Huijie Fan
TMUAD: Enhancing Logical Capabilities in Unified Anomaly Detection Models with a Text Memory Bank
Aug 29, 2025Abstract:Anomaly detection, which aims to identify anomalies deviating from normal patterns, is challenging due to the limited amount of normal data available. Unlike most existing unified methods that rely on carefully designed image feature extractors and memory banks to capture logical relationships between objects, we introduce a text memory bank to enhance the detection of logical anomalies. Specifically, we propose a Three-Memory framework for Unified structural and logical Anomaly Detection (TMUAD). First, we build a class-level text memory bank for logical anomaly detection by the proposed logic-aware text extractor, which can capture rich logical descriptions of objects from input images. Second, we construct an object-level image memory bank that preserves complete object contours by extracting features from segmented objects. Third, we employ visual encoders to extract patch-level image features for constructing a patch-level memory bank for structural anomaly detection. These three complementary memory banks are used to retrieve and compare normal images that are most similar to the query image, compute anomaly scores at multiple levels, and fuse them into a final anomaly score. By unifying structural and logical anomaly detection through collaborative memory banks, TMUAD achieves state-of-the-art performance across seven publicly available datasets involving industrial and medical domains. The model and code are available at https://github.com/SIA-IDE/TMUAD.
Distribution-aware Forgetting Compensation for Exemplar-Free Lifelong Person Re-identification
Apr 22, 2025Abstract:Lifelong Person Re-identification (LReID) suffers from a key challenge in preserving old knowledge while adapting to new information. The existing solutions include rehearsal-based and rehearsal-free methods to address this challenge. Rehearsal-based approaches rely on knowledge distillation, continuously accumulating forgetting during the distillation process. Rehearsal-free methods insufficiently learn the distribution of each domain, leading to forgetfulness over time. To solve these issues, we propose a novel Distribution-aware Forgetting Compensation (DAFC) model that explores cross-domain shared representation learning and domain-specific distribution integration without using old exemplars or knowledge distillation. We propose a Text-driven Prompt Aggregation (TPA) that utilizes text features to enrich prompt elements and guide the prompt model to learn fine-grained representations for each instance. This can enhance the differentiation of identity information and establish the foundation for domain distribution awareness. Then, Distribution-based Awareness and Integration (DAI) is designed to capture each domain-specific distribution by a dedicated expert network and adaptively consolidate them into a shared region in high-dimensional space. In this manner, DAI can consolidate and enhance cross-domain shared representation learning while alleviating catastrophic forgetting. Furthermore, we develop a Knowledge Consolidation Mechanism (KCM) that comprises instance-level discrimination and cross-domain consistency alignment strategies to facilitate model adaptive learning of new knowledge from the current domain and promote knowledge consolidation learning between acquired domain-specific distributions, respectively. Experimental results show that our DAFC outperforms state-of-the-art methods. Our code is available at https://github.com/LiuShiBen/DAFC.
GAFusion: Adaptive Fusing LiDAR and Camera with Multiple Guidance for 3D Object Detection
Nov 01, 2024



Abstract:Recent years have witnessed the remarkable progress of 3D multi-modality object detection methods based on the Bird's-Eye-View (BEV) perspective. However, most of them overlook the complementary interaction and guidance between LiDAR and camera. In this work, we propose a novel multi-modality 3D objection detection method, named GAFusion, with LiDAR-guided global interaction and adaptive fusion. Specifically, we introduce sparse depth guidance (SDG) and LiDAR occupancy guidance (LOG) to generate 3D features with sufficient depth information. In the following, LiDAR-guided adaptive fusion transformer (LGAFT) is developed to adaptively enhance the interaction of different modal BEV features from a global perspective. Meanwhile, additional downsampling with sparse height compression and multi-scale dual-path transformer (MSDPT) are designed to enlarge the receptive fields of different modal features. Finally, a temporal fusion module is introduced to aggregate features from previous frames. GAFusion achieves state-of-the-art 3D object detection results with 73.6$\%$ mAP and 74.9$\%$ NDS on the nuScenes test set.
Selective Aggregation for Low-Rank Adaptation in Federated Learning
Oct 02, 2024



Abstract:We investigate LoRA in federated learning through the lens of the asymmetry analysis of the learned $A$ and $B$ matrices. In doing so, we uncover that $A$ matrices are responsible for learning general knowledge, while $B$ matrices focus on capturing client-specific knowledge. Based on this finding, we introduce Federated Share-A Low-Rank Adaptation (FedSA-LoRA), which employs two low-rank trainable matrices $A$ and $B$ to model the weight update, but only $A$ matrices are shared with the server for aggregation. Moreover, we delve into the relationship between the learned $A$ and $B$ matrices in other LoRA variants, such as rsLoRA and VeRA, revealing a consistent pattern. Consequently, we extend our FedSA-LoRA method to these LoRA variants, resulting in FedSA-rsLoRA and FedSA-VeRA. In this way, we establish a general paradigm for integrating LoRA with FL, offering guidance for future work on subsequent LoRA variants combined with FL. Extensive experimental results on natural language understanding and generation tasks demonstrate the effectiveness of the proposed method.
Attribute-Text Guided Forgetting Compensation for Lifelong Person Re-Identification
Sep 30, 2024Abstract:Lifelong person re-identification (LReID) aims to continuously learn from non-stationary data to match individuals in different environments. Each task is affected by variations in illumination and person-related information (such as pose and clothing), leading to task-wise domain gaps. Current LReID methods focus on task-specific knowledge and ignore intrinsic task-shared representations within domain gaps, limiting model performance. Bridging task-wise domain gaps is crucial for improving anti-forgetting and generalization capabilities, especially when accessing limited old classes during training. To address these issues, we propose a novel attribute-text guided forgetting compensation (ATFC) model, which explores text-driven global representations of identity-related information and attribute-related local representations of identity-free information for LReID. Due to the lack of paired text-image data, we design an attribute-text generator (ATG) to dynamically generate a text descriptor for each instance. We then introduce a text-guided aggregation network (TGA) to explore robust text-driven global representations for each identity and knowledge transfer. Furthermore, we propose an attribute compensation network (ACN) to investigate attribute-related local representations, which distinguish similar identities and bridge domain gaps. Finally, we develop an attribute anti-forgetting (AF) loss and knowledge transfer (KT) loss to minimize domain gaps and achieve knowledge transfer, improving model performance. Extensive experiments demonstrate that our ATFC method achieves superior performance, outperforming existing LReID methods by over 9.0$\%$/7.4$\%$ in average mAP/R-1 on the seen dataset.
GMT: A Robust Global Association Model for Multi-Target Multi-Camera Tracking
Jul 01, 2024



Abstract:In the task of multi-target multi-camera (MTMC) tracking of pedestrians, the data association problem is a key issue and main challenge, especially with complications arising from camera movements, lighting variations, and obstructions. However, most MTMC models adopt two-step approaches, thus heavily depending on the results of the first-step tracking in practical applications. Moreover, the same targets crossing different cameras may exhibit significant appearance variations, which further increases the difficulty of cross-camera matching. To address the aforementioned issues, we propose a global online MTMC tracking model that addresses the dependency on the first tracking stage in two-step methods and enhances cross-camera matching. Specifically, we propose a transformer-based global MTMC association module to explore target associations across different cameras and frames, generating global trajectories directly. Additionally, to integrate the appearance and spatio-temporal features of targets, we propose a feature extraction and fusion module for MTMC tracking. This module enhances feature representation and establishes correlations between the features of targets across multiple cameras. To accommodate high scene diversity and complex lighting condition variations, we have established the VisionTrack dataset, which enables the development of models that are more generalized and robust to various environments. Our model demonstrates significant improvements over comparison methods on the VisionTrack dataset and others.
Diverse Representation Embedding for Lifelong Person Re-Identification
Apr 02, 2024Abstract:Lifelong Person Re-Identification (LReID) aims to continuously learn from successive data streams, matching individuals across multiple cameras. The key challenge for LReID is how to effectively preserve old knowledge while incrementally learning new information, which is caused by task-level domain gaps and limited old task datasets. Existing methods based on CNN backbone are insufficient to explore the representation of each instance from different perspectives, limiting model performance on limited old task datasets and new task datasets. Unlike these methods, we propose a Diverse Representations Embedding (DRE) framework that first explores a pure transformer for LReID. The proposed DRE preserves old knowledge while adapting to new information based on instance-level and task-level layout. Concretely, an Adaptive Constraint Module (ACM) is proposed to implement integration and push away operations between multiple overlapping representations generated by transformer-based backbone, obtaining rich and discriminative representations for each instance to improve adaptive ability of LReID. Based on the processed diverse representations, we propose Knowledge Update (KU) and Knowledge Preservation (KP) strategies at the task-level layout by introducing the adjustment model and the learner model. KU strategy enhances the adaptive learning ability of learner models for new information under the adjustment model prior, and KP strategy preserves old knowledge operated by representation-level alignment and logit-level supervision in limited old task datasets while guaranteeing the adaptive learning information capacity of the LReID model. Compared to state-of-the-art methods, our method achieves significantly improved performance in holistic, large-scale, and occluded datasets.
Residual Denoising Diffusion Models
Aug 25, 2023



Abstract:Current diffusion-based image restoration methods feed degraded input images as conditions into the noise estimation network. However, interpreting this diffusion process is challenging since it essentially generates the target image from the noise. To establish a unified and more interpretable model for image generation and restoration, we propose residual denoising diffusion models (RDDM). In contrast to existing diffusion models (e.g., DDPM or DDIM) that focus solely on noise estimation, our RDDM predicts residuals to represent directional diffusion from the target domain to the input domain, while concurrently estimating noise to account for random perturbations in the diffusion process. The introduction of residuals allows us to redefine the forward diffusion process, wherein the target image progressively diffuses into a purely noisy image or a noise-carrying input image, thus unifying image generation and restoration. We demonstrate that our sampling process is consistent with that of DDPM and DDIM through coefficient transformation, and propose a partially path-independent generation process to better understand the reverse process. Notably, with native support for conditional inputs, our RDDM enables a generic UNet, trained with only an $\ell _1$ loss and a batch size of 1, to compete with state-of-the-art image restoration methods. We provide code and pre-trained models to encourage further exploration, application, and development of our innovative framework (https://github.com/nachifur/RDDM).
Automatic label correction based on CCESD
Oct 05, 2020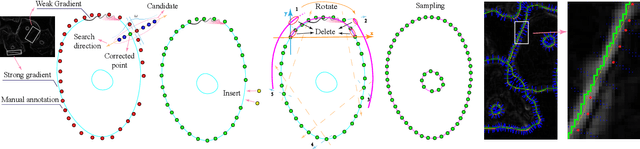
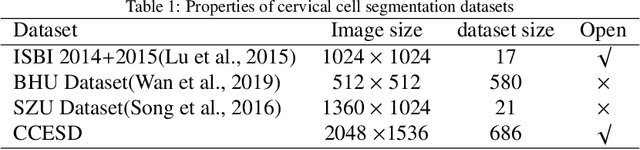
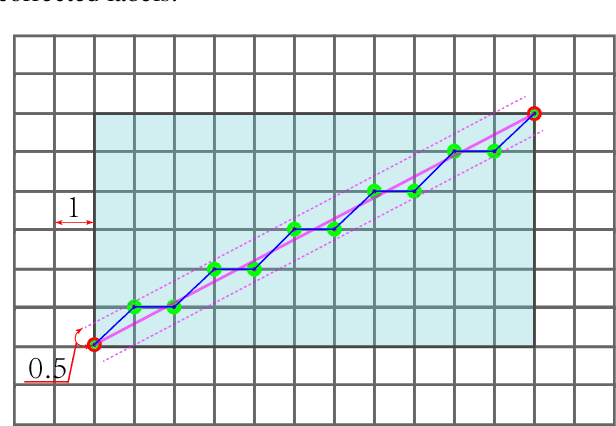
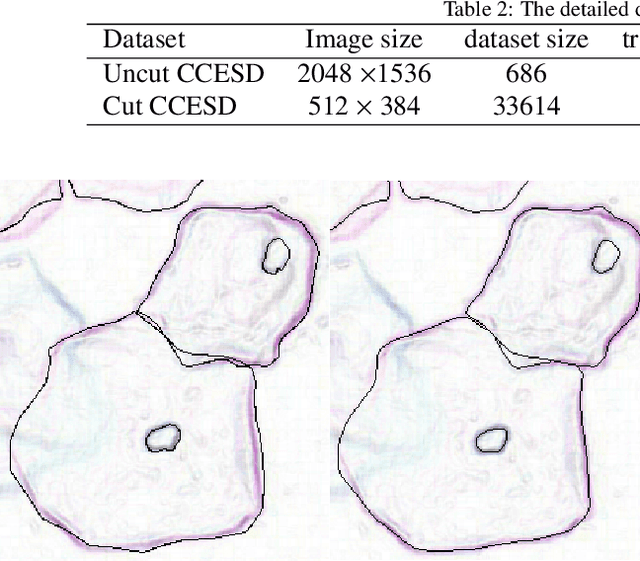
Abstract:In the computer-aided diagnosis of cervical precancerous lesions, it is essential for accurate cell segmentation. For a cervical cell image with multi-cell overlap (n>3), blurry and noisy background, and low contrast, it is difficult for a professional doctor to obtain an ultra-high-precision labeled image. On the other hand, it is possible for the annotator to draw the outline of the cell as accurately as possible. However, if the label edge position is inaccurate, the accuracy of the training model will decrease, and it will have a great impact on the accuracy of the model evaluation. We designed an automatic label correction algorithm based on gradient guidance, which can solve the effects of poor edge position accuracy and differences between different annotators during manual labeling. At the same time, an open cervical cell edge segmentation dataset (CCESD) with higher labeling accuracy was constructed. We also use deep learning models to generate the baseline performance on CCESD. Using the modified labeling data to train multiple models compared to the original labeling data can be improved 7% average precision (AP). The implementation is available at https://github.com/nachifur-ljw/label_correction_based_CCESD.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge