Felix Grün
Error-related Potential driven Reinforcement Learning for adaptive Brain-Computer Interfaces
Feb 25, 2025



Abstract:Brain-computer interfaces (BCIs) provide alternative communication methods for individuals with motor disabilities by allowing control and interaction with external devices. Non-invasive BCIs, especially those using electroencephalography (EEG), are practical and safe for various applications. However, their performance is often hindered by EEG non-stationarities, caused by changing mental states or device characteristics like electrode impedance. This challenge has spurred research into adaptive BCIs that can handle such variations. In recent years, interest has grown in using error-related potentials (ErrPs) to enhance BCI performance. ErrPs, neural responses to errors, can be detected non-invasively and have been integrated into different BCI paradigms to improve performance through error correction or adaptation. This research introduces a novel adaptive ErrP-based BCI approach using reinforcement learning (RL). We demonstrate the feasibility of an RL-driven adaptive framework incorporating ErrPs and motor imagery. Utilizing two RL agents, the framework adapts dynamically to EEG non-stationarities. Validation was conducted using a publicly available motor imagery dataset and a fast-paced game designed to boost user engagement. Results show the framework's promise, with RL agents learning control policies from user interactions and achieving robust performance across datasets. However, a critical insight from the game-based protocol revealed that motor imagery in a high-speed interaction paradigm was largely ineffective for participants, highlighting task design limitations in real-time BCI applications. These findings underscore the potential of RL for adaptive BCIs while pointing out practical constraints related to task complexity and user responsiveness.
Towards Scenario- and Capability-Driven Dataset Development and Evaluation: An Approach in the Context of Mapless Automated Driving
Apr 30, 2024


Abstract:The foundational role of datasets in defining the capabilities of deep learning models has led to their rapid proliferation. At the same time, published research focusing on the process of dataset development for environment perception in automated driving has been scarce, thereby reducing the applicability of openly available datasets and impeding the development of effective environment perception systems. Sensor-based, mapless automated driving is one of the contexts where this limitation is evident. While leveraging real-time sensor data, instead of pre-defined HD maps promises enhanced adaptability and safety by effectively navigating unexpected environmental changes, it also increases the demands on the scope and complexity of the information provided by the perception system. To address these challenges, we propose a scenario- and capability-based approach to dataset development. Grounded in the principles of ISO 21448 (safety of the intended functionality, SOTIF), extended by ISO/TR 4804, our approach facilitates the structured derivation of dataset requirements. This not only aids in the development of meaningful new datasets but also enables the effective comparison of existing ones. Applying this methodology to a broad range of existing lane detection datasets, we identify significant limitations in current datasets, particularly in terms of real-world applicability, a lack of labeling of critical features, and an absence of comprehensive information for complex driving maneuvers.
Invariance to Quantile Selection in Distributional Continuous Control
Dec 29, 2022Abstract:In recent years distributional reinforcement learning has produced many state of the art results. Increasingly sample efficient Distributional algorithms for the discrete action domain have been developed over time that vary primarily in the way they parameterize their approximations of value distributions, and how they quantify the differences between those distributions. In this work we transfer three of the most well-known and successful of those algorithms (QR-DQN, IQN and FQF) to the continuous action domain by extending two powerful actor-critic algorithms (TD3 and SAC) with distributional critics. We investigate whether the relative performance of the methods for the discrete action space translates to the continuous case. To that end we compare them empirically on the pybullet implementations of a set of continuous control tasks. Our results indicate qualitative invariance regarding the number and placement of distributional atoms in the deterministic, continuous action setting.
Automatic Liver and Tumor Segmentation of CT and MRI Volumes using Cascaded Fully Convolutional Neural Networks
Feb 23, 2017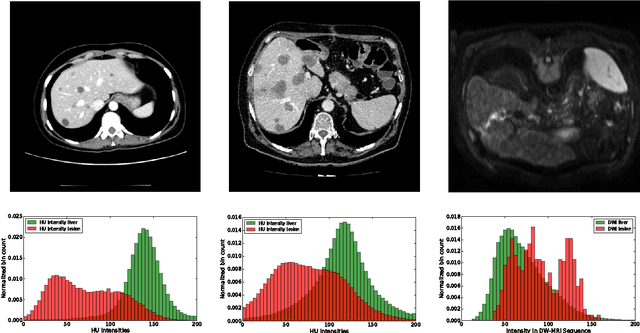
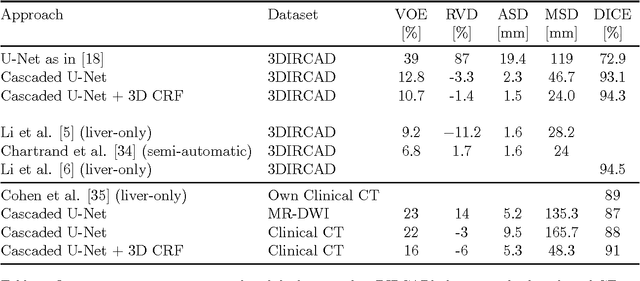
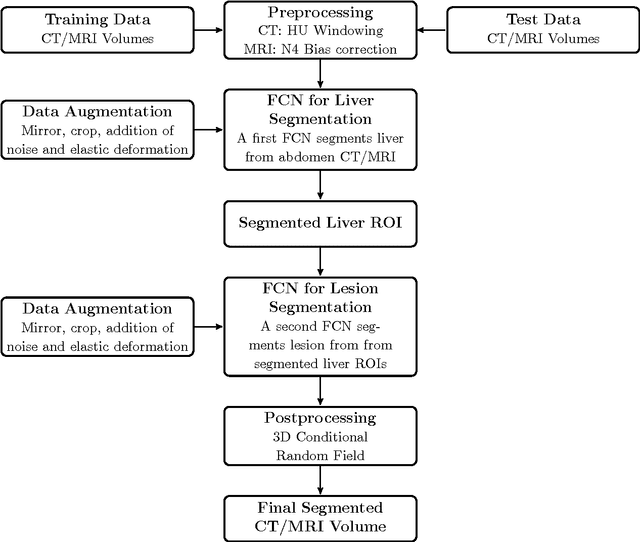
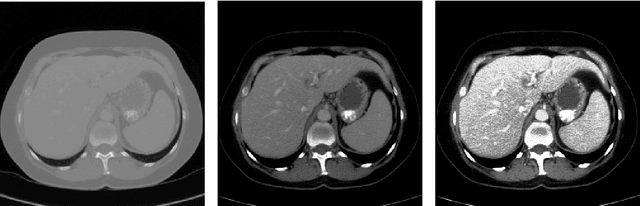
Abstract:Automatic segmentation of the liver and hepatic lesions is an important step towards deriving quantitative biomarkers for accurate clinical diagnosis and computer-aided decision support systems. This paper presents a method to automatically segment liver and lesions in CT and MRI abdomen images using cascaded fully convolutional neural networks (CFCNs) enabling the segmentation of a large-scale medical trial or quantitative image analysis. We train and cascade two FCNs for a combined segmentation of the liver and its lesions. In the first step, we train a FCN to segment the liver as ROI input for a second FCN. The second FCN solely segments lesions within the predicted liver ROIs of step 1. CFCN models were trained on an abdominal CT dataset comprising 100 hepatic tumor volumes. Validations on further datasets show that CFCN-based semantic liver and lesion segmentation achieves Dice scores over 94% for liver with computation times below 100s per volume. We further experimentally demonstrate the robustness of the proposed method on an 38 MRI liver tumor volumes and the public 3DIRCAD dataset.
SurvivalNet: Predicting patient survival from diffusion weighted magnetic resonance images using cascaded fully convolutional and 3D convolutional neural networks
Feb 20, 2017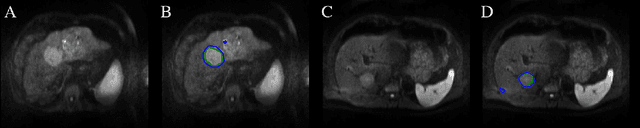
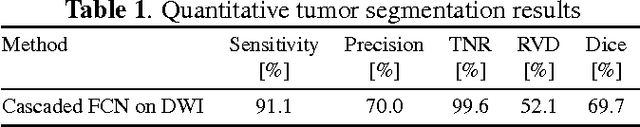

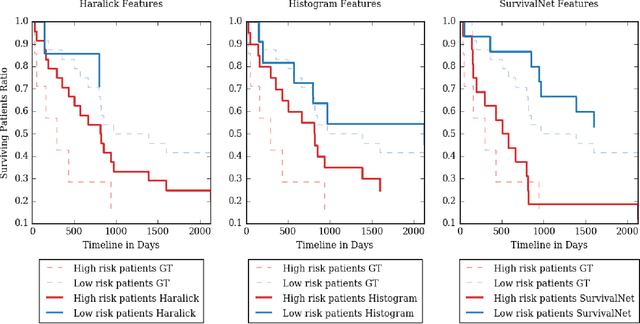
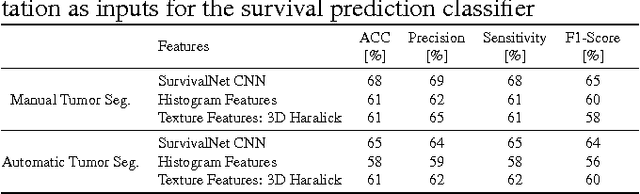
Abstract:Automatic non-invasive assessment of hepatocellular carcinoma (HCC) malignancy has the potential to substantially enhance tumor treatment strategies for HCC patients. In this work we present a novel framework to automatically characterize the malignancy of HCC lesions from DWI images. We predict HCC malignancy in two steps: As a first step we automatically segment HCC tumor lesions using cascaded fully convolutional neural networks (CFCN). A 3D neural network (SurvivalNet) then predicts the HCC lesions' malignancy from the HCC tumor segmentation. We formulate this task as a classification problem with classes being "low risk" and "high risk" represented by longer or shorter survival times than the median survival. We evaluated our method on DWI of 31 HCC patients. Our proposed framework achieves an end-to-end accuracy of 65% with a Dice score for the automatic lesion segmentation of 69% and an accuracy of 68% for tumor malignancy classification based on expert annotations. We compared the SurvivalNet to classical handcrafted features such as Histogram and Haralick and show experimentally that SurvivalNet outperforms the handcrafted features in HCC malignancy classification. End-to-end assessment of tumor malignancy based on our proposed fully automatic framework corresponds to assessment based on expert annotations with high significance (p>0.95).
A Taxonomy and Library for Visualizing Learned Features in Convolutional Neural Networks
Jun 24, 2016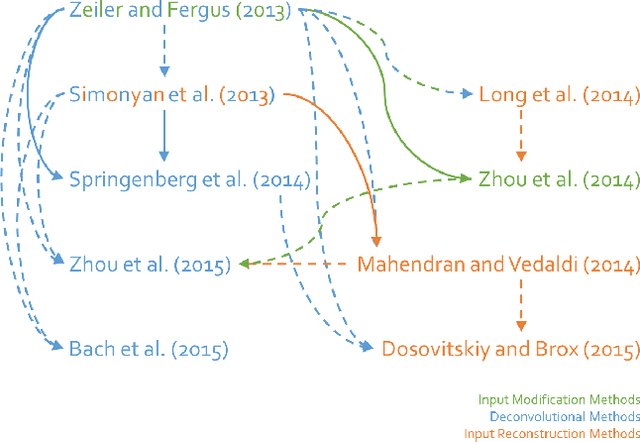
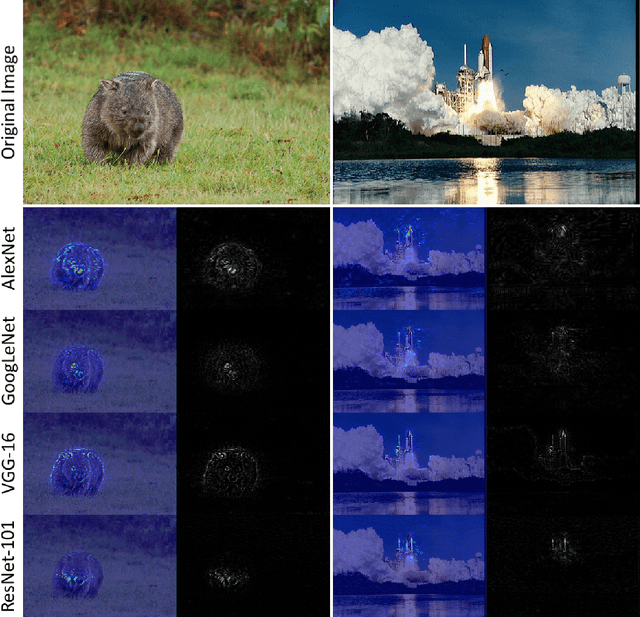
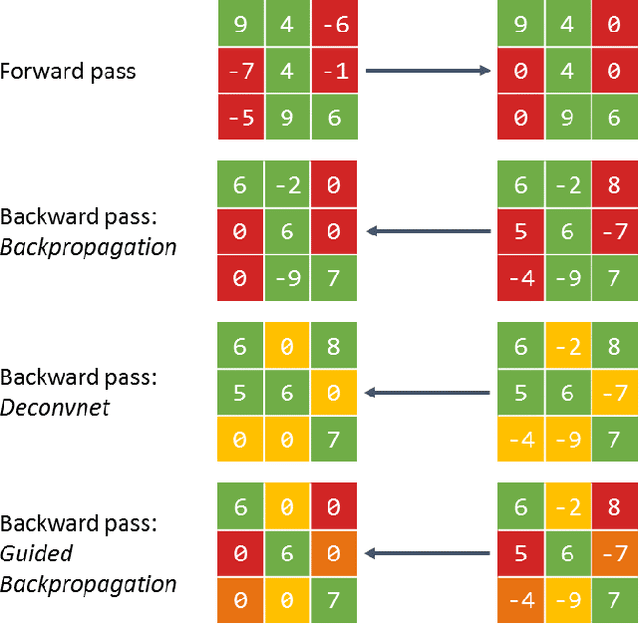
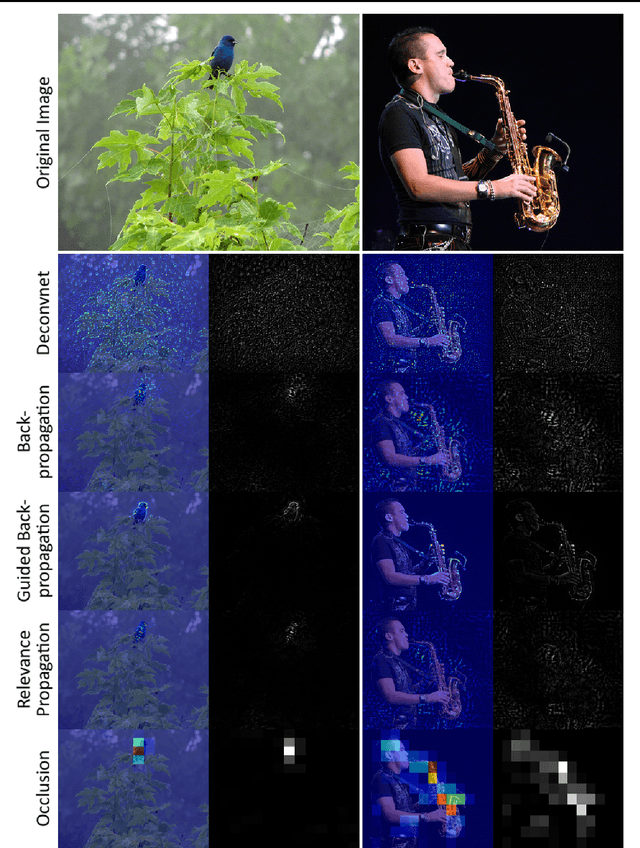

Abstract:Over the last decade, Convolutional Neural Networks (CNN) saw a tremendous surge in performance. However, understanding what a network has learned still proves to be a challenging task. To remedy this unsatisfactory situation, a number of groups have recently proposed different methods to visualize the learned models. In this work we suggest a general taxonomy to classify and compare these methods, subdividing the literature into three main categories and providing researchers with a terminology to base their works on. Furthermore, we introduce the FeatureVis library for MatConvNet: an extendable, easy to use open source library for visualizing CNNs. It contains implementations from each of the three main classes of visualization methods and serves as a useful tool for an enhanced understanding of the features learned by intermediate layers, as well as for the analysis of why a network might fail for certain examples.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge