Jake Lee
Jet Propulsion Laboratory, California Institute of Technology
Fully Automatic Trace Gas Plume Detection
May 05, 2026Abstract:Future imaging spectrometers will increase data volumes by orders of magnitude, requiring automated detection of trace gas point sources. We present a fully automated framework that combines machine learning-based morphological analysis with physics-based spectroscopic fitting to detect plumes without human participation. Applied to EMIT imaging spectrometer data, the system operates in two modes: "daily digest" that runs automatically on all downlinked data, flagging the largest events for immediate response, and a retrospective analysis that identifies plumes missed by prior human review. The daily digest demonstrates that a significant fraction of the largest plumes can be detected automatically with negligible false positives, while retrospective analysis suggests at least 25% of plumes may have been overlooked. In addition to the previously observed methane point sources, we extend detection to three understudied trace gases: NH3, NO2 and the first observations of carbon monoxide (CO) plume in EMIT imagery.
Adaptive MSD-Splitting: Enhancing C4.5 and Random Forests for Skewed Continuous Attributes
Apr 21, 2026Abstract:The discretization of continuous numerical attributes remains a persistent computational bottleneck in the induction of decision trees, particularly as dataset dimensions scale. Building upon the recently proposed MSD-Splitting technique -- which bins continuous data using the empirical mean and standard deviation to dramatically improve the efficiency and accuracy of the C4.5 algorithm -- we introduce Adaptive MSD-Splitting (AMSD). While standard MSD-Splitting is highly effective for approximately symmetric distributions, its rigid adherence to fixed one-standard-deviation cutoffs can lead to catastrophic information loss in highly skewed data, a common artifact in real-world biomedical and financial datasets. AMSD addresses this by dynamically adjusting the standard deviation multiplier based on feature skewness, narrowing intervals in dense regions to preserve discriminative resolution. Furthermore, we integrate AMSD into ensemble methods, specifically presenting the Random Forest-AMSD (RF-AMSD) framework. Empirical evaluations on the Census Income, Heart Disease, Breast Cancer, and Forest Covertype datasets demonstrate that AMSD yields a 2-4% accuracy improvement over standard MSD-Splitting, while maintaining near-identical O(N) time complexity reductions compared to the O(N log N) exhaustive search. Our Random Forest extension achieves state-of-the-art accuracy at a fraction of standard computational costs, confirming the viability of adaptive statistical binning in large-scale ensemble learning architectures.
Agentic generative AI for media content discovery at the national football league
Oct 08, 2025Abstract:Generative AI has unlocked new possibilities in content discovery and management. Through collaboration with the National Football League (NFL), we demonstrate how a generative-AI based workflow enables media researchers and analysts to query relevant historical plays using natural language rather than traditional filter-and-click interfaces. The agentic workflow takes a user query as input, breaks it into elements, and translates them into the underlying database query language. Accuracy and latency are further improved through carefully designed semantic caching. The solution achieves over 95 percent accuracy and reduces the average time to find relevant videos from 10 minutes to 30 seconds, significantly increasing the NFL's operational efficiency and allowing users to focus on producing creative content and engaging storylines.
Onboard Science Instrument Autonomy for the Detection of Microscopy Biosignatures on the Ocean Worlds Life Surveyor
Apr 25, 2023Abstract:The quest to find extraterrestrial life is a critical scientific endeavor with civilization-level implications. Icy moons in our solar system are promising targets for exploration because their liquid oceans make them potential habitats for microscopic life. However, the lack of a precise definition of life poses a fundamental challenge to formulating detection strategies. To increase the chances of unambiguous detection, a suite of complementary instruments must sample multiple independent biosignatures (e.g., composition, motility/behavior, and visible structure). Such an instrument suite could generate 10,000x more raw data than is possible to transmit from distant ocean worlds like Enceladus or Europa. To address this bandwidth limitation, Onboard Science Instrument Autonomy (OSIA) is an emerging discipline of flight systems capable of evaluating, summarizing, and prioritizing observational instrument data to maximize science return. We describe two OSIA implementations developed as part of the Ocean Worlds Life Surveyor (OWLS) prototype instrument suite at the Jet Propulsion Laboratory. The first identifies life-like motion in digital holographic microscopy videos, and the second identifies cellular structure and composition via innate and dye-induced fluorescence. Flight-like requirements and computational constraints were used to lower barriers to infusion, similar to those available on the Mars helicopter, "Ingenuity." We evaluated the OSIA's performance using simulated and laboratory data and conducted a live field test at the hypersaline Mono Lake planetary analog site. Our study demonstrates the potential of OSIA for enabling biosignature detection and provides insights and lessons learned for future mission concepts aimed at exploring the outer solar system.
Mars Image Content Classification: Three Years of NASA Deployment and Recent Advances
Feb 09, 2021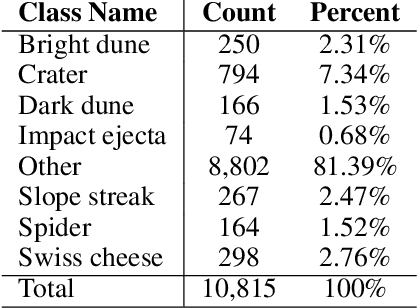
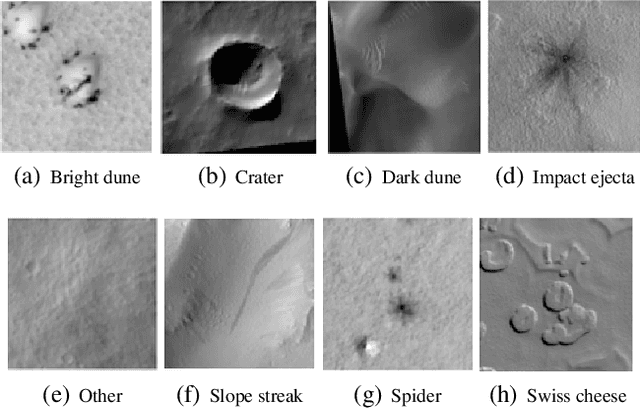
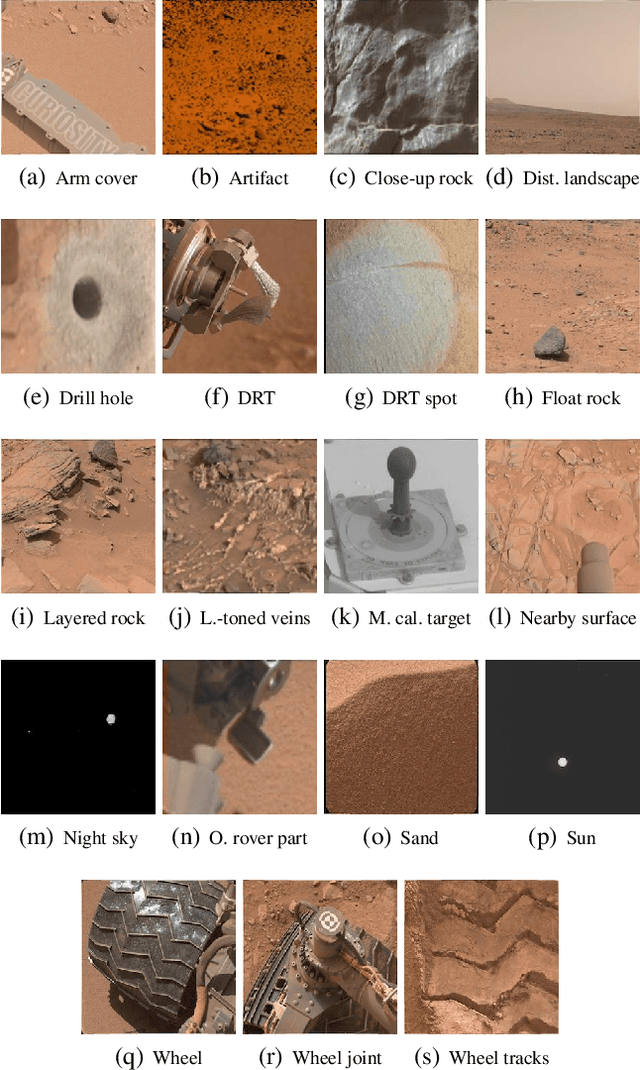


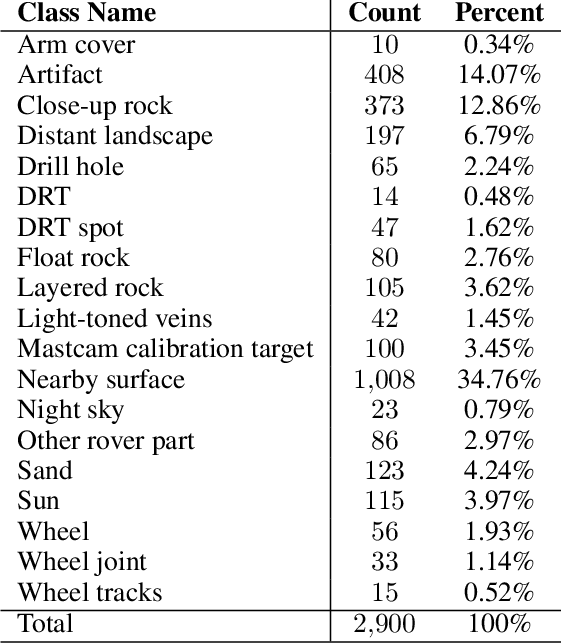
Abstract:The NASA Planetary Data System hosts millions of images acquired from the planet Mars. To help users quickly find images of interest, we have developed and deployed content-based classification and search capabilities for Mars orbital and surface images. The deployed systems are publicly accessible using the PDS Image Atlas. We describe the process of training, evaluating, calibrating, and deploying updates to two CNN classifiers for images collected by Mars missions. We also report on three years of deployment including usage statistics, lessons learned, and plans for the future.
What Does CNN Shift Invariance Look Like? A Visualization Study
Nov 09, 2020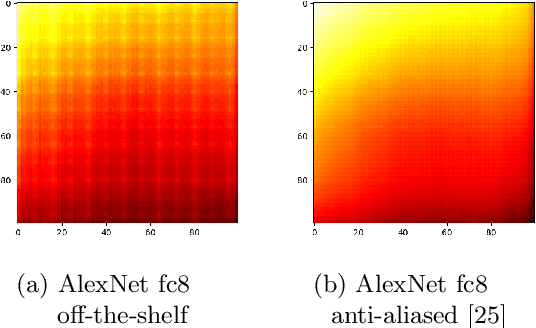
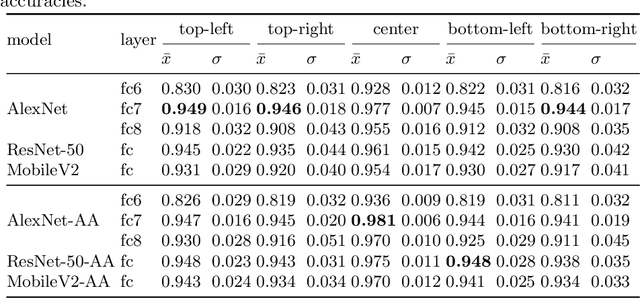
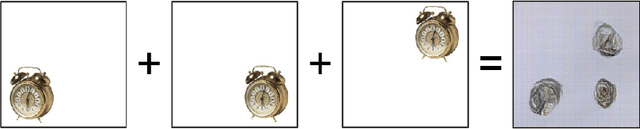
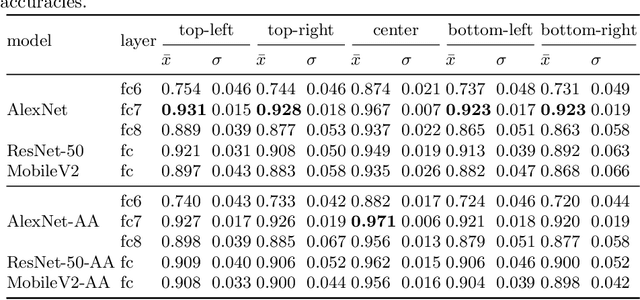
Abstract:Feature extraction with convolutional neural networks (CNNs) is a popular method to represent images for machine learning tasks. These representations seek to capture global image content, and ideally should be independent of geometric transformations. We focus on measuring and visualizing the shift invariance of extracted features from popular off-the-shelf CNN models. We present the results of three experiments comparing representations of millions of images with exhaustively shifted objects, examining both local invariance (within a few pixels) and global invariance (across the image frame). We conclude that features extracted from popular networks are not globally invariant, and that biases and artifacts exist within this variance. Additionally, we determine that anti-aliased models significantly improve local invariance but do not impact global invariance. Finally, we provide a code repository for experiment reproduction, as well as a website to interact with our results at https://jakehlee.github.io/visualize-invariance.
Interpretable Discovery in Large Image Data Sets
Jun 21, 2018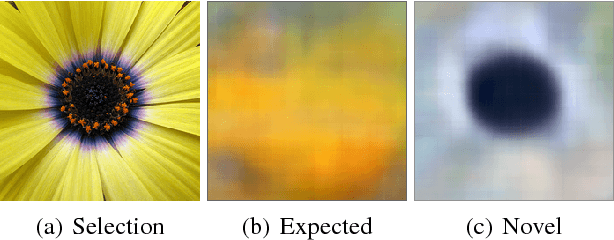
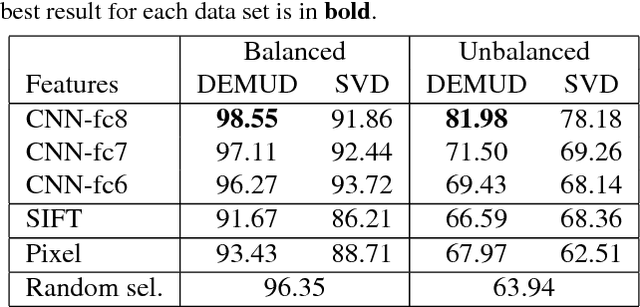
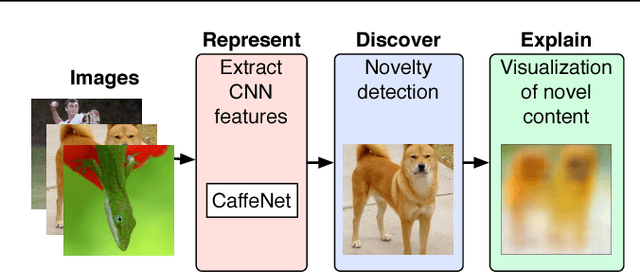
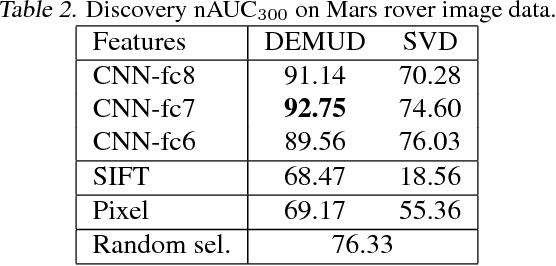
Abstract:Automated detection of new, interesting, unusual, or anomalous images within large data sets has great value for applications from surveillance (e.g., airport security) to science (observations that don't fit a given theory can lead to new discoveries). Many image data analysis systems are turning to convolutional neural networks (CNNs) to represent image content due to their success in achieving high classification accuracy rates. However, CNN representations are notoriously difficult for humans to interpret. We describe a new strategy that combines novelty detection with CNN image features to achieve rapid discovery with interpretable explanations of novel image content. We applied this technique to familiar images from ImageNet as well as to a scientific image collection from planetary science.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge