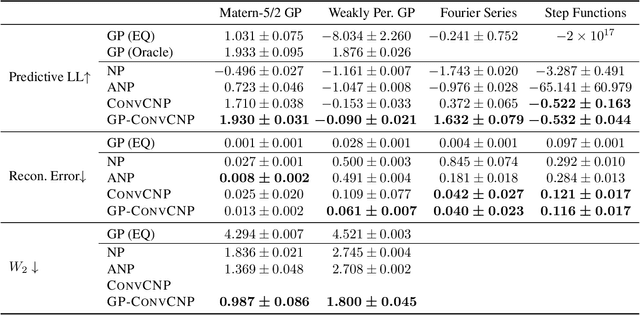
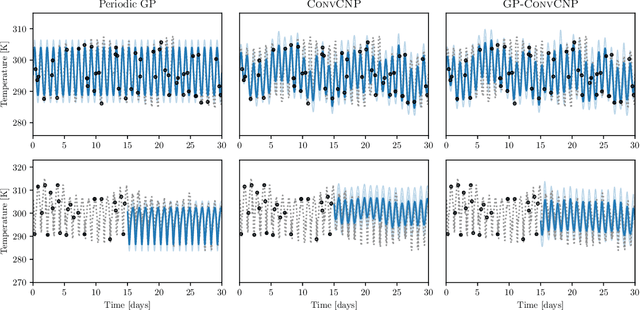
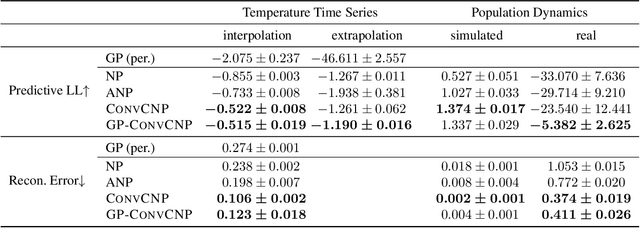
Abstract:Neural Processes (NPs) are a family of conditional generative models that are able to model a distribution over functions, in a way that allows them to perform predictions at test time conditioned on a number of context points. A recent addition to this family, Convolutional Conditional Neural Processes (ConvCNP), have shown remarkable improvement in performance over prior art, but we find that they sometimes struggle to generalize when applied to time series data. In particular, they are not robust to distribution shifts and fail to extrapolate observed patterns into the future. By incorporating a Gaussian Process into the model, we are able to remedy this and at the same time improve performance within distribution. As an added benefit, the Gaussian Process reintroduces the possibility to sample from the model, a key feature of other members in the NP family.
Common Limitations of Image Processing Metrics: A Picture Story
Apr 13, 2021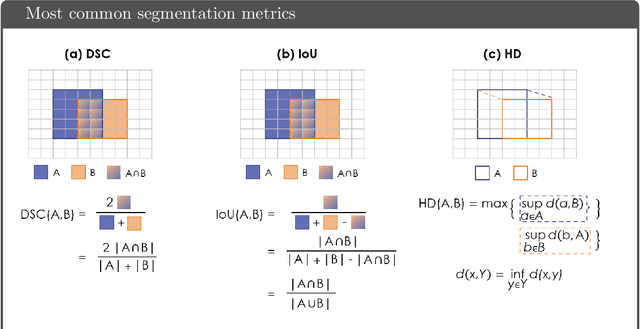
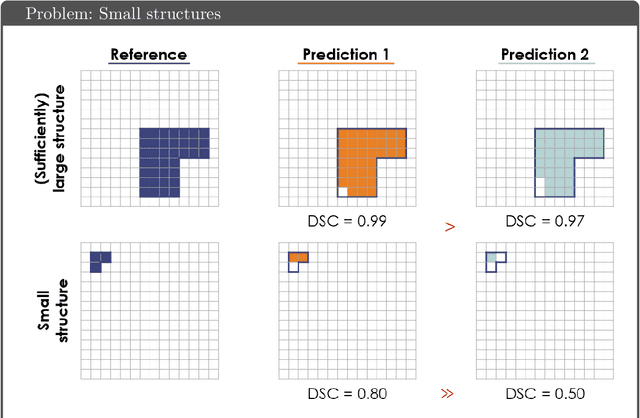
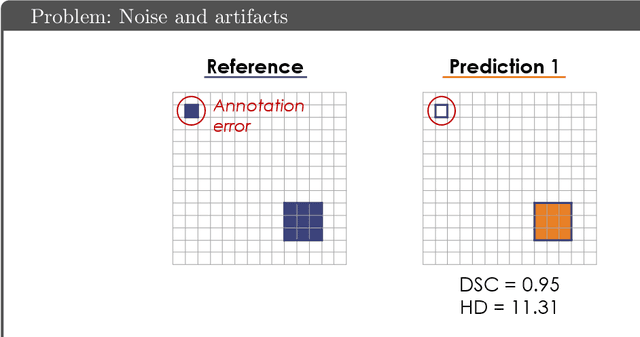
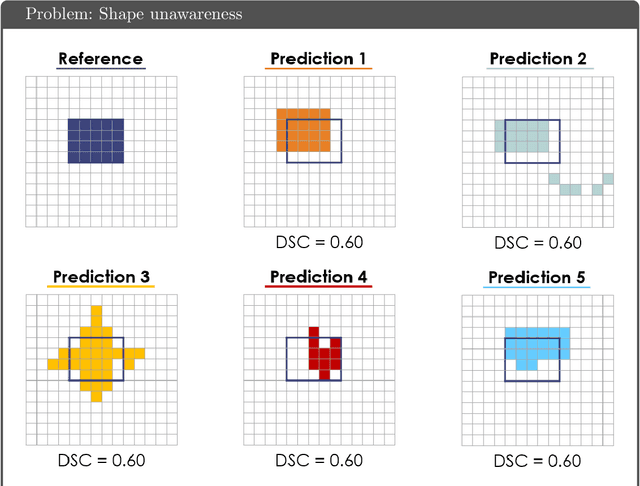
Abstract:While the importance of automatic image analysis is increasing at an enormous pace, recent meta-research revealed major flaws with respect to algorithm validation. Specifically, performance metrics are key for objective, transparent and comparative performance assessment, but relatively little attention has been given to the practical pitfalls when using specific metrics for a given image analysis task. A common mission of several international initiatives is therefore to provide researchers with guidelines and tools to choose the performance metrics in a problem-aware manner. This dynamically updated document has the purpose to illustrate important limitations of performance metrics commonly applied in the field of image analysis. The current version is based on a Delphi process on metrics conducted by an international consortium of image analysis experts.
Segmenting two-dimensional structures with strided tensor networks
Feb 13, 2021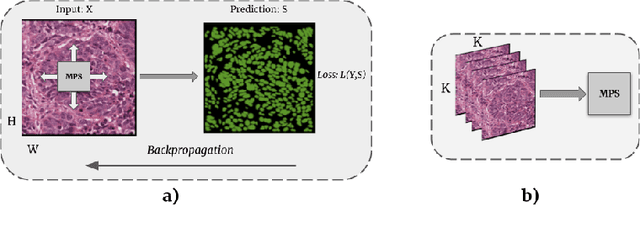
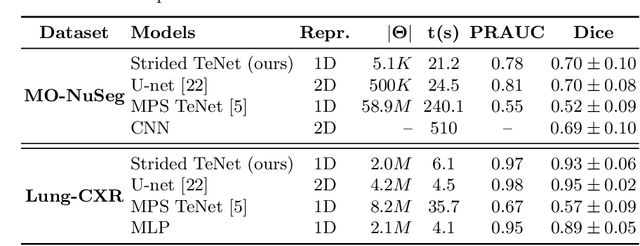
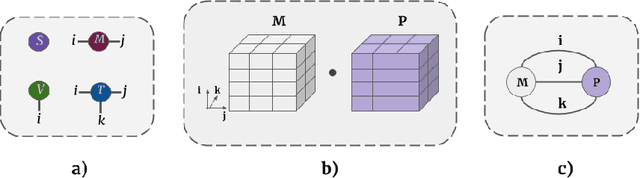
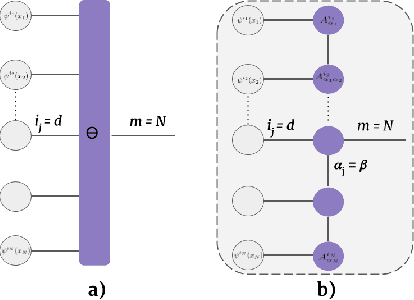
Abstract:Tensor networks provide an efficient approximation of operations involving high dimensional tensors and have been extensively used in modelling quantum many-body systems. More recently, supervised learning has been attempted with tensor networks, primarily focused on tasks such as image classification. In this work, we propose a novel formulation of tensor networks for supervised image segmentation which allows them to operate on high resolution medical images. We use the matrix product state (MPS) tensor network on non-overlapping patches of a given input image to predict the segmentation mask by learning a pixel-wise linear classification rule in a high dimensional space. The proposed model is end-to-end trainable using backpropagation. It is implemented as a Strided Tensor Network to reduce the parameter complexity. The performance of the proposed method is evaluated on two public medical imaging datasets and compared to relevant baselines. The evaluation shows that the strided tensor network yields competitive performance compared to CNN-based models while using fewer resources. Additionally, based on the experiments we discuss the feasibility of using fully linear models for segmentation tasks.
A Case for the Score: Identifying Image Anomalies using Variational Autoencoder Gradients
Nov 28, 2019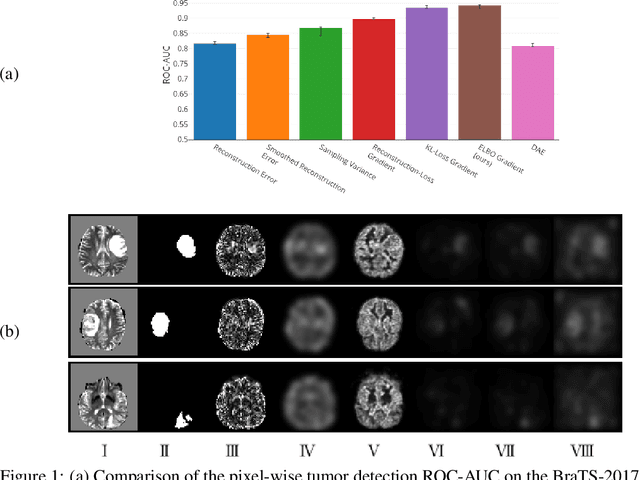
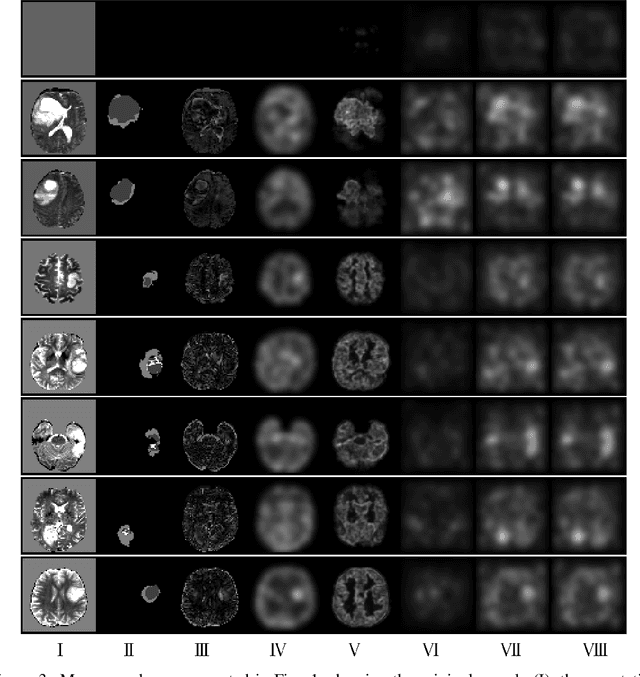
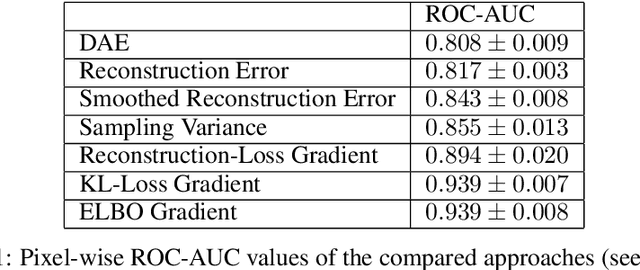
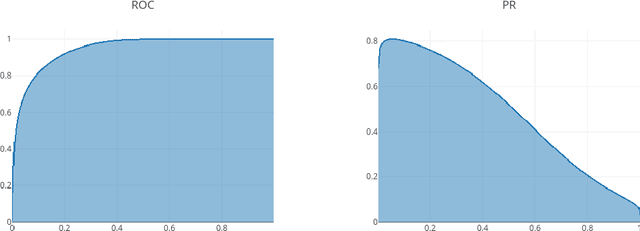

Abstract:Through training on unlabeled data, anomaly detection has the potential to impact computer-aided diagnosis by outlining suspicious regions. Previous work on deep-learning-based anomaly detection has primarily focused on the reconstruction error. We argue instead, that pixel-wise anomaly ratings derived from a Variational Autoencoder based score approximation yield a theoretically better grounded and more faithful estimate. In our experiments, Variational Autoencoder gradient-based rating outperforms other approaches on unsupervised pixel-wise tumor detection on the BraTS-2017 dataset with a ROC-AUC of 0.94.
High- and Low-level image component decomposition using VAEs for improved reconstruction and anomaly detection
Nov 27, 2019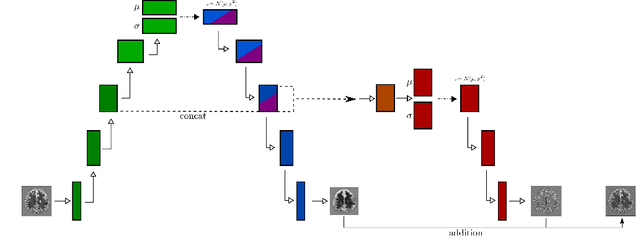
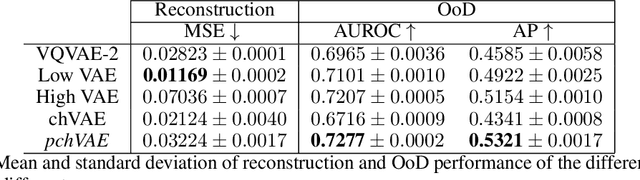
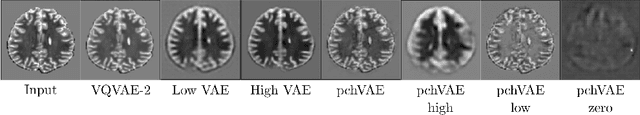
Abstract:Variational Auto-Encoders have often been used for unsupervised pretraining, feature extraction and out-of-distribution and anomaly detection in the medical field. However, VAEs often lack the ability to produce sharp images and learn high-level features. We propose to alleviate these issues by adding a new branch to conditional hierarchical VAEs. This enforces a division between higher-level and lower-level features. Despite the additional computational overhead compared to a normal VAE it results in sharper and better reconstructions and can capture the data distribution similarly well (indicated by a similar or slightly better OoD detection performance).
Unsupervised Anomaly Localization using Variational Auto-Encoders
Jul 11, 2019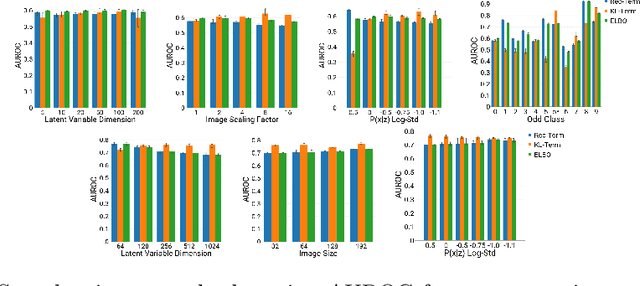
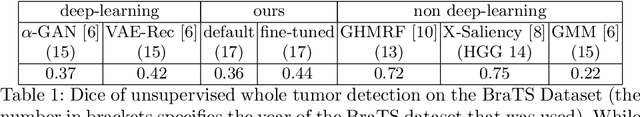
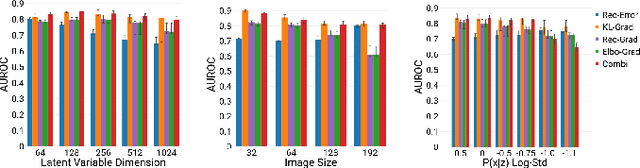
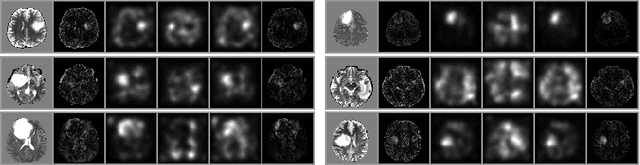
Abstract:An assumption-free automatic check of medical images for potentially overseen anomalies would be a valuable assistance for a radiologist. Deep learning and especially Variational Auto-Encoders (VAEs) have shown great potential in the unsupervised learning of data distributions. In principle, this allows for such a check and even the localization of parts in the image that are most suspicious. Currently, however, the reconstruction-based localization by design requires adjusting the model architecture to the specific problem looked at during evaluation. This contradicts the principle of building assumption-free models. We propose complementing the localization part with a term derived from the Kullback-Leibler (KL)-divergence. For validation, we perform a series of experiments on FashionMNIST as well as on a medical task including >1000 healthy and >250 brain tumor patients. Results show that the proposed formalism outperforms the state of the art VAE-based localization of anomalies across many hyperparameter settings and also shows a competitive max performance.
Deep Probabilistic Modeling of Glioma Growth
Jul 09, 2019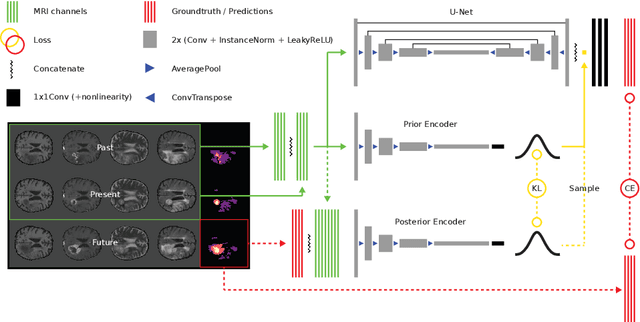
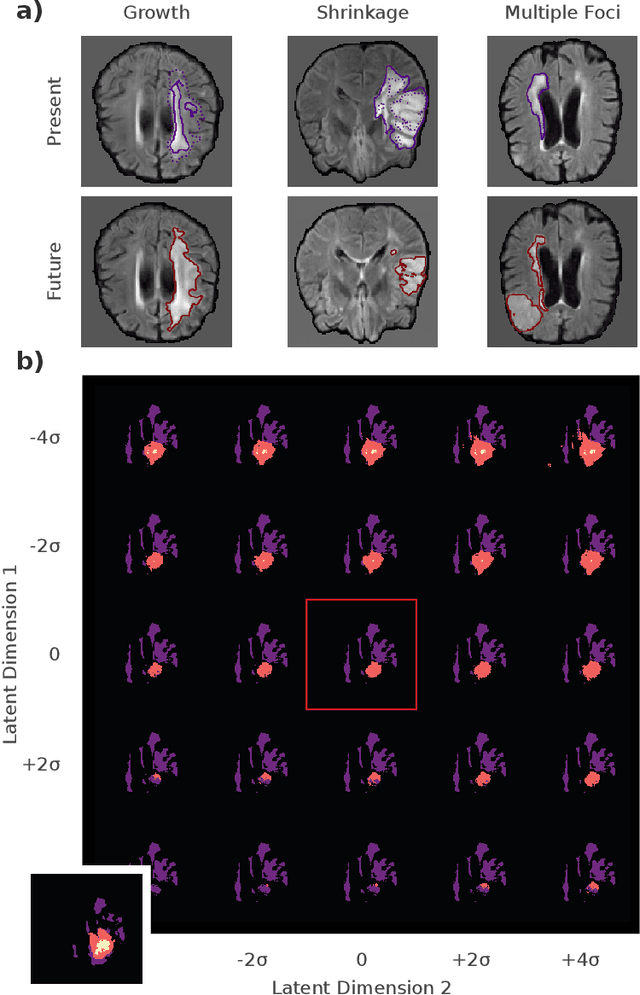
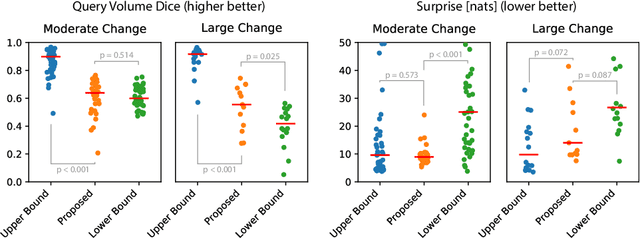
Abstract:Existing approaches to modeling the dynamics of brain tumor growth, specifically glioma, employ biologically inspired models of cell diffusion, using image data to estimate the associated parameters. In this work, we propose an alternative approach based on recent advances in probabilistic segmentation and representation learning that implicitly learns growth dynamics directly from data without an underlying explicit model. We present evidence that our approach is able to learn a distribution of plausible future tumor appearances conditioned on past observations of the same tumor.
nnU-Net: Breaking the Spell on Successful Medical Image Segmentation
Apr 17, 2019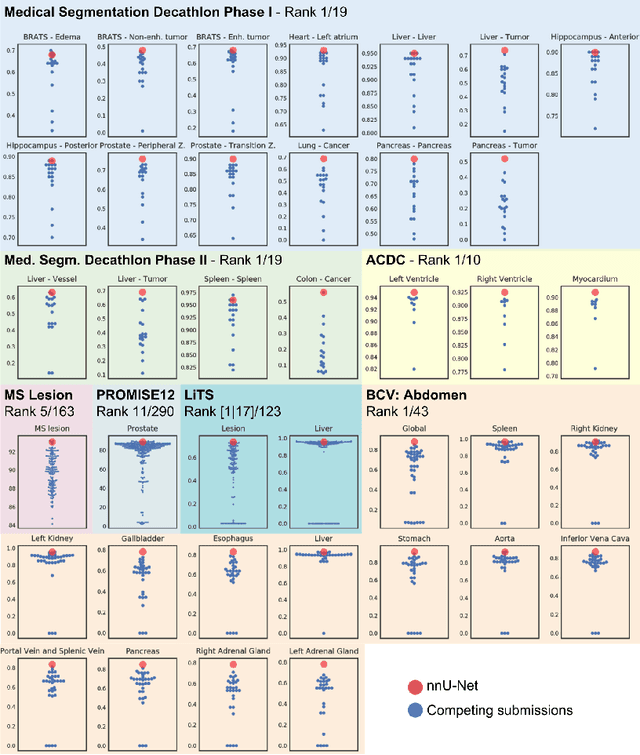
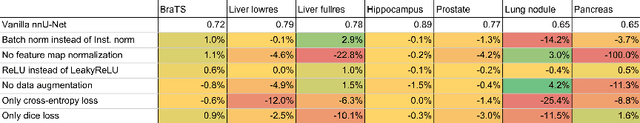
Abstract:Fueled by the diversity of datasets, semantic segmentation is a popular subfield in medical image analysis with a vast number of new methods being proposed each year. This ever-growing jungle of methodologies, however, becomes increasingly impenetrable. At the same time, many proposed methods fail to generalize beyond the experiments they were demonstrated on, thus hampering the process of developing a segmentation algorithm on a new dataset. Here we present nnU-Net ('no-new-Net'), a framework that automatically adapts itself to any given new dataset. While this process was completely human-driven so far, we make a first attempt to automate necessary adaptations such as preprocessing, the exact patch size, batch size, and inference settings based on the properties of a given dataset. Remarkably, nnU-Net strips away the architectural bells and whistles that are typically proposed in the literature and relies on just a simple U-Net architecture embedded in a robust training scheme. Out of the box, nnU-Net achieves state of the art performance on six well-established segmentation challenges. Source code is available at https://github.com/MIC-DKFZ/nnunet.
Segmentation of Roots in Soil with U-Net
Mar 18, 2019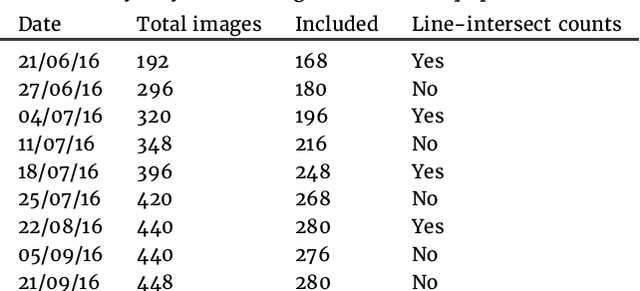
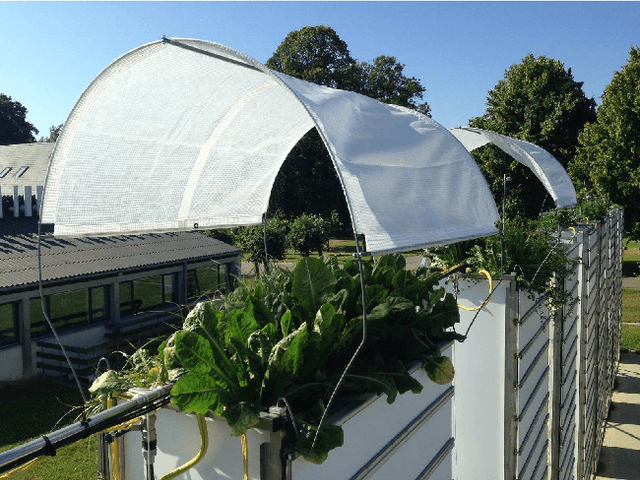
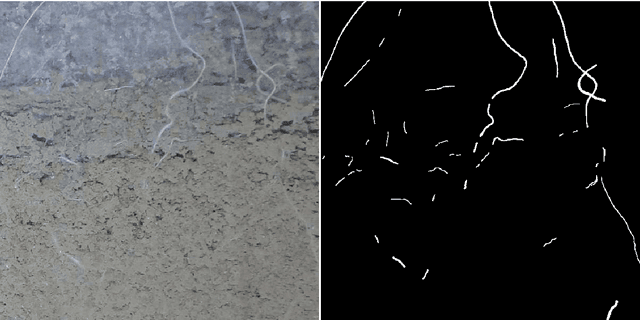


Abstract:Plant root research can provide a way to attain stress-tolerant crops that produce greater yield in a diverse array of conditions. Phenotyping roots in soil is often challenging due to the roots being difficult to access and the use of time consuming manual methods. Rhizotrons allow visual inspection of root growth through transparent surfaces. Agronomists currently manually label photographs of roots obtained from rhizotrons using a line-intersect method to obtain root length density and rooting depth measurements which are essential for their experiments. We investigate the effectiveness of an automated image segmentation method based on the U-Net Convolutional Neural Network (CNN) architecture to enable such measurements. We design a data-set of 50 annotated Chicory (Cichorium intybus L.) root images which we use to train, validate and test the system and compare against a baseline built using the Frangi vesselness filter. We obtain metrics using manual annotations and line-intersect counts. Our results on the held out data show our proposed automated segmentation system to be a viable solution for detecting and quantifying roots. We evaluate our system using 867 images for which we have obtained line-intersect counts, attaining a Spearman rank correlation of 0.9748 and an $r^2$ of 0.9217. We also achieve an $F_1$ of 0.7 when comparing the automated segmentation to the manual annotations, with our automated segmentation system producing segmentations with higher quality than the manual annotations for large portions of the image.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge