Gustavo Carneiro
Australian Institute for Machine Learning
Residual Pattern Learning for Pixel-wise Out-of-Distribution Detection in Semantic Segmentation
Nov 26, 2022



Abstract:Semantic segmentation models classify pixels into a set of known (``in-distribution'') visual classes. When deployed in an open world, the reliability of these models depends on their ability not only to classify in-distribution pixels but also to detect out-of-distribution (OoD) pixels. Historically, the poor OoD detection performance of these models has motivated the design of methods based on model re-training using synthetic training images that include OoD visual objects. Although successful, these re-trained methods have two issues: 1) their in-distribution segmentation accuracy may drop during re-training, and 2) their OoD detection accuracy does not generalise well to new contexts (e.g., country surroundings) outside the training set (e.g., city surroundings). In this paper, we mitigate these issues with: (i) a new residual pattern learning (RPL) module that assists the segmentation model to detect OoD pixels without affecting the inlier segmentation performance; and (ii) a novel context-robust contrastive learning (CoroCL) that enforces RPL to robustly detect OoD pixels among various contexts. Our approach improves by around 10\% FPR and 7\% AuPRC the previous state-of-the-art in Fishyscapes, Segment-Me-If-You-Can, and RoadAnomaly datasets. Our code is available at: https://github.com/yyliu01/RPL.
Knowing What to Label for Few Shot Microscopy Image Cell Segmentation
Nov 18, 2022



Abstract:In microscopy image cell segmentation, it is common to train a deep neural network on source data, containing different types of microscopy images, and then fine-tune it using a support set comprising a few randomly selected and annotated training target images. In this paper, we argue that the random selection of unlabelled training target images to be annotated and included in the support set may not enable an effective fine-tuning process, so we propose a new approach to optimise this image selection process. Our approach involves a new scoring function to find informative unlabelled target images. In particular, we propose to measure the consistency in the model predictions on target images against specific data augmentations. However, we observe that the model trained with source datasets does not reliably evaluate consistency on target images. To alleviate this problem, we propose novel self-supervised pretext tasks to compute the scores of unlabelled target images. Finally, the top few images with the least consistency scores are added to the support set for oracle (i.e., expert) annotation and later used to fine-tune the model to the target images. In our evaluations that involve the segmentation of five different types of cell images, we demonstrate promising results on several target test sets compared to the random selection approach as well as other selection approaches, such as Shannon's entropy and Monte-Carlo dropout.
Bootstrapping the Relationship Between Images and Their Clean and Noisy Labels
Oct 17, 2022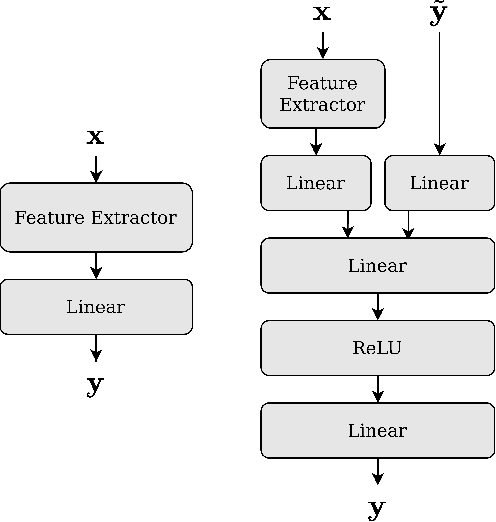
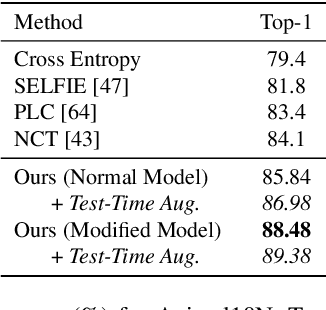
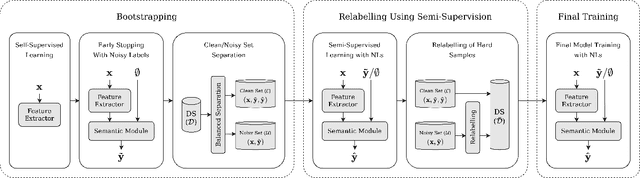
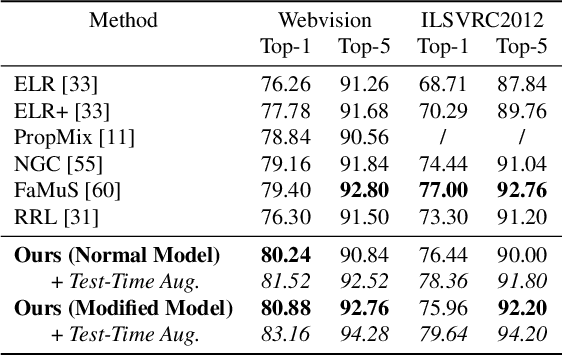
Abstract:Many state-of-the-art noisy-label learning methods rely on learning mechanisms that estimate the samples' clean labels during training and discard their original noisy labels. However, this approach prevents the learning of the relationship between images, noisy labels and clean labels, which has been shown to be useful when dealing with instance-dependent label noise problems. Furthermore, methods that do aim to learn this relationship require cleanly annotated subsets of data, as well as distillation or multi-faceted models for training. In this paper, we propose a new training algorithm that relies on a simple model to learn the relationship between clean and noisy labels without the need for a cleanly labelled subset of data. Our algorithm follows a 3-stage process, namely: 1) self-supervised pre-training followed by an early-stopping training of the classifier to confidently predict clean labels for a subset of the training set; 2) use the clean set from stage (1) to bootstrap the relationship between images, noisy labels and clean labels, which we exploit for effective relabelling of the remaining training set using semi-supervised learning; and 3) supervised training of the classifier with all relabelled samples from stage (2). By learning this relationship, we achieve state-of-the-art performance in asymmetric and instance-dependent label noise problems.
Knowledge Distillation to Ensemble Global and Interpretable Prototype-Based Mammogram Classification Models
Sep 26, 2022Abstract:State-of-the-art (SOTA) deep learning mammogram classifiers, trained with weakly-labelled images, often rely on global models that produce predictions with limited interpretability, which is a key barrier to their successful translation into clinical practice. On the other hand, prototype-based models improve interpretability by associating predictions with training image prototypes, but they are less accurate than global models and their prototypes tend to have poor diversity. We address these two issues with the proposal of BRAIxProtoPNet++, which adds interpretability to a global model by ensembling it with a prototype-based model. BRAIxProtoPNet++ distills the knowledge of the global model when training the prototype-based model with the goal of increasing the classification accuracy of the ensemble. Moreover, we propose an approach to increase prototype diversity by guaranteeing that all prototypes are associated with different training images. Experiments on weakly-labelled private and public datasets show that BRAIxProtoPNet++ has higher classification accuracy than SOTA global and prototype-based models. Using lesion localisation to assess model interpretability, we show BRAIxProtoPNet++ is more effective than other prototype-based models and post-hoc explanation of global models. Finally, we show that the diversity of the prototypes learned by BRAIxProtoPNet++ is superior to SOTA prototype-based approaches.
Multi-view Local Co-occurrence and Global Consistency Learning Improve Mammogram Classification Generalisation
Sep 21, 2022Abstract:When analysing screening mammograms, radiologists can naturally process information across two ipsilateral views of each breast, namely the cranio-caudal (CC) and mediolateral-oblique (MLO) views. These multiple related images provide complementary diagnostic information and can improve the radiologist's classification accuracy. Unfortunately, most existing deep learning systems, trained with globally-labelled images, lack the ability to jointly analyse and integrate global and local information from these multiple views. By ignoring the potentially valuable information present in multiple images of a screening episode, one limits the potential accuracy of these systems. Here, we propose a new multi-view global-local analysis method that mimics the radiologist's reading procedure, based on a global consistency learning and local co-occurrence learning of ipsilateral views in mammograms. Extensive experiments show that our model outperforms competing methods, in terms of classification accuracy and generalisation, on a large-scale private dataset and two publicly available datasets, where models are exclusively trained and tested with global labels.
On the Optimal Combination of Cross-Entropy and Soft Dice Losses for Lesion Segmentation with Out-of-Distribution Robustness
Sep 14, 2022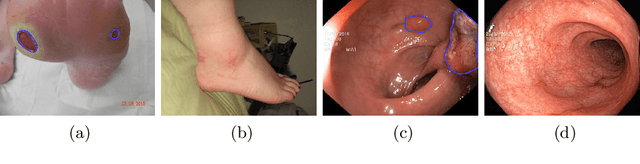
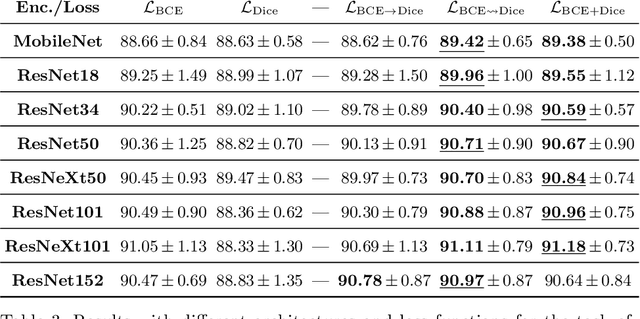
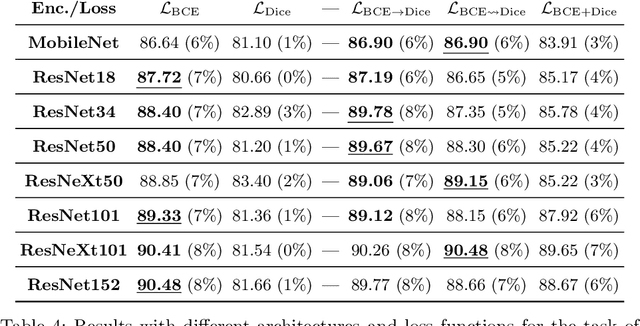
Abstract:We study the impact of different loss functions on lesion segmentation from medical images. Although the Cross-Entropy (CE) loss is the most popular option when dealing with natural images, for biomedical image segmentation the soft Dice loss is often preferred due to its ability to handle imbalanced scenarios. On the other hand, the combination of both functions has also been successfully applied in this kind of tasks. A much less studied problem is the generalization ability of all these losses in the presence of Out-of-Distribution (OoD) data. This refers to samples appearing in test time that are drawn from a different distribution than training images. In our case, we train our models on images that always contain lesions, but in test time we also have lesion-free samples. We analyze the impact of the minimization of different loss functions on in-distribution performance, but also its ability to generalize to OoD data, via comprehensive experiments on polyp segmentation from endoscopic images and ulcer segmentation from diabetic feet images. Our findings are surprising: CE-Dice loss combinations that excel in segmenting in-distribution images have a poor performance when dealing with OoD data, which leads us to recommend the adoption of the CE loss for this kind of problems, due to its robustness and ability to generalize to OoD samples. Code associated to our experiments can be found at https://github.com/agaldran/lesion_losses_ood .
Instance-Dependent Noisy Label Learning via Graphical Modelling
Sep 02, 2022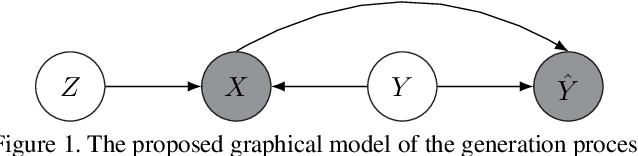
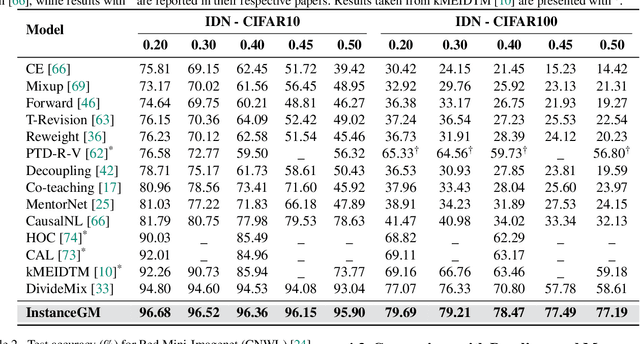
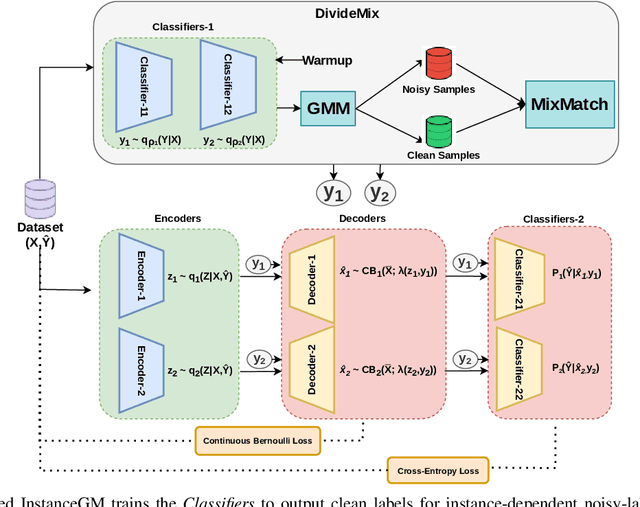
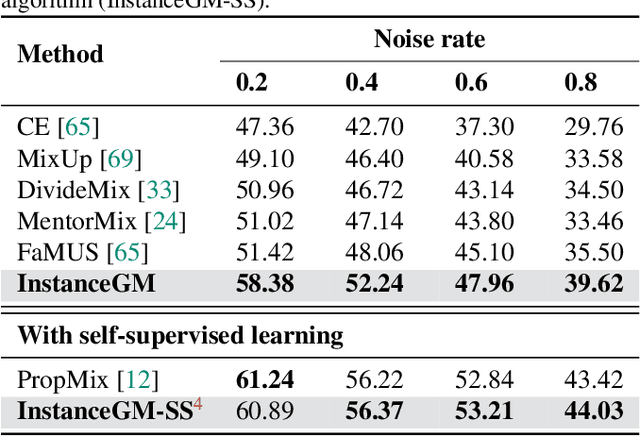
Abstract:Noisy labels are unavoidable yet troublesome in the ecosystem of deep learning because models can easily overfit them. There are many types of label noise, such as symmetric, asymmetric and instance-dependent noise (IDN), with IDN being the only type that depends on image information. Such dependence on image information makes IDN a critical type of label noise to study, given that labelling mistakes are caused in large part by insufficient or ambiguous information about the visual classes present in images. Aiming to provide an effective technique to address IDN, we present a new graphical modelling approach called InstanceGM, that combines discriminative and generative models. The main contributions of InstanceGM are: i) the use of the continuous Bernoulli distribution to train the generative model, offering significant training advantages, and ii) the exploration of a state-of-the-art noisy-label discriminative classifier to generate clean labels from instance-dependent noisy-label samples. InstanceGM is competitive with current noisy-label learning approaches, particularly in IDN benchmarks using synthetic and real-world datasets, where our method shows better accuracy than the competitors in most experiments.
Maximising the Utility of Validation Sets for Imbalanced Noisy-label Meta-learning
Aug 27, 2022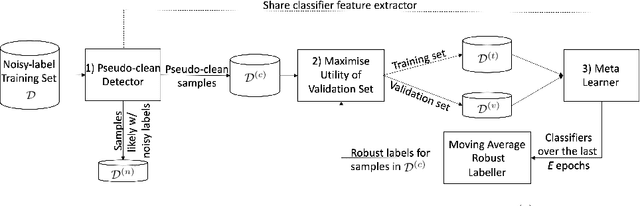
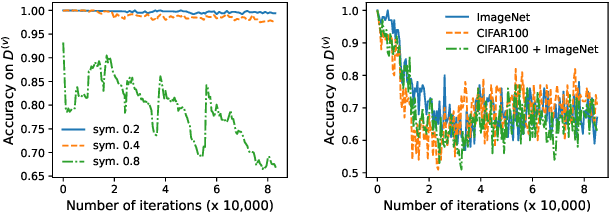
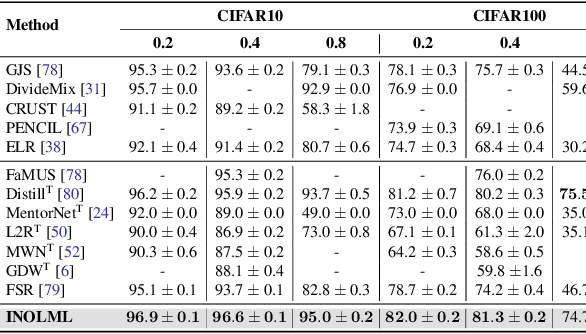
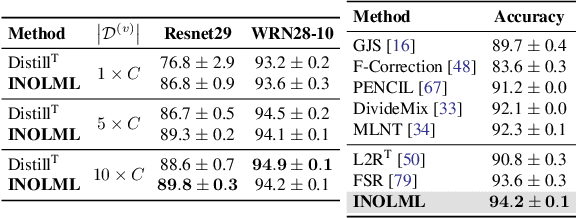
Abstract:Meta-learning is an effective method to handle imbalanced and noisy-label learning, but it depends on a validation set containing randomly selected, manually labelled and balanced distributed samples. The random selection and manual labelling and balancing of this validation set is not only sub-optimal for meta-learning, but it also scales poorly with the number of classes. Hence, recent meta-learning papers have proposed ad-hoc heuristics to automatically build and label this validation set, but these heuristics are still sub-optimal for meta-learning. In this paper, we analyse the meta-learning algorithm and propose new criteria to characterise the utility of the validation set, based on: 1) the informativeness of the validation set; 2) the class distribution balance of the set; and 3) the correctness of the labels of the set. Furthermore, we propose a new imbalanced noisy-label meta-learning (INOLML) algorithm that automatically builds a validation set by maximising its utility using the criteria above. Our method shows significant improvements over previous meta-learning approaches and sets the new state-of-the-art on several benchmarks.
A Study on the Impact of Data Augmentation for Training Convolutional Neural Networks in the Presence of Noisy Labels
Aug 23, 2022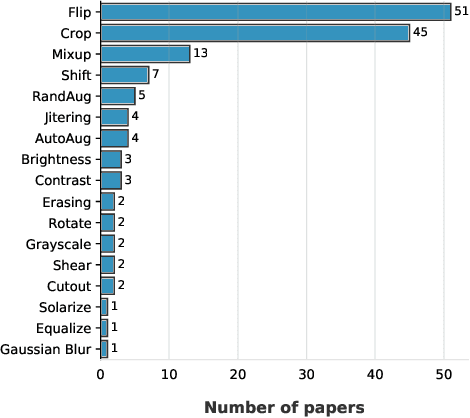
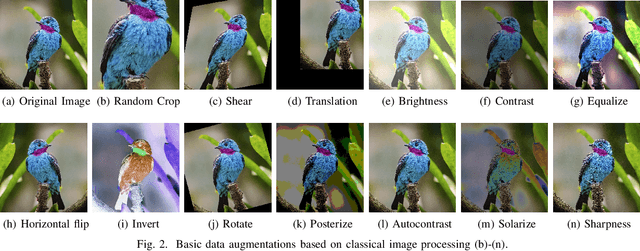
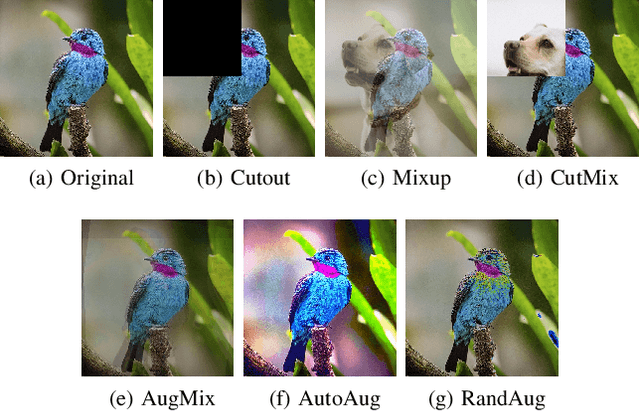


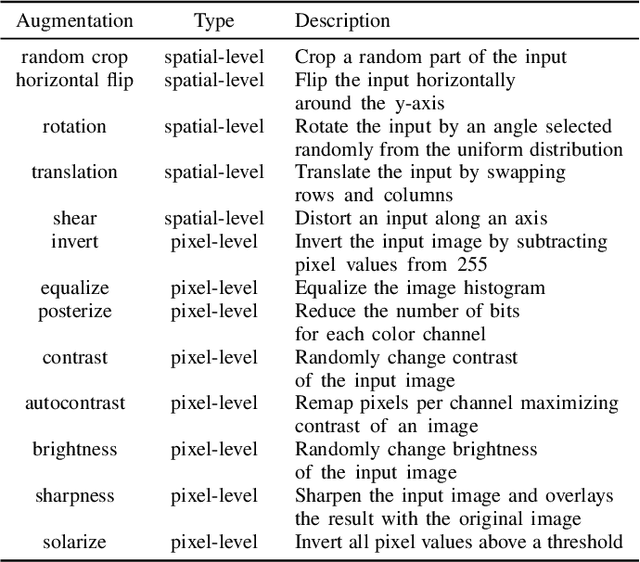
Abstract:Label noise is common in large real-world datasets, and its presence harms the training process of deep neural networks. Although several works have focused on the training strategies to address this problem, there are few studies that evaluate the impact of data augmentation as a design choice for training deep neural networks. In this work, we analyse the model robustness when using different data augmentations and their improvement on the training with the presence of noisy labels. We evaluate state-of-the-art and classical data augmentation strategies with different levels of synthetic noise for the datasets MNist, CIFAR-10, CIFAR-100, and the real-world dataset Clothing1M. We evaluate the methods using the accuracy metric. Results show that the appropriate selection of data augmentation can drastically improve the model robustness to label noise, increasing up to 177.84% of relative best test accuracy compared to the baseline with no augmentation, and an increase of up to 6% in absolute value with the state-of-the-art DivideMix training strategy.
An Evolutionary Approach for Creating of Diverse Classifier Ensembles
Aug 23, 2022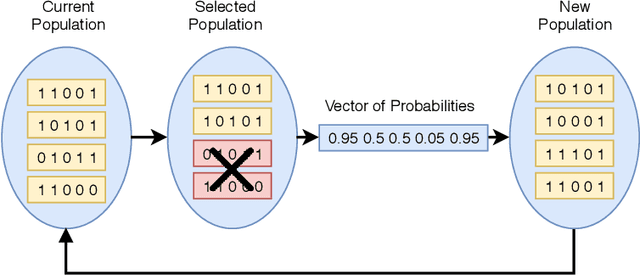

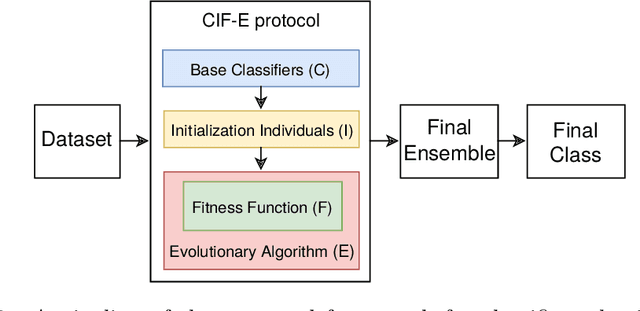
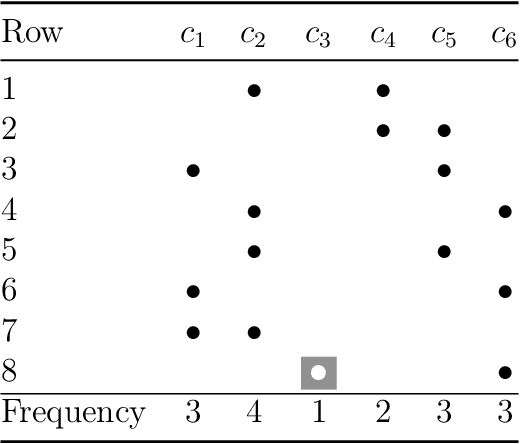
Abstract:Classification is one of the most studied tasks in data mining and machine learning areas and many works in the literature have been presented to solve classification problems for multiple fields of knowledge such as medicine, biology, security, and remote sensing. Since there is no single classifier that achieves the best results for all kinds of applications, a good alternative is to adopt classifier fusion strategies. A key point in the success of classifier fusion approaches is the combination of diversity and accuracy among classifiers belonging to an ensemble. With a large amount of classification models available in the literature, one challenge is the choice of the most suitable classifiers to compose the final classification system, which generates the need of classifier selection strategies. We address this point by proposing a framework for classifier selection and fusion based on a four-step protocol called CIF-E (Classifiers, Initialization, Fitness function, and Evolutionary algorithm). We implement and evaluate 24 varied ensemble approaches following the proposed CIF-E protocol and we are able to find the most accurate approach. A comparative analysis has also been performed among the best approaches and many other baselines from the literature. The experiments show that the proposed evolutionary approach based on Univariate Marginal Distribution Algorithm (UMDA) can outperform the state-of-the-art literature approaches in many well-known UCI datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge