Dina Demner-Fushman
Clinical Language Understanding Evaluation (CLUE)
Sep 28, 2022



Abstract:Clinical language processing has received a lot of attention in recent years, resulting in new models or methods for disease phenotyping, mortality prediction, and other tasks. Unfortunately, many of these approaches are tested under different experimental settings (e.g., data sources, training and testing splits, metrics, evaluation criteria, etc.) making it difficult to compare approaches and determine state-of-the-art. To address these issues and facilitate reproducibility and comparison, we present the Clinical Language Understanding Evaluation (CLUE) benchmark with a set of four clinical language understanding tasks, standard training, development, validation and testing sets derived from MIMIC data, as well as a software toolkit. It is our hope that these data will enable direct comparison between approaches, improve reproducibility, and reduce the barrier-to-entry for developing novel models or methods for these clinical language understanding tasks.
CHQ-Summ: A Dataset for Consumer Healthcare Question Summarization
Jun 15, 2022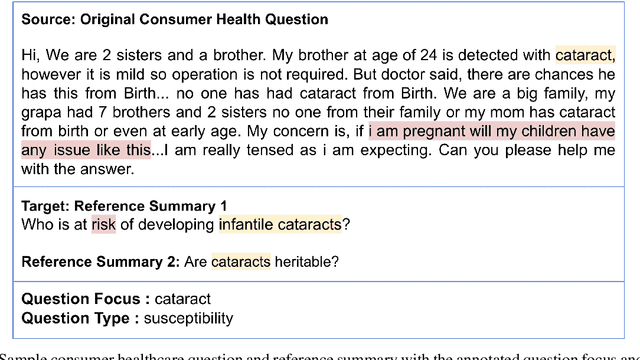
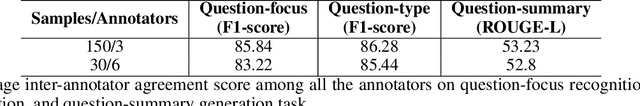
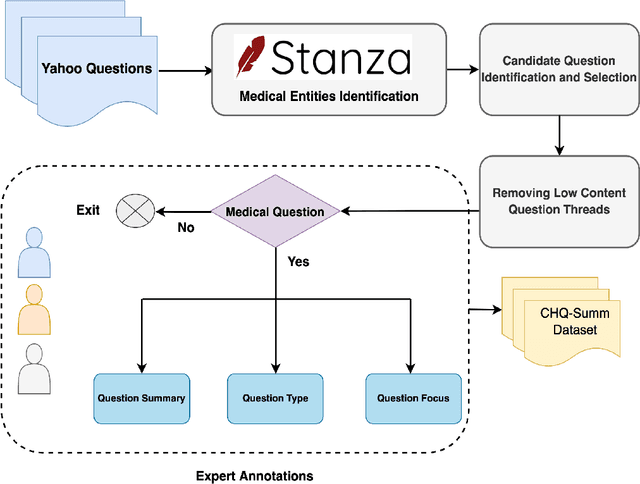
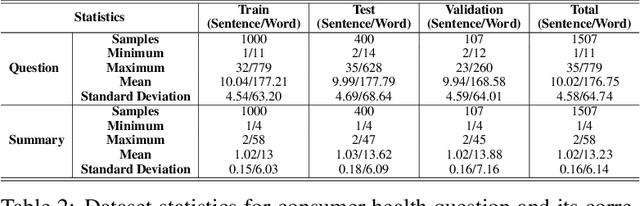
Abstract:The quest for seeking health information has swamped the web with consumers' health-related questions. Generally, consumers use overly descriptive and peripheral information to express their medical condition or other healthcare needs, contributing to the challenges of natural language understanding. One way to address this challenge is to summarize the questions and distill the key information of the original question. To address this issue, we introduce a new dataset, CHQ-Summ that contains 1507 domain-expert annotated consumer health questions and corresponding summaries. The dataset is derived from the community question-answering forum and therefore provides a valuable resource for understanding consumer health-related posts on social media. We benchmark the dataset on multiple state-of-the-art summarization models to show the effectiveness of the dataset.
A Dataset for Medical Instructional Video Classification and Question Answering
Jan 30, 2022Abstract:This paper introduces a new challenge and datasets to foster research toward designing systems that can understand medical videos and provide visual answers to natural language questions. We believe medical videos may provide the best possible answers to many first aids, medical emergency, and medical education questions. Toward this, we created the MedVidCL and MedVidQA datasets and introduce the tasks of Medical Video Classification (MVC) and Medical Visual Answer Localization (MVAL), two tasks that focus on cross-modal (medical language and medical video) understanding. The proposed tasks and datasets have the potential to support the development of sophisticated downstream applications that can benefit the public and medical practitioners. Our datasets consist of 6,117 annotated videos for the MVC task and 3,010 annotated questions and answers timestamps from 899 videos for the MVAL task. These datasets have been verified and corrected by medical informatics experts. We have also benchmarked each task with the created MedVidCL and MedVidQA datasets and proposed the multimodal learning methods that set competitive baselines for future research.
Reinforcement Learning for Abstractive Question Summarization with Question-aware Semantic Rewards
Jul 01, 2021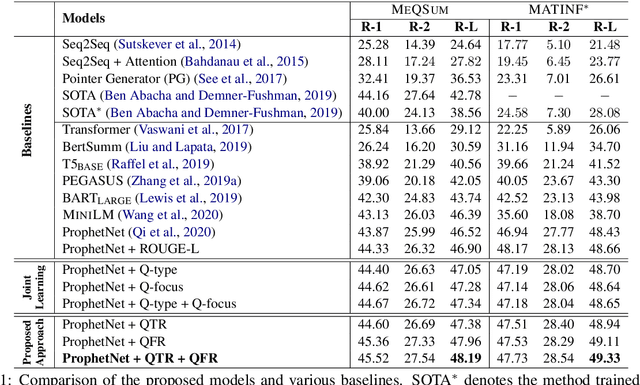
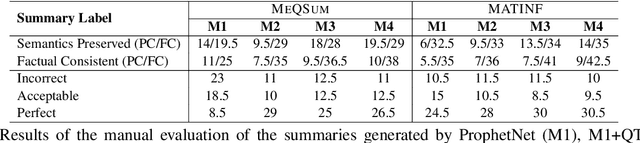

Abstract:The growth of online consumer health questions has led to the necessity for reliable and accurate question answering systems. A recent study showed that manual summarization of consumer health questions brings significant improvement in retrieving relevant answers. However, the automatic summarization of long questions is a challenging task due to the lack of training data and the complexity of the related subtasks, such as the question focus and type recognition. In this paper, we introduce a reinforcement learning-based framework for abstractive question summarization. We propose two novel rewards obtained from the downstream tasks of (i) question-type identification and (ii) question-focus recognition to regularize the question generation model. These rewards ensure the generation of semantically valid questions and encourage the inclusion of key medical entities/foci in the question summary. We evaluated our proposed method on two benchmark datasets and achieved higher performance over state-of-the-art models. The manual evaluation of the summaries reveals that the generated questions are more diverse and have fewer factual inconsistencies than the baseline summaries
Question-aware Transformer Models for Consumer Health Question Summarization
Jun 01, 2021
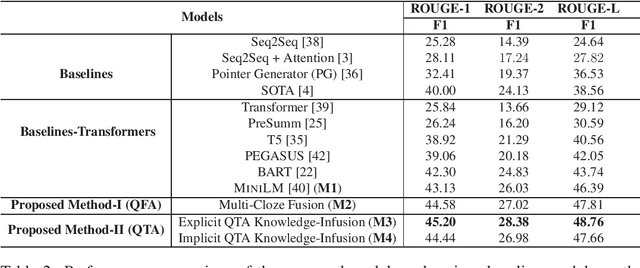

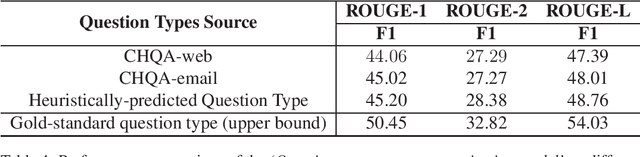
Abstract:Searching for health information online is becoming customary for more and more consumers every day, which makes the need for efficient and reliable question answering systems more pressing. An important contributor to the success rates of these systems is their ability to fully understand the consumers' questions. However, these questions are frequently longer than needed and mention peripheral information that is not useful in finding relevant answers. Question summarization is one of the potential solutions to simplifying long and complex consumer questions before attempting to find an answer. In this paper, we study the task of abstractive summarization for real-world consumer health questions. We develop an abstractive question summarization model that leverages the semantic interpretation of a question via recognition of medical entities, which enables the generation of informative summaries. Towards this, we propose multiple Cloze tasks (i.e. the task of filing missing words in a given context) to identify the key medical entities that enforce the model to have better coverage in question-focus recognition. Additionally, we infuse the decoder inputs with question-type information to generate question-type driven summaries. When evaluated on the MeQSum benchmark corpus, our framework outperformed the state-of-the-art method by 10.2 ROUGE-L points. We also conducted a manual evaluation to assess the correctness of the generated summaries.
Searching for Scientific Evidence in a Pandemic: An Overview of TREC-COVID
Apr 19, 2021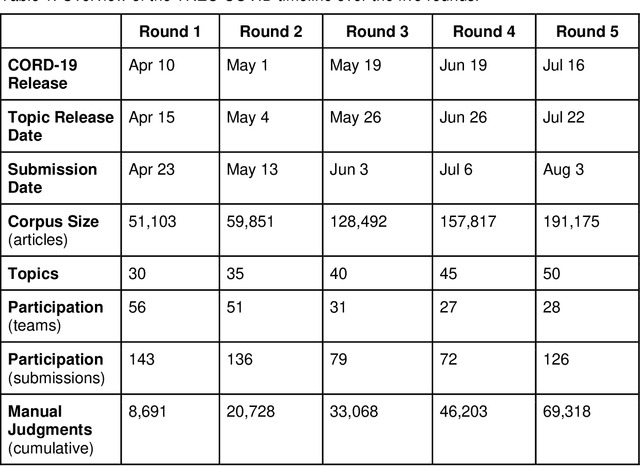
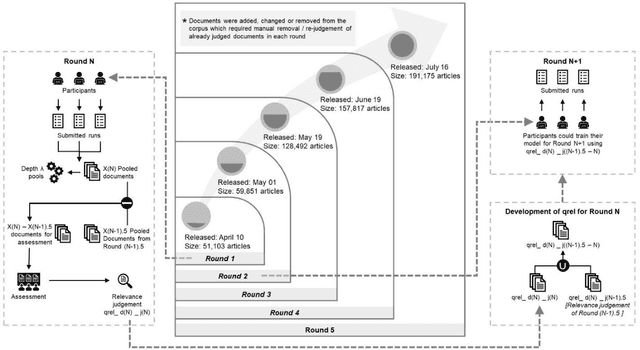

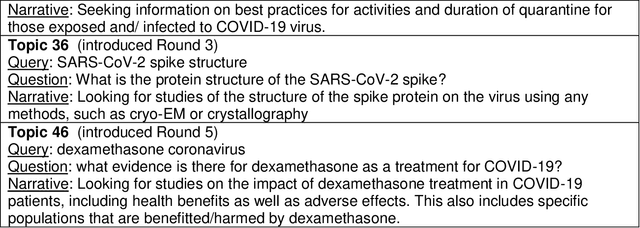
Abstract:We present an overview of the TREC-COVID Challenge, an information retrieval (IR) shared task to evaluate search on scientific literature related to COVID-19. The goals of TREC-COVID include the construction of a pandemic search test collection and the evaluation of IR methods for COVID-19. The challenge was conducted over five rounds from April to July, 2020, with participation from 92 unique teams and 556 individual submissions. A total of 50 topics (sets of related queries) were used in the evaluation, starting at 30 topics for Round 1 and adding 5 new topics per round to target emerging topics at that state of the still-emerging pandemic. This paper provides a comprehensive overview of the structure and results of TREC-COVID. Specifically, the paper provides details on the background, task structure, topic structure, corpus, participation, pooling, assessment, judgments, results, top-performing systems, lessons learned, and benchmark datasets.
Question-Driven Summarization of Answers to Consumer Health Questions
May 20, 2020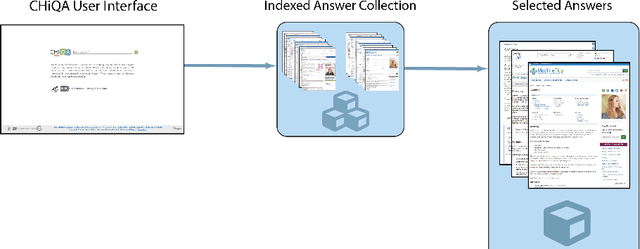
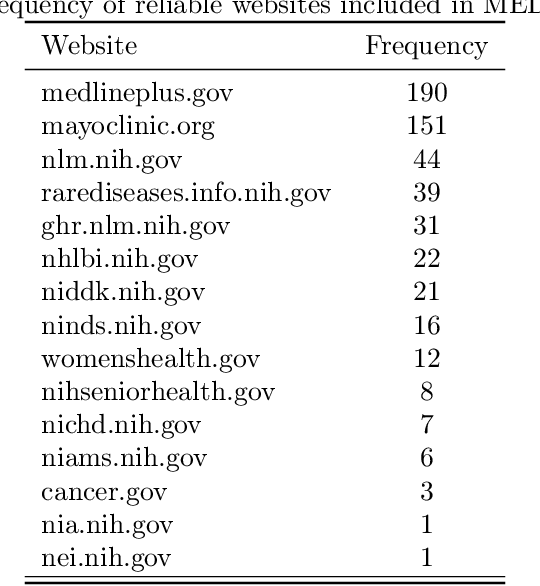
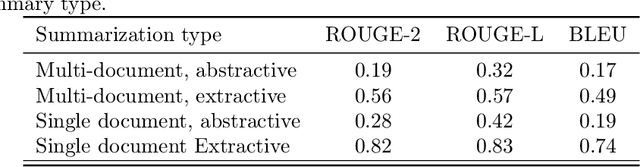
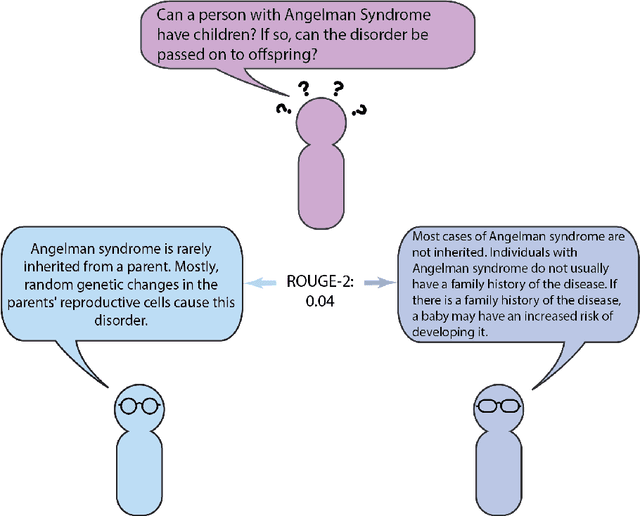
Abstract:Automatic summarization of natural language is a widely studied area in computer science, one that is broadly applicable to anyone who routinely needs to understand large quantities of information. For example, in the medical domain, recent developments in deep learning approaches to automatic summarization have the potential to make health information more easily accessible to patients and consumers. However, to evaluate the quality of automatically generated summaries of health information, gold-standard, human generated summaries are required. Using answers provided by the National Library of Medicine's consumer health question answering system, we present the MEDIQA Answer Summarization dataset, the first summarization collection containing question-driven summaries of answers to consumer health questions. This dataset can be used to evaluate single or multi-document summaries generated by algorithms using extractive or abstractive approaches. In order to benchmark the dataset, we include results of baseline and state-of-the-art deep learning summarization models, demonstrating that this dataset can be used to effectively evaluate question-driven machine-generated summaries and promote further machine learning research in medical question answering.
Bridging the Knowledge Gap: Enhancing Question Answering with World and Domain Knowledge
Oct 16, 2019
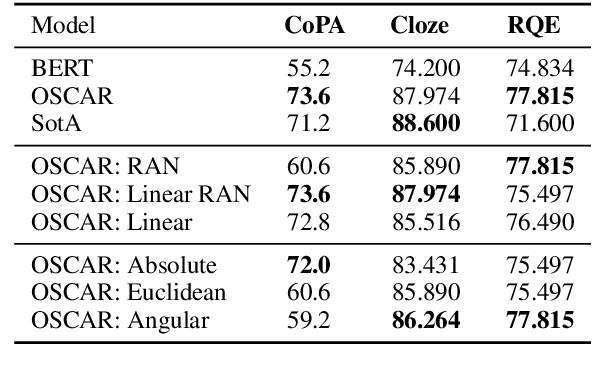
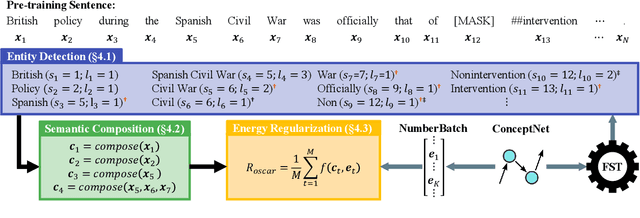
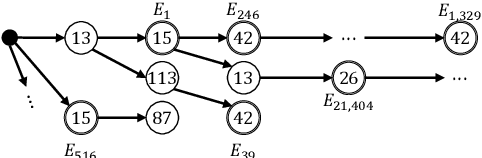
Abstract:In this paper we present OSCAR (Ontology-based Semantic Composition Augmented Regularization), a method for injecting task-agnostic knowledge from an Ontology or knowledge graph into a neural network during pretraining. We evaluated the impact of including OSCAR when pretraining BERT with Wikipedia articles by measuring the performance when fine-tuning on two question answering tasks involving world knowledge and causal reasoning and one requiring domain (healthcare) knowledge and obtained 33:3%, 18:6%, and 4% improved accuracy compared to pretraining BERT without OSCAR and obtaining new state-of-the-art results on two of the tasks.
Understanding Spatial Language in Radiology: Representation Framework, Annotation, and Spatial Relation Extraction from Chest X-ray Reports using Deep Learning
Aug 13, 2019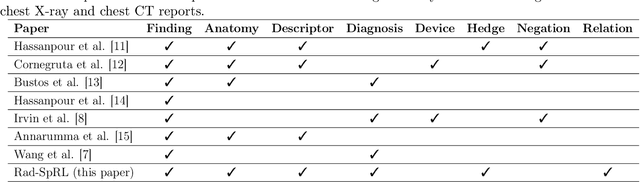
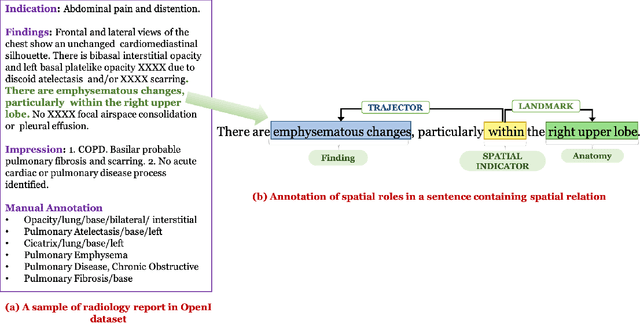

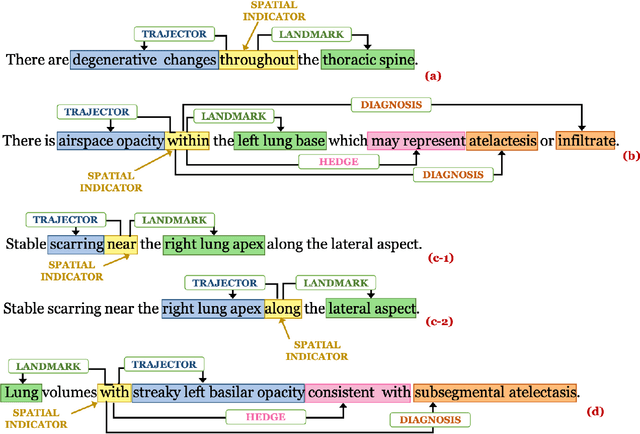
Abstract:We define a representation framework for extracting spatial information from radiology reports (Rad-SpRL). We annotated a total of 2000 chest X-ray reports with 4 spatial roles corresponding to the common radiology entities. Our focus is on extracting detailed information of a radiologist's interpretation containing a radiographic finding, its anatomical location, corresponding probable diagnoses, as well as associated hedging terms. For this, we propose a deep learning-based natural language processing (NLP) method involving both word and character-level encodings. Specifically, we utilize a bidirectional long short-term memory (Bi-LSTM) conditional random field (CRF) model for extracting the spatial roles. The model achieved average F1 measures of 90.28 and 94.61 for extracting the Trajector and Landmark roles respectively whereas the performance was moderate for Diagnosis and Hedge roles with average F1 of 71.47 and 73.27 respectively. The corpus will soon be made available upon request.
A Question-Entailment Approach to Question Answering
Jan 23, 2019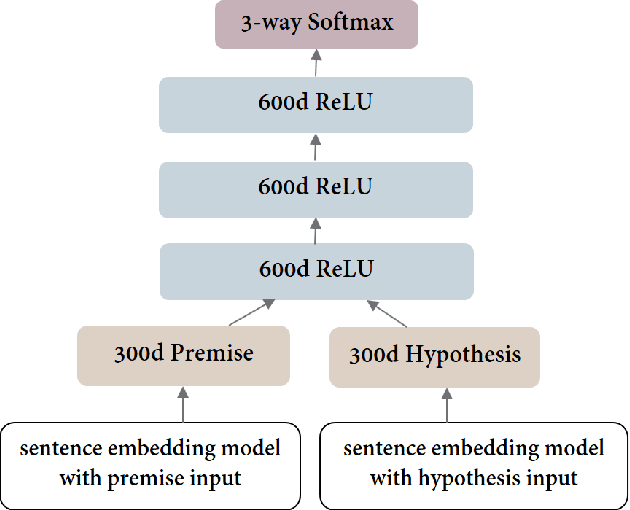
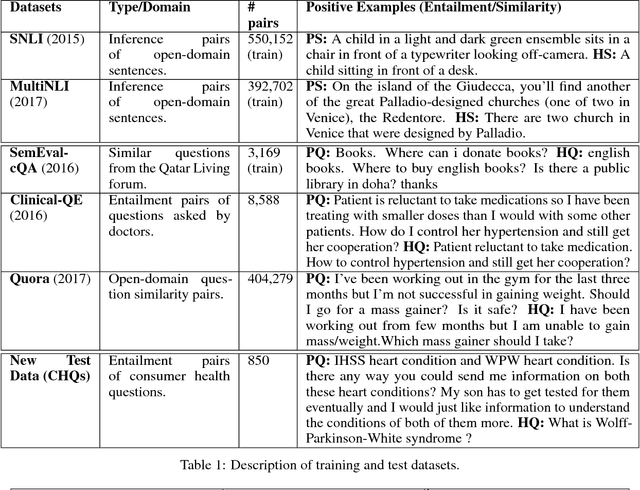
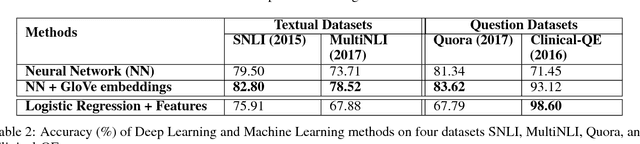

Abstract:One of the challenges in large-scale information retrieval (IR) is to develop fine-grained and domain-specific methods to answer natural language questions. Despite the availability of numerous sources and datasets for answer retrieval, Question Answering (QA) remains a challenging problem due to the difficulty of the question understanding and answer extraction tasks. One of the promising tracks investigated in QA is to map new questions to formerly answered questions that are `similar'. In this paper, we propose a novel QA approach based on Recognizing Question Entailment (RQE) and we describe the QA system and resources that we built and evaluated on real medical questions. First, we compare machine learning and deep learning methods for RQE using different kinds of datasets, including textual inference, question similarity and entailment in both the open and clinical domains. Second, we combine IR models with the best RQE method to select entailed questions and rank the retrieved answers. To study the end-to-end QA approach, we built the MedQuAD collection of 47,457 question-answer pairs from trusted medical sources, that we introduce and share in the scope of this paper. Following the evaluation process used in TREC 2017 LiveQA, we find that our approach exceeds the best results of the medical task with a 29.8% increase over the best official score. The evaluation results also support the relevance of question entailment for QA and highlight the effectiveness of combining IR and RQE for future QA efforts. Our findings also show that relying on a restricted set of reliable answer sources can bring a substantial improvement in medical QA.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge