Gustavo Carneiro
Australian Institute for Machine Learning
3D Semantic Mapping from Arthroscopy using Out-of-distribution Pose and Depth and In-distribution Segmentation Training
Jun 10, 2021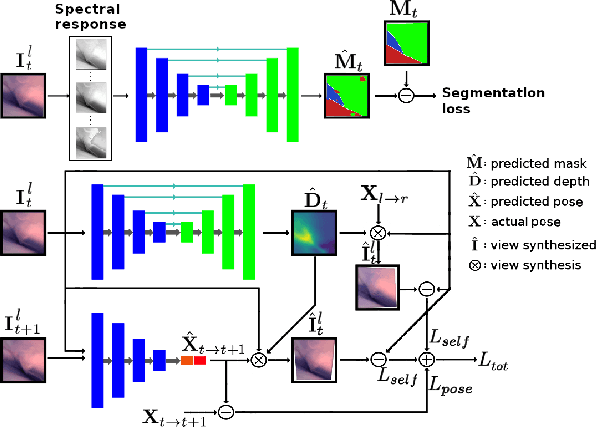
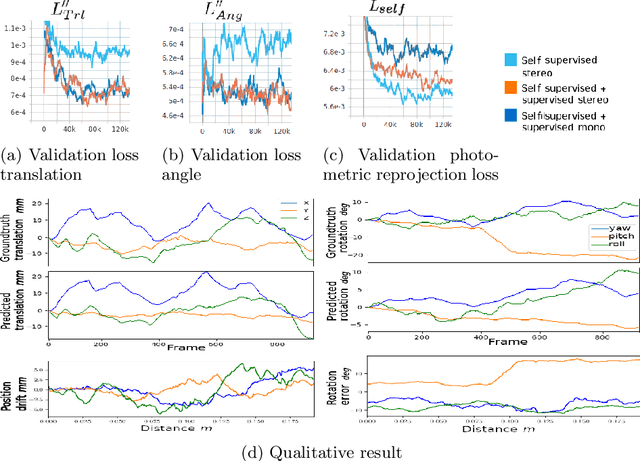

Abstract:Minimally invasive surgery (MIS) has many documented advantages, but the surgeon's limited visual contact with the scene can be problematic. Hence, systems that can help surgeons navigate, such as a method that can produce a 3D semantic map, can compensate for the limitation above. In theory, we can borrow 3D semantic mapping techniques developed for robotics, but this requires finding solutions to the following challenges in MIS: 1) semantic segmentation, 2) depth estimation, and 3) pose estimation. In this paper, we propose the first 3D semantic mapping system from knee arthroscopy that solves the three challenges above. Using out-of-distribution non-human datasets, where pose could be labeled, we jointly train depth+pose estimators using selfsupervised and supervised losses. Using an in-distribution human knee dataset, we train a fully-supervised semantic segmentation system to label arthroscopic image pixels into femur, ACL, and meniscus. Taking testing images from human knees, we combine the results from these two systems to automatically create 3D semantic maps of the human knee. The result of this work opens the pathway to the generation of intraoperative 3D semantic mapping, registration with pre-operative data, and robotic-assisted arthroscopy
Probabilistic task modelling for meta-learning
Jun 09, 2021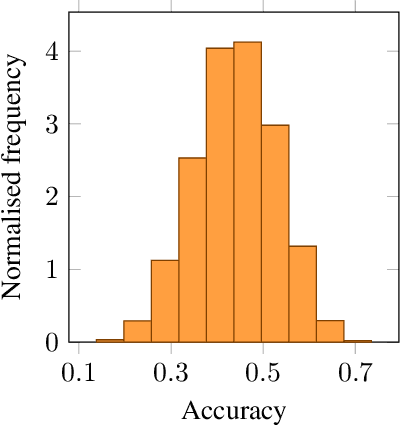
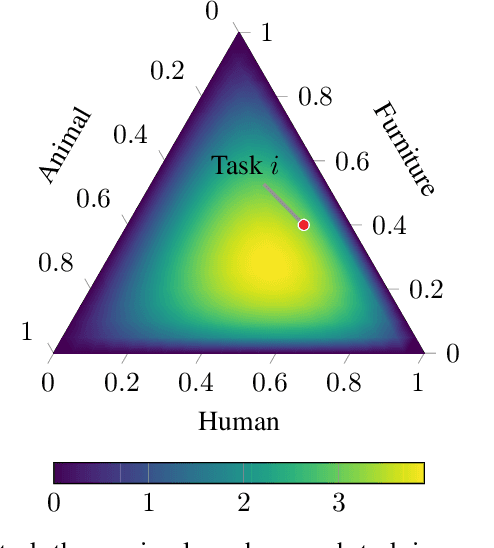
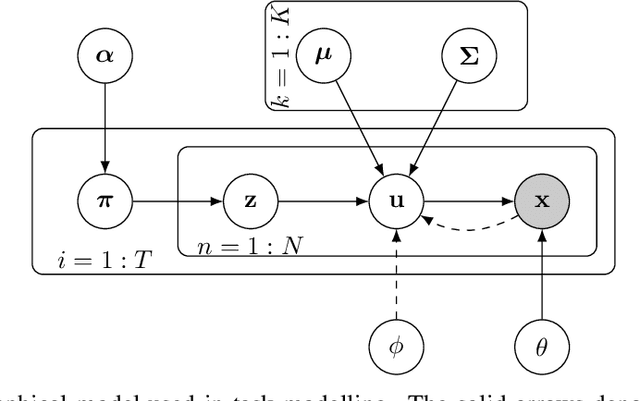
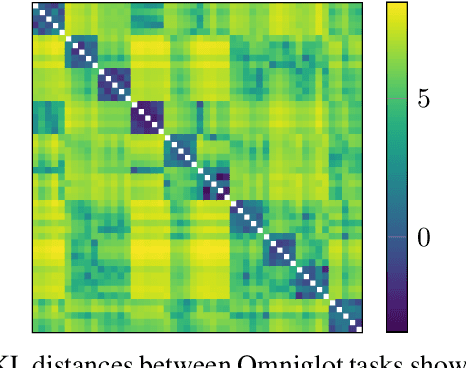
Abstract:We propose probabilistic task modelling -- a generative probabilistic model for collections of tasks used in meta-learning. The proposed model combines variational auto-encoding and latent Dirichlet allocation to model each task as a mixture of Gaussian distribution in an embedding space. Such modelling provides an explicit representation of a task through its task-theme mixture. We present an efficient approximation inference technique based on variational inference method for empirical Bayes parameter estimation. We perform empirical evaluations to validate the task uncertainty and task distance produced by the proposed method through correlation diagrams of the prediction accuracy on testing tasks. We also carry out experiments of task selection in meta-learning to demonstrate how the task relatedness inferred from the proposed model help to facilitate meta-learning algorithms.
Self-supervised Lesion Change Detection and Localisation in Longitudinal Multiple Sclerosis Brain Imaging
Jun 02, 2021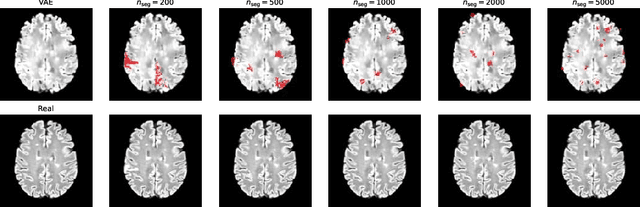
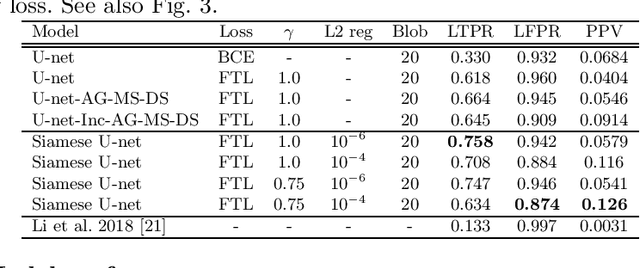
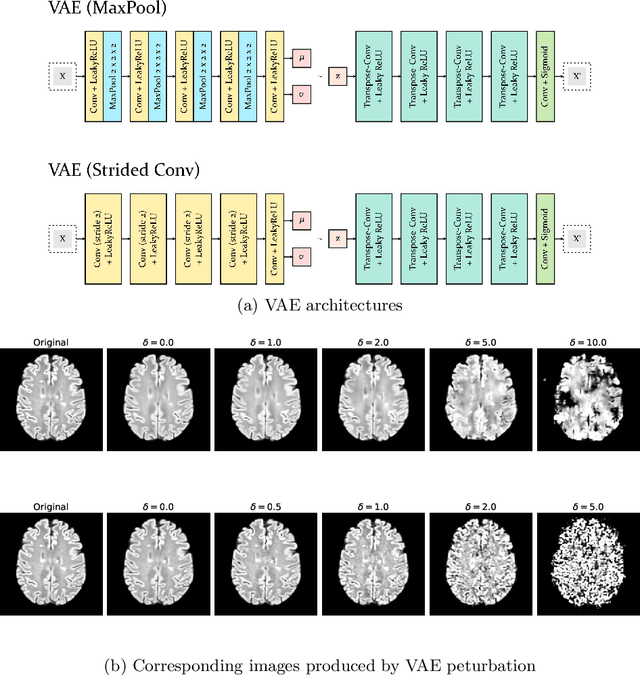
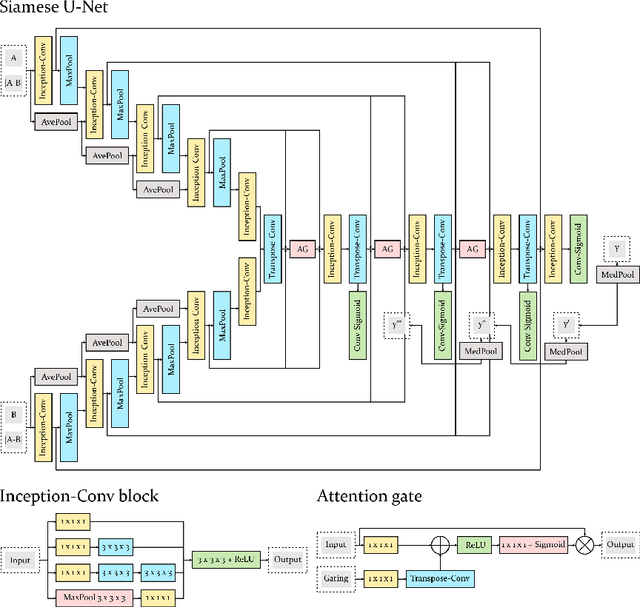
Abstract:Longitudinal imaging forms an essential component in the management and follow-up of many medical conditions. The presence of lesion changes on serial imaging can have significant impact on clinical decision making, highlighting the important role for automated change detection. Lesion changes can represent anomalies in serial imaging, which implies a limited availability of annotations and a wide variety of possible changes that need to be considered. Hence, we introduce a new unsupervised anomaly detection and localisation method trained exclusively with serial images that do not contain any lesion changes. Our training automatically synthesises lesion changes in serial images, introducing detection and localisation pseudo-labels that are used to self-supervise the training of our model. Given the rarity of these lesion changes in the synthesised images, we train the model with the imbalance robust focal Tversky loss. When compared to supervised models trained on different datasets, our method shows competitive performance in the detection and localisation of new demyelinating lesions on longitudinal magnetic resonance imaging in multiple sclerosis patients. Code for the models will be made available on GitHub.
ScanMix: Learning from Severe Label Noise via Semantic Clustering and Semi-Supervised Learning
Mar 21, 2021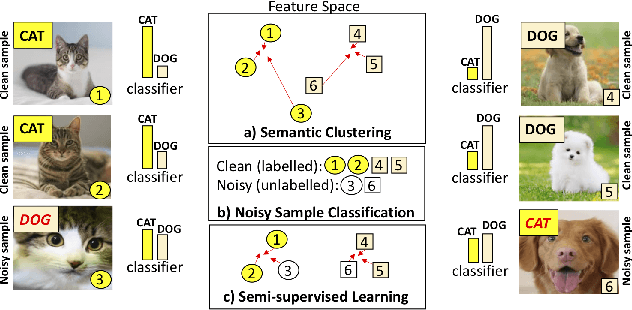
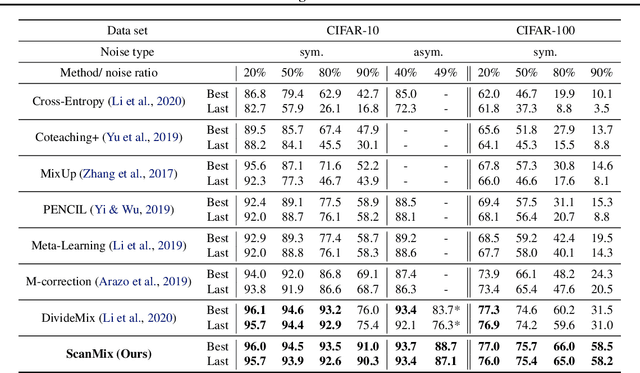
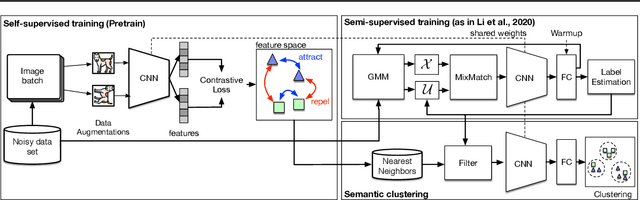
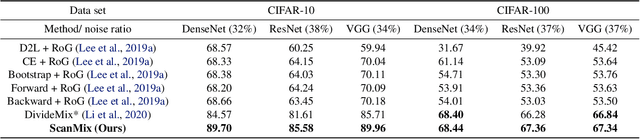
Abstract:In this paper, we address the problem of training deep neural networks in the presence of severe label noise. Our proposed training algorithm ScanMix, combines semantic clustering with semi-supervised learning (SSL) to improve the feature representations and enable an accurate identification of noisy samples, even in severe label noise scenarios. To be specific, ScanMix is designed based on the expectation maximisation (EM) framework, where the E-step estimates the value of a latent variable to cluster the training images based on their appearance representations and classification results, and the M-step optimises the SSL classification and learns effective feature representations via semantic clustering. In our evaluations, we show state-of-the-art results on standard benchmarks for symmetric, asymmetric and semantic label noise on CIFAR-10 and CIFAR-100, as well as large scale real label noise on WebVision. Most notably, for the benchmarks contaminated with large noise rates (80% and above), our results are up to 27% better than the related work. The code is available at https://github.com/ragavsachdeva/ScanMix.
LongReMix: Robust Learning with High Confidence Samples in a Noisy Label Environment
Mar 06, 2021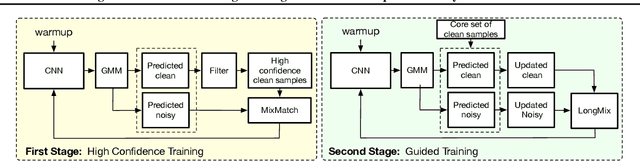
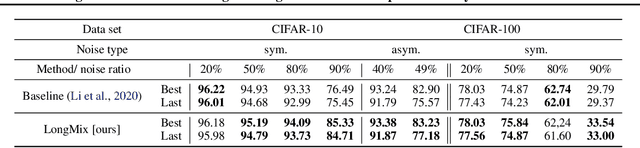
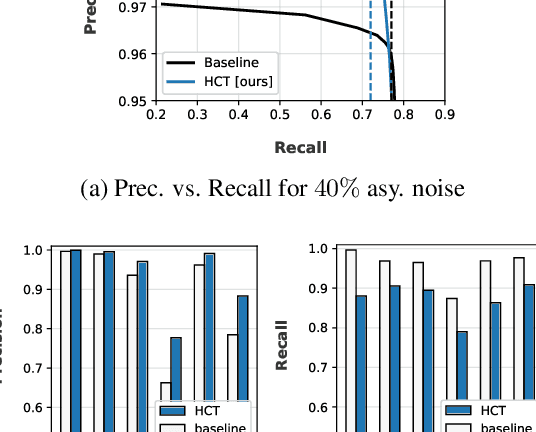
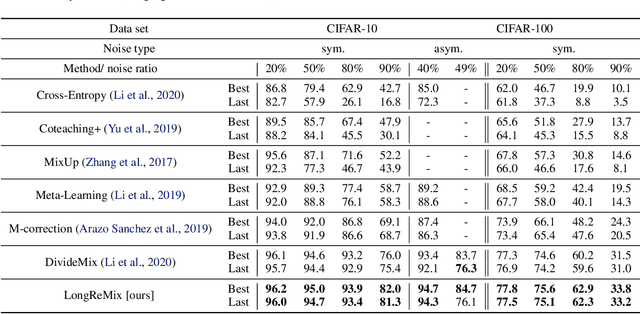
Abstract:Deep neural network models are robust to a limited amount of label noise, but their ability to memorise noisy labels in high noise rate problems is still an open issue. The most competitive noisy-label learning algorithms rely on a 2-stage process comprising an unsupervised learning to classify training samples as clean or noisy, followed by a semi-supervised learning that minimises the empirical vicinal risk (EVR) using a labelled set formed by samples classified as clean, and an unlabelled set with samples classified as noisy. In this paper, we hypothesise that the generalisation of such 2-stage noisy-label learning methods depends on the precision of the unsupervised classifier and the size of the training set to minimise the EVR. We empirically validate these two hypotheses and propose the new 2-stage noisy-label training algorithm LongReMix. We test LongReMix on the noisy-label benchmarks CIFAR-10, CIFAR-100, WebVision, Clothing1M, and Food101-N. The results show that our LongReMix generalises better than competing approaches, particularly in high label noise problems. Furthermore, our approach achieves state-of-the-art performance in most datasets. The code will be available upon paper acceptance.
Noisy Label Learning for Large-scale Medical Image Classification
Mar 06, 2021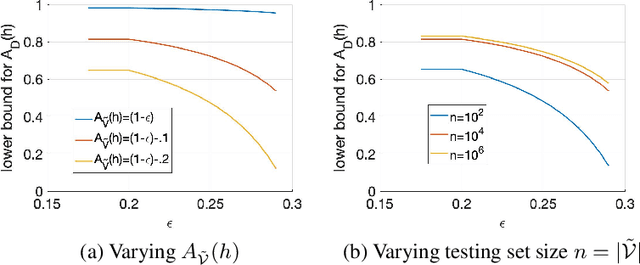
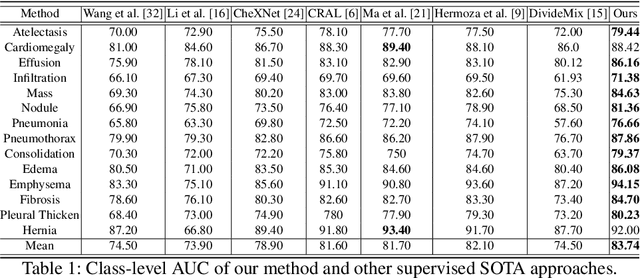
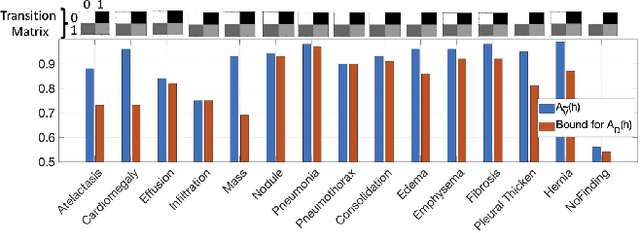

Abstract:The classification accuracy of deep learning models depends not only on the size of their training sets, but also on the quality of their labels. In medical image classification, large-scale datasets are becoming abundant, but their labels will be noisy when they are automatically extracted from radiology reports using natural language processing tools. Given that deep learning models can easily overfit these noisy-label samples, it is important to study training approaches that can handle label noise. In this paper, we adapt a state-of-the-art (SOTA) noisy-label multi-class training approach to learn a multi-label classifier for the dataset Chest X-ray14, which is a large scale dataset known to contain label noise in the training set. Given that this dataset also has label noise in the testing set, we propose a new theoretically sound method to estimate the performance of the model on a hidden clean testing data, given the result on the noisy testing data. Using our clean data performance estimation, we notice that the majority of label noise on Chest X-ray14 is present in the class 'No Finding', which is intuitively correct because this is the most likely class to contain one or more of the 14 diseases due to labelling mistakes.
Self-supervised Mean Teacher for Semi-supervised Chest X-ray Classification
Mar 05, 2021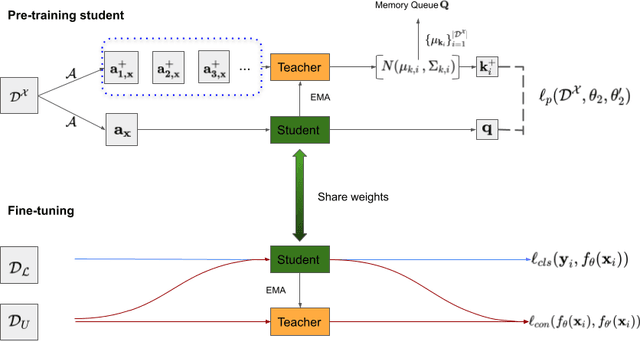
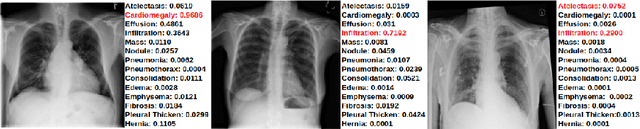
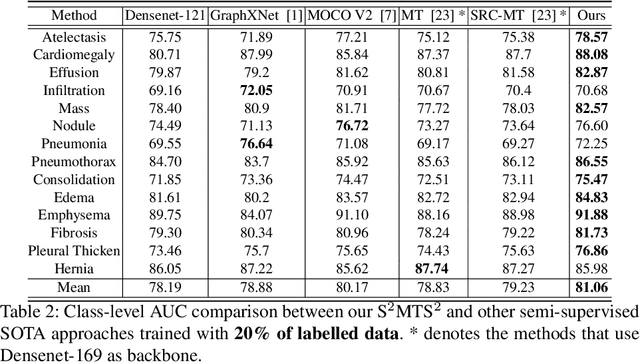
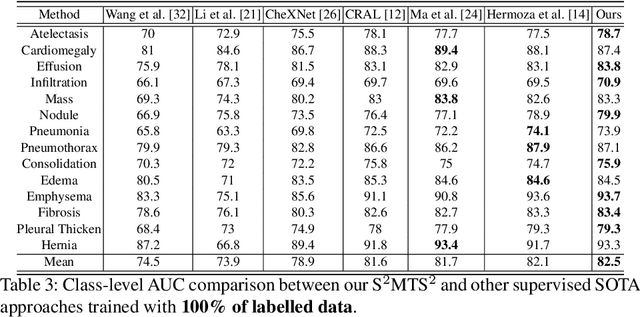
Abstract:The training of deep learning models generally requires a large amount of annotated data for effective convergence and generalisation. However, obtaining high-quality annotations is a laboursome and expensive process due to the need of expert radiologists for the labelling task. The study of semi-supervised learning in medical image analysis is then of crucial importance given that it is much less expensive to obtain unlabelled images than to acquire images labelled by expert radiologists.Essentially, semi-supervised methods leverage large sets of unlabelled data to enable better training convergence and generalisation than if we use only the small set of labelled images.In this paper, we propose the Self-supervised Mean Teacher for Semi-supervised (S$^2$MTS$^2$) learning that combines self-supervised mean-teacher pre-training with semi-supervised fine-tuning. The main innovation of S$^2$MTS$^2$ is the self-supervised mean-teacher pre-training based on the joint contrastive learning, which uses an infinite number of pairs of positive query and key features to improve the mean-teacher representation. The model is then fine-tuned using the exponential moving average teacher framework trained with semi-supervised learning.We validate S$^2$MTS$^2$ on the thorax disease multi-label classification problem from the dataset Chest X-ray14, where we show that it outperforms the previous SOTA semi-supervised learning methods by a large margin.
Constrained Contrastive Distribution Learning for Unsupervised Anomaly Detection and Localisation in Medical Images
Mar 05, 2021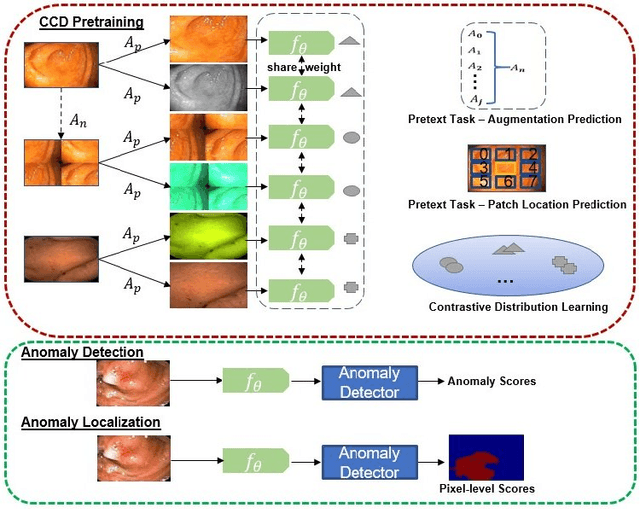
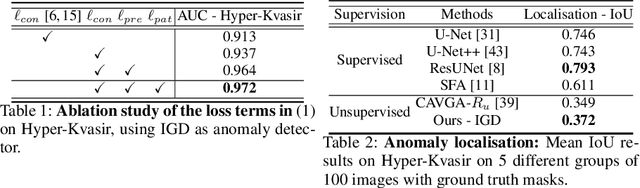
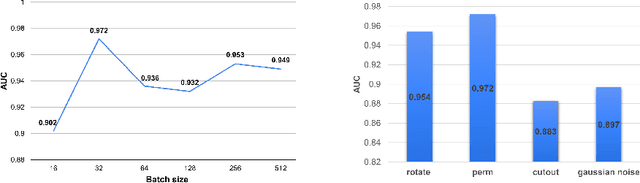
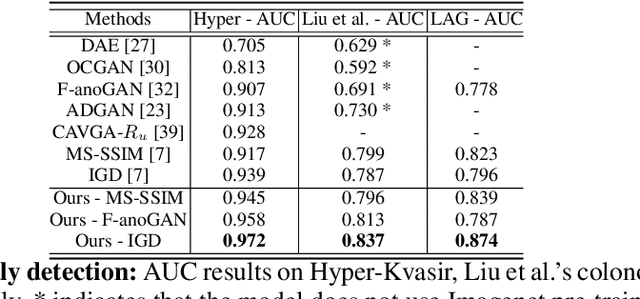
Abstract:Unsupervised anomaly detection (UAD) learns one-class classifiers exclusively with normal (i.e., healthy) images to detect any abnormal (i.e., unhealthy) samples that do not conform to the expected normal patterns. UAD has two main advantages over its fully supervised counterpart. Firstly, it is able to directly leverage large datasets available from health screening programs that contain mostly normal image samples, avoiding the costly manual labelling of abnormal samples and the subsequent issues involved in training with extremely class-imbalanced data. Further, UAD approaches can potentially detect and localise any type of lesions that deviate from the normal patterns. One significant challenge faced by UAD methods is how to learn effective low-dimensional image representations to detect and localise subtle abnormalities, generally consisting of small lesions. To address this challenge, we propose a novel self-supervised representation learning method, called Constrained Contrastive Distribution learning for anomaly detection (CCD), which learns fine-grained feature representations by simultaneously predicting the distribution of augmented data and image contexts using contrastive learning with pretext constraints. The learned representations can be leveraged to train more anomaly-sensitive detection models. Extensive experiment results show that our method outperforms current state-of-the-art UAD approaches on three different colonoscopy and fundus screening datasets. Our code is available at https://github.com/tianyu0207/CCD.
Post-hoc Overall Survival Time Prediction from Brain MRI
Feb 22, 2021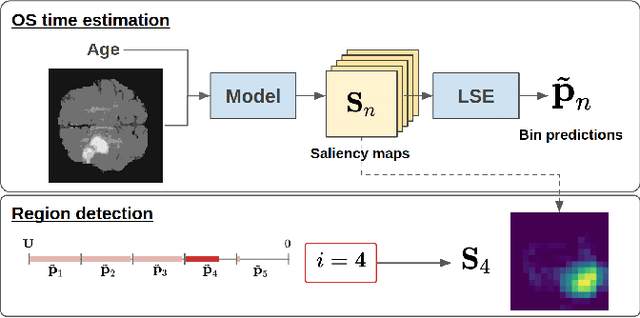
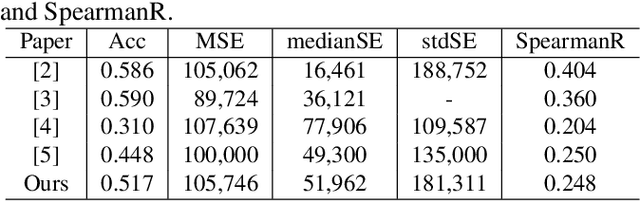
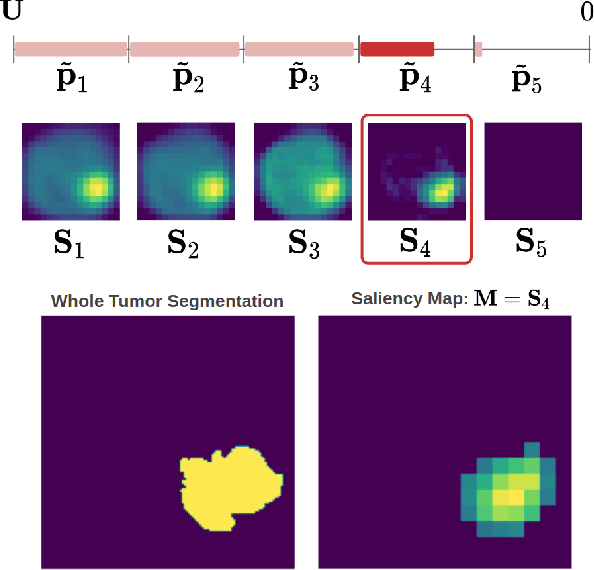
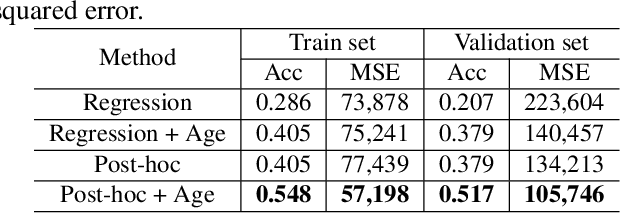
Abstract:Overall survival (OS) time prediction is one of the most common estimates of the prognosis of gliomas and is used to design an appropriate treatment planning. State-of-the-art (SOTA) methods for OS time prediction follow a pre-hoc approach that require computing the segmentation map of the glioma tumor sub-regions (necrotic, edema tumor, enhancing tumor) for estimating OS time. However, the training of the segmentation methods require ground truth segmentation labels which are tedious and expensive to obtain. Given that most of the large-scale data sets available from hospitals are unlikely to contain such precise segmentation, those SOTA methods have limited applicability. In this paper, we introduce a new post-hoc method for OS time prediction that does not require segmentation map annotation for training. Our model uses medical image and patient demographics (represented by age) as inputs to estimate the OS time and to estimate a saliency map that localizes the tumor as a way to explain the OS time prediction in a post-hoc manner. It is worth emphasizing that although our model can localize tumors, it uses only the ground truth OS time as training signal, i.e., no segmentation labels are needed. We evaluate our post-hoc method on the Multimodal Brain Tumor Segmentation Challenge (BraTS) 2019 data set and show that it achieves competitive results compared to pre-hoc methods with the advantage of not requiring segmentation labels for training.
Similarity of Classification Tasks
Jan 27, 2021Abstract:Recent advances in meta-learning has led to remarkable performances on several few-shot learning benchmarks. However, such success often ignores the similarity between training and testing tasks, resulting in a potential bias evaluation. We, therefore, propose a generative approach based on a variant of Latent Dirichlet Allocation to analyse task similarity to optimise and better understand the performance of meta-learning. We demonstrate that the proposed method can provide an insightful evaluation for meta-learning algorithms on two few-shot classification benchmarks that matches common intuition: the more similar the higher performance. Based on this similarity measure, we propose a task-selection strategy for meta-learning and show that it can produce more accurate classification results than methods that randomly select training tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge