Elgar Fleisch
Vocal Prognostic Digital Biomarkers in Monitoring Chronic Heart Failure: A Longitudinal Observational Study
Mar 31, 2026Abstract:Objective: This study aimed to evaluate which voice features can predict health deterioration in patients with chronic HF. Background: Heart failure (HF) is a chronic condition with progressive deterioration and acute decompensations, often requiring hospitalization and imposing substantial healthcare and economic burdens. Current standard-of-care (SoC) home monitoring, such as weight tracking, lacks predictive accuracy and requires high patient engagement. Voice is a promising non-invasive biomarker, though prior studies have mainly focused on acute HF stages. Methods: In a 2-month longitudinal study, 32 patients with HF collected daily voice recordings and SoC measures of weight and blood pressure at home, with biweekly questionnaires for health status. Acoustic analysis generated detailed vowel and speech features. Time-series features were extracted from aggregated lookback windows (e.g., 7 days) to predict next-day health status. Explainable machine learning with nested cross-validation identified top vocal biomarkers, and a case study illustrated model application. Results: A total of 21,863 recordings were analyzed. Acoustic vowel features showed strong correlations with health status. Time-series voice features within the lookback window outperformed corresponding standard care measures, achieving peak sensitivity and specificity of 0.826 and 0.782 versus 0.783 and 0.567 for SoC metrics. Key prognostic voice features identifying deterioration included delayed energy shift, low energy variability, and higher shimmer variability in vowels, along with reduced speaking and articulation rate, lower phonation ratio, decreased voice quality, and increased formant variability in speech. Conclusion: Voice-based monitoring offers a non-invasive approach to detect early health changes in chronic HF, supporting proactive and personalized care.
Patient-Level Multimodal Question Answering from Multi-Site Auscultation Recordings
Mar 09, 2026Abstract:Auscultation is a vital diagnostic tool, yet its utility is often limited by subjective interpretation. While general-purpose Audio-Language Models (ALMs) excel in general domains, they struggle with the nuances of physiological signals. We propose a framework that aligns multi-site auscultation recordings directly with a frozen Large Language Model (LLM) embedding space via gated cross-attention. By leveraging the LLM's latent world knowledge, our approach moves beyond isolated classification toward holistic, patient-level assessment. On the CaReSound benchmark, our model achieves a state-of-the-art 0.865 F1-macro and 0.952 BERTScore. We demonstrate that lightweight, domain-specific encoders rival large-scale ALMs and that multi-site aggregation provides spatial redundancy that mitigates temporal truncation. This alignment of medical acoustics with text foundations offers a scalable path for bridging signal processing and clinical assessment.
EvoMorph: Counterfactual Explanations for Continuous Time-Series Extrinsic Regression Applied to Photoplethysmography
Jan 15, 2026Abstract:Wearable devices enable continuous, population-scale monitoring of physiological signals, such as photoplethysmography (PPG), creating new opportunities for data-driven clinical assessment. Time-series extrinsic regression (TSER) models increasingly leverage PPG signals to estimate clinically relevant outcomes, including heart rate, respiratory rate, and oxygen saturation. For clinical reasoning and trust, however, single point estimates alone are insufficient: clinicians must also understand whether predictions are stable under physiologically plausible variations and to what extent realistic, attainable changes in physiological signals would meaningfully alter a model's prediction. Counterfactual explanations (CFE) address these "what-if" questions, yet existing time series CFE generation methods are largely restricted to classification, overlook waveform morphology, and often produce physiologically implausible signals, limiting their applicability to continuous biomedical time series. To address these limitations, we introduce EvoMorph, a multi-objective evolutionary framework for generating physiologically plausible and diverse CFE for TSER applications. EvoMorph optimizes morphology-aware objectives defined on interpretable signal descriptors and applies transformations to preserve the waveform structure. We evaluated EvoMorph on three PPG datasets (heart rate, respiratory rate, and oxygen saturation) against a nearest-unlike-neighbor baseline. In addition, in a case study, we evaluated EvoMorph as a tool for uncertainty quantification by relating counterfactual sensitivity to bootstrap-ensemble uncertainty and data-density measures. Overall, EvoMorph enables the generation of physiologically-aware counterfactuals for continuous biomedical signals and supports uncertainty-aware interpretability, advancing trustworthy model analysis for clinical time-series applications.
Comparative Efficacy of Commercial Wearables for Circadian Rhythm Home Monitoring from Activity, Heart Rate, and Core Body Temperature
Apr 04, 2024



Abstract:Circadian rhythms govern biological patterns that follow a 24-hour cycle. Dysfunctions in circadian rhythms can contribute to various health problems, such as sleep disorders. Current circadian rhythm assessment methods, often invasive or subjective, limit circadian rhythm monitoring to laboratories. Hence, this study aims to investigate scalable consumer-centric wearables for circadian rhythm monitoring outside traditional laboratories. In a two-week longitudinal study conducted in real-world settings, 36 participants wore an Actigraph, a smartwatch, and a core body temperature sensor to collect activity, temperature, and heart rate data. We evaluated circadian rhythms calculated from commercial wearables by comparing them with circadian rhythm reference measures, i.e., Actigraph activities and chronotype questionnaire scores. The circadian rhythm metric acrophases, determined from commercial wearables using activity, heart rate, and temperature data, significantly correlated with the acrophase derived from Actigraph activities (r=0.96, r=0.87, r=0.79; all p<0.001) and chronotype questionnaire (r=-0.66, r=-0.73, r=-0.61; all p<0.001). The acrophases obtained concurrently from consumer sensors significantly predicted the chronotype (R2=0.64; p<0.001). Our study validates commercial sensors for circadian rhythm assessment, highlighting their potential to support maintaining healthy rhythms and provide scalable and timely health monitoring in real-life scenarios.
"Are you okay, honey?": Recognizing Emotions among Couples Managing Diabetes in Daily Life using Multimodal Real-World Smartwatch Data
Aug 22, 2022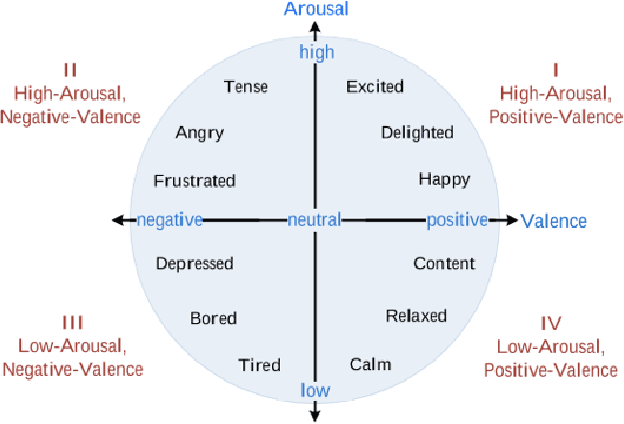
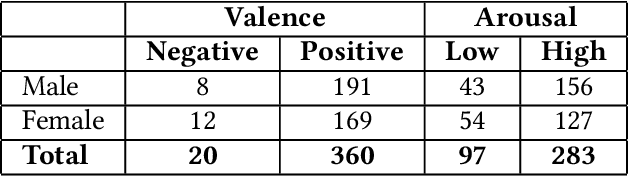
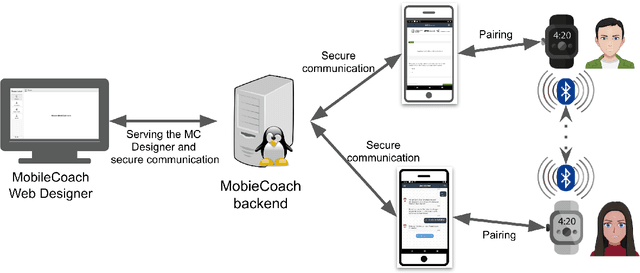
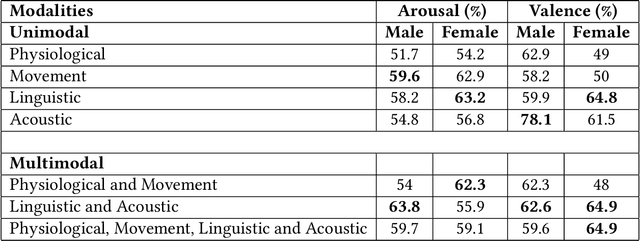
Abstract:Couples generally manage chronic diseases together and the management takes an emotional toll on both patients and their romantic partners. Consequently, recognizing the emotions of each partner in daily life could provide an insight into their emotional well-being in chronic disease management. Currently, the process of assessing each partner's emotions is manual, time-intensive, and costly. Despite the existence of works on emotion recognition among couples, none of these works have used data collected from couples' interactions in daily life. In this work, we collected 85 hours (1,021 5-minute samples) of real-world multimodal smartwatch sensor data (speech, heart rate, accelerometer, and gyroscope) and self-reported emotion data (n=612) from 26 partners (13 couples) managing diabetes mellitus type 2 in daily life. We extracted physiological, movement, acoustic, and linguistic features, and trained machine learning models (support vector machine and random forest) to recognize each partner's self-reported emotions (valence and arousal). Our results from the best models (balanced accuracies of 63.8% and 78.1% for arousal and valence respectively) are better than chance and our prior work that also used data from German-speaking, Swiss-based couples, albeit, in the lab. This work contributes toward building automated emotion recognition systems that would eventually enable partners to monitor their emotions in daily life and enable the delivery of interventions to improve their emotional well-being.
Driver Identification via the Steering Wheel
Sep 09, 2019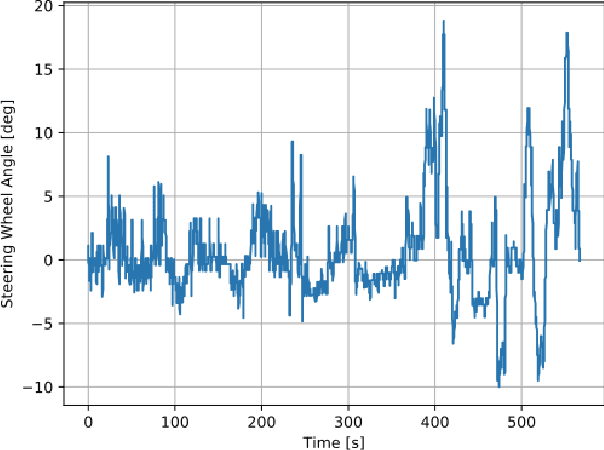
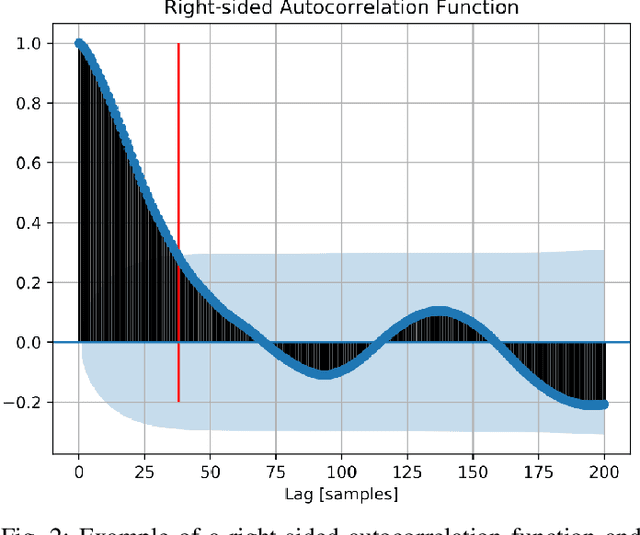
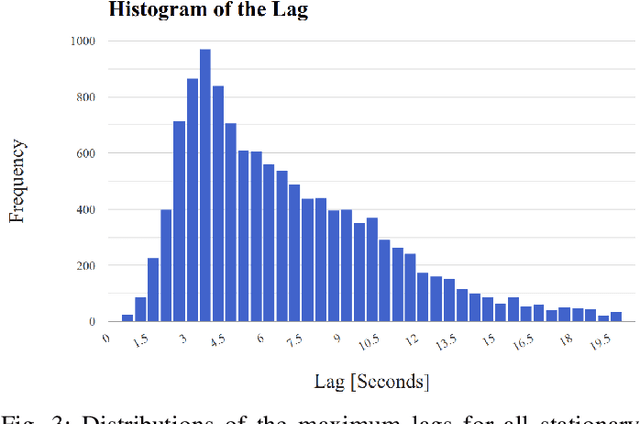
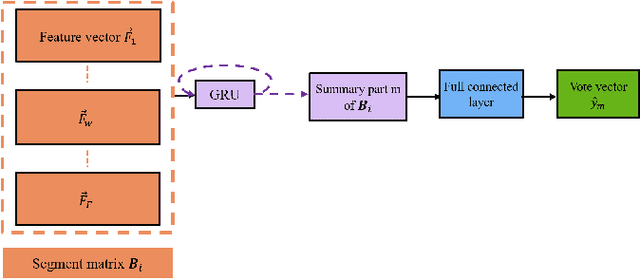
Abstract:Driver identification has emerged as a vital research field, where both practitioners and researchers investigate the potential of driver identification to enable a personalized driving experience. Within recent years, a selection of studies have reported that individuals could be perfectly identified based on their driving behavior under controlled conditions. However, research investigating the potential of driver identification under naturalistic conditions claim accuracies only marginally higher than random guess. The paper at hand provides a comprehensive summary of the recent work, highlighting the main discrepancies in the design of the machine learning approaches, primarily the window length parameter that was considered. Key findings further indicate that the longitudinal vehicle control information is particularly useful for driver identification, leaving the research gap on the extent to which the lateral vehicle control can be used for reliable identification. Building upon existing work, we provide a novel approach for the design of the window length parameter that provides evidence that reliable driver identification can be achieved with data limited to the steering wheel only. The results and insights in this paper are based on data collected from the largest naturalistic driving study conducted in this field. Overall, a neural network based on GRUs was found to provide better identification performance than traditional methods, increasing the prediction accuracy from under 15\% to over 65\% for 15 drivers. When leveraging the full field study dataset, comprising 72 drivers, the accuracy of identification prediction of the approach improved a random guess approach by a factor of 25.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge