Changhao Sun
OBBStacking: An Ensemble Method for Remote Sensing Object Detection
Sep 27, 2022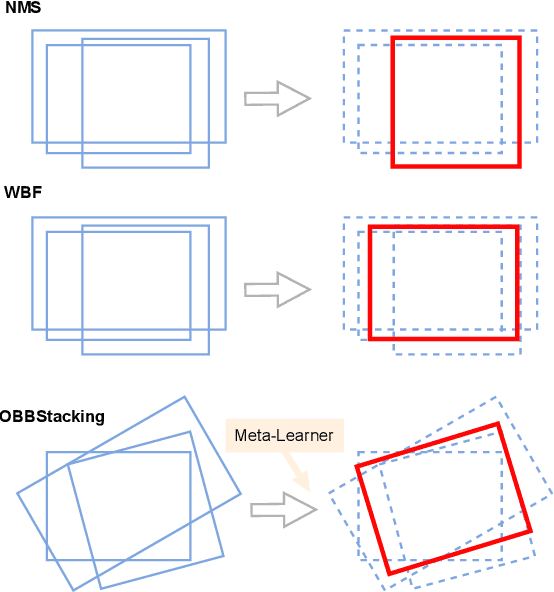
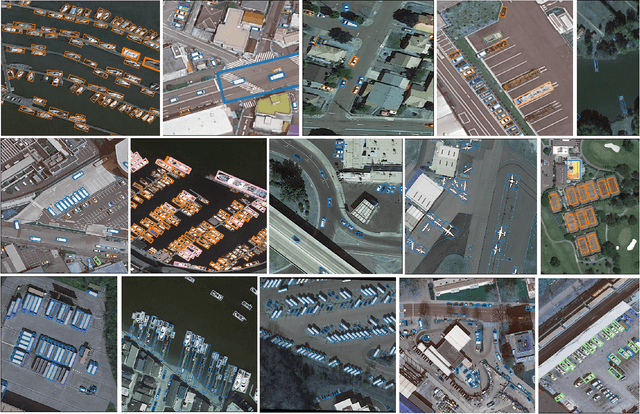
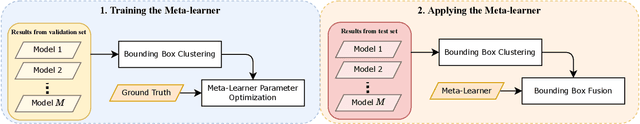

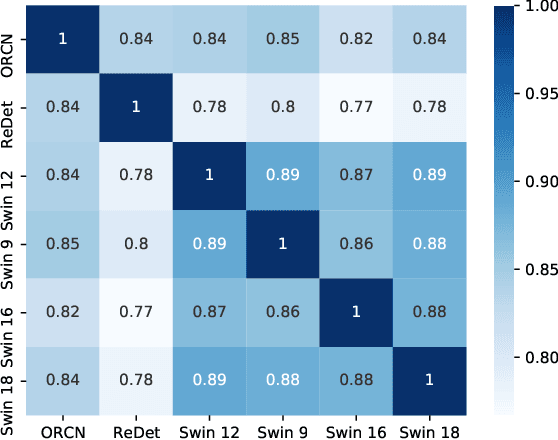
Abstract:Ensemble methods are a reliable way to combine several models to achieve superior performance. However, research on the application of ensemble methods in the remote sensing object detection scenario is mostly overlooked. Two problems arise. First, one unique characteristic of remote sensing object detection is the Oriented Bounding Boxes (OBB) of the objects and the fusion of multiple OBBs requires further research attention. Second, the widely used deep learning object detectors provide a score for each detected object as an indicator of confidence, but how to use these indicators effectively in an ensemble method remains a problem. Trying to address these problems, this paper proposes OBBStacking, an ensemble method that is compatible with OBBs and combines the detection results in a learned fashion. This ensemble method helps take 1st place in the Challenge Track \textit{Fine-grained Object Recognition in High-Resolution Optical Images}, which was featured in \textit{2021 Gaofen Challenge on Automated High-Resolution Earth Observation Image Interpretation}. The experiments on DOTA dataset and FAIR1M dataset demonstrate the improved performance of OBBStacking and the features of OBBStacking are analyzed.
IL-MCAM: An interactive learning and multi-channel attention mechanism-based weakly supervised colorectal histopathology image classification approach
Jun 07, 2022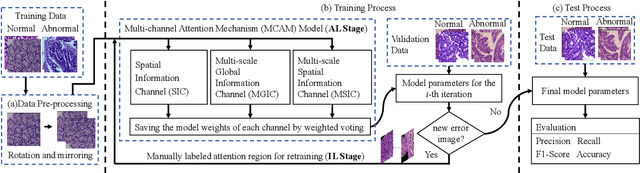
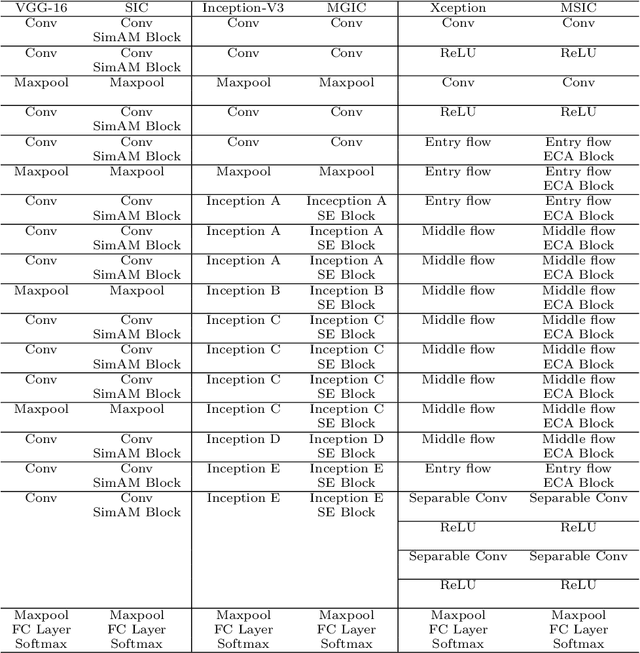
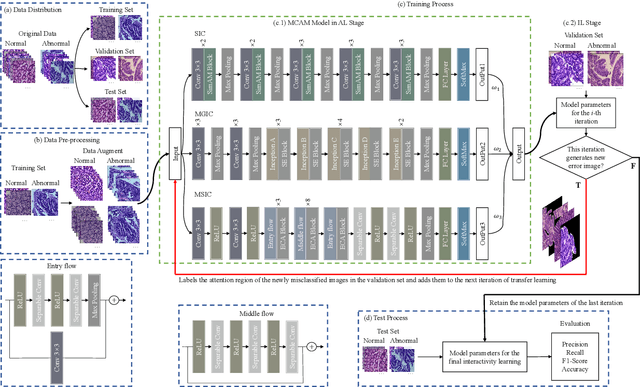
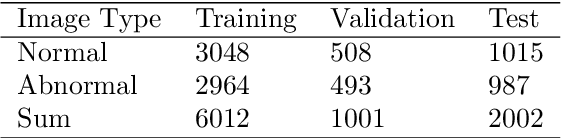
Abstract:In recent years, colorectal cancer has become one of the most significant diseases that endanger human health. Deep learning methods are increasingly important for the classification of colorectal histopathology images. However, existing approaches focus more on end-to-end automatic classification using computers rather than human-computer interaction. In this paper, we propose an IL-MCAM framework. It is based on attention mechanisms and interactive learning. The proposed IL-MCAM framework includes two stages: automatic learning (AL) and interactivity learning (IL). In the AL stage, a multi-channel attention mechanism model containing three different attention mechanism channels and convolutional neural networks is used to extract multi-channel features for classification. In the IL stage, the proposed IL-MCAM framework continuously adds misclassified images to the training set in an interactive approach, which improves the classification ability of the MCAM model. We carried out a comparison experiment on our dataset and an extended experiment on the HE-NCT-CRC-100K dataset to verify the performance of the proposed IL-MCAM framework, achieving classification accuracies of 98.98% and 99.77%, respectively. In addition, we conducted an ablation experiment and an interchangeability experiment to verify the ability and interchangeability of the three channels. The experimental results show that the proposed IL-MCAM framework has excellent performance in the colorectal histopathological image classification tasks.
CVM-Cervix: A Hybrid Cervical Pap-Smear Image Classification Framework Using CNN, Visual Transformer and Multilayer Perceptron
Jun 02, 2022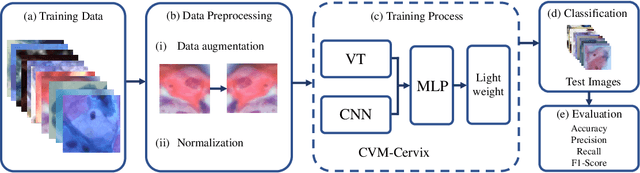
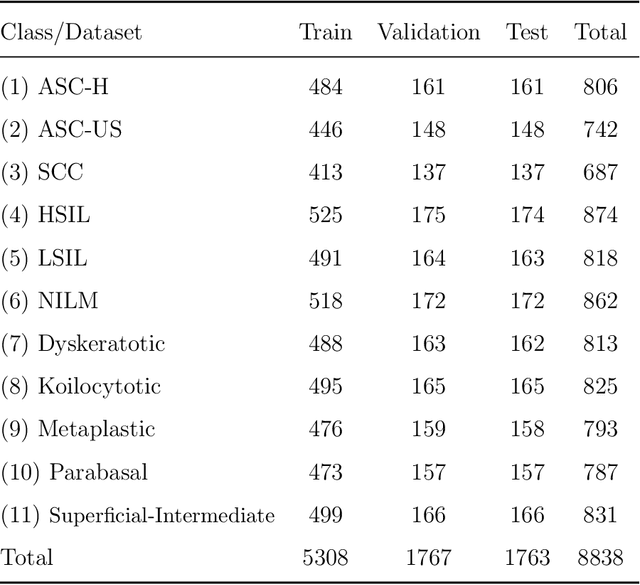
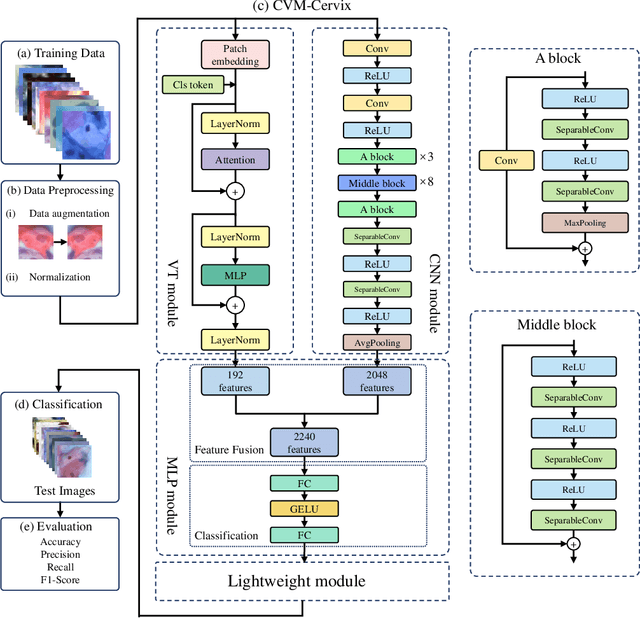
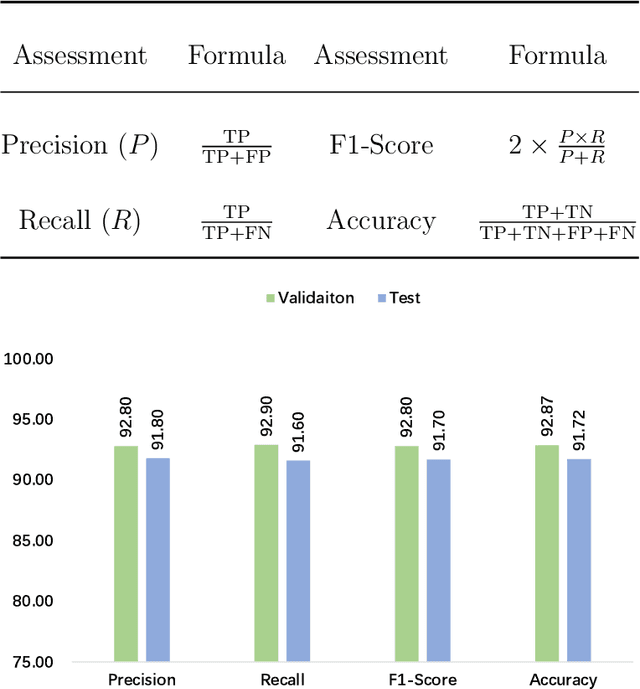
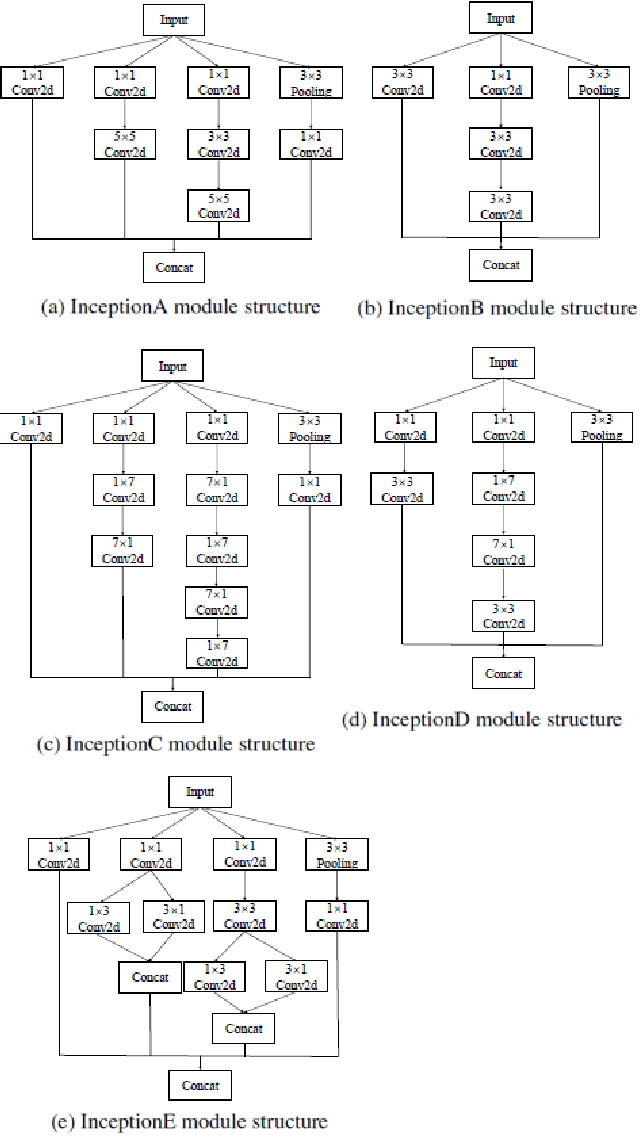
Abstract:Cervical cancer is the seventh most common cancer among all the cancers worldwide and the fourth most common cancer among women. Cervical cytopathology image classification is an important method to diagnose cervical cancer. Manual screening of cytopathology images is time-consuming and error-prone. The emergence of the automatic computer-aided diagnosis system solves this problem. This paper proposes a framework called CVM-Cervix based on deep learning to perform cervical cell classification tasks. It can analyze pap slides quickly and accurately. CVM-Cervix first proposes a Convolutional Neural Network module and a Visual Transformer module for local and global feature extraction respectively, then a Multilayer Perceptron module is designed to fuse the local and global features for the final classification. Experimental results show the effectiveness and potential of the proposed CVM-Cervix in the field of cervical Pap smear image classification. In addition, according to the practical needs of clinical work, we perform a lightweight post-processing to compress the model.
A New Gastric Histopathology Subsize Image Database (GasHisSDB) for Classification Algorithm Test: from Linear Regression to Visual Transformer
Jun 18, 2021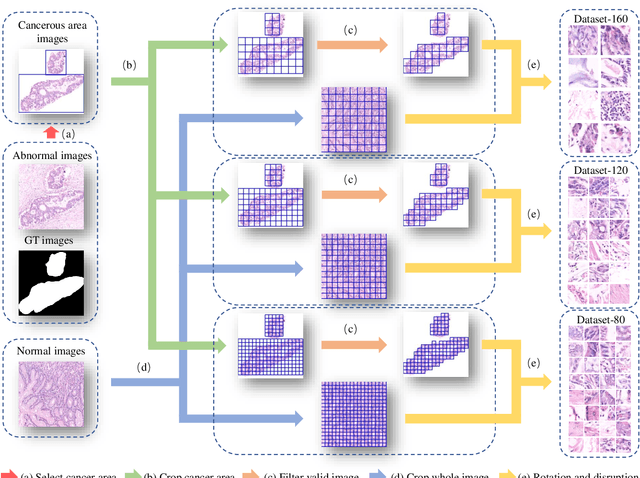
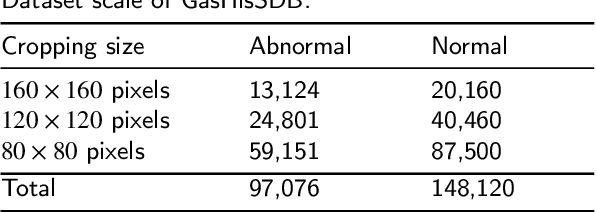
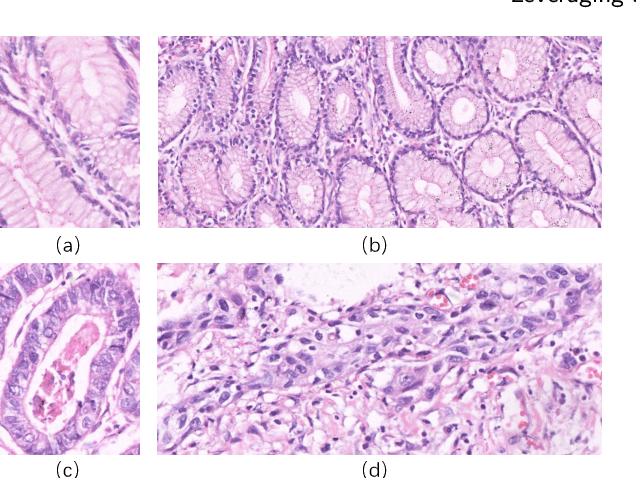
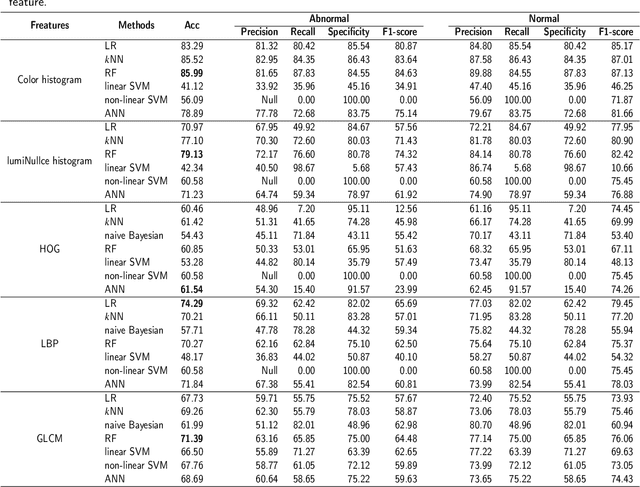
Abstract:GasHisSDB is a New Gastric Histopathology Subsize Image Database with a total of 245196 images. GasHisSDB is divided into 160*160 pixels sub-database, 120*120 pixels sub-database and 80*80 pixels sub-database. GasHisSDB is made to realize the function of valuating image classification. In order to prove that the methods of different periods in the field of image classification have discrepancies on GasHisSDB, we select a variety of classifiers for evaluation. Seven classical machine learning classifiers, three CNN classifiers and a novel transformer-based classifier are selected for testing on image classification tasks. GasHisSDB is available at the URL:https://github.com/NEUhwm/GasHisSDB.git.
GasHis-Transformer: A Multi-scale Visual Transformer Approach for Gastric Histopathology Image Classification
May 25, 2021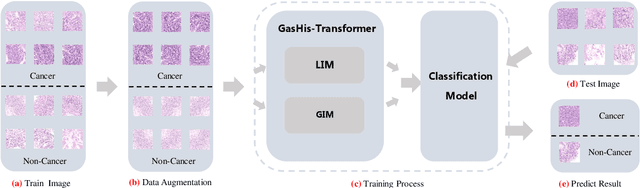
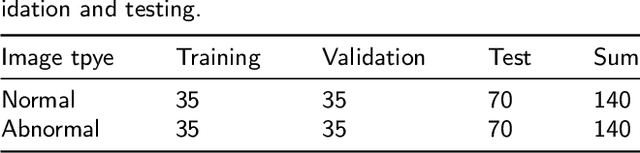

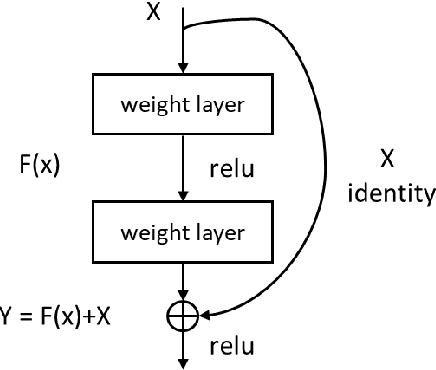
Abstract:Existing deep learning methods for diagnosis of gastric cancer commonly use convolutional neural networks (CNN). Recently, the Visual Transformer (VT) has attracted a major attention because of its performance and efficiency, but its applications are mostly in the field of computer vision. In this paper, a multi-scale visual transformer model, referred to as GasHis-Transformer, is proposed for gastric histopathology image classification (GHIC), which enables the automatic classification of microscopic gastric images into abnormal and normal cases. The GasHis-Transformer model consists of two key modules: a global information module (GIM) and a local information module (LIM) to extract pathological features effectively. In our experiments, a public hematoxylin and eosin (H&E) stained gastric histopathology dataset with 280 abnormal or normal images using the GasHis-Transformer model is applied to estimate precision, recall, F1-score, and accuracy on the testing set as 98.0%, 100.0%, 96.0% and 98.0% respectively. Furthermore, a critical study is conducted to evaluate the robustness of GasHis-Transformer according to add ten different noises including adversarial attack and traditional image noise. In addition, a clinically meaningful study is executed to test the gastric cancer identification of GasHis-Transformerwith 420 abnormal images and achieves 96.2% accuracy. Finally, a comparative study is performed to test the generalizability with both H&E and Immunohistochemical (IHC) stained images on a lymphoma image dataset, a breast cancer dataset and a cervical cancer dataset, producing comparable F1-scores (85.6%, 82.8% and 65.7%, respectively) and accuracy (83.9%, 89.4% and 65.7%, respectively) respectively. In conclusion, GasHis-Transformerdemonstrates a high classification performance and shows its significant potential in histopathology image analysis.
Is Image Size Important? A Robustness Comparison of Deep Learning Methods for Multi-scale Cell Image Classification Tasks: from Convolutional Neural Networks to Visual Transformers
May 16, 2021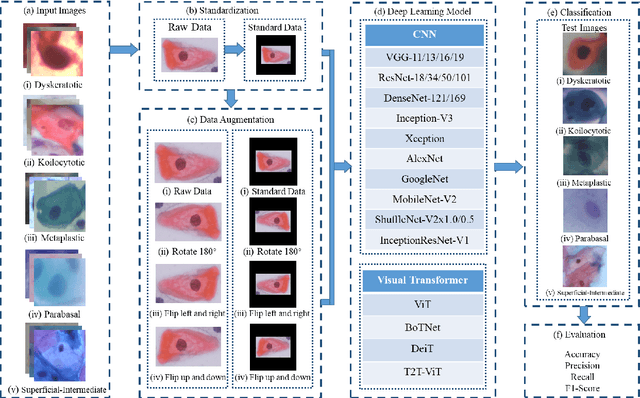

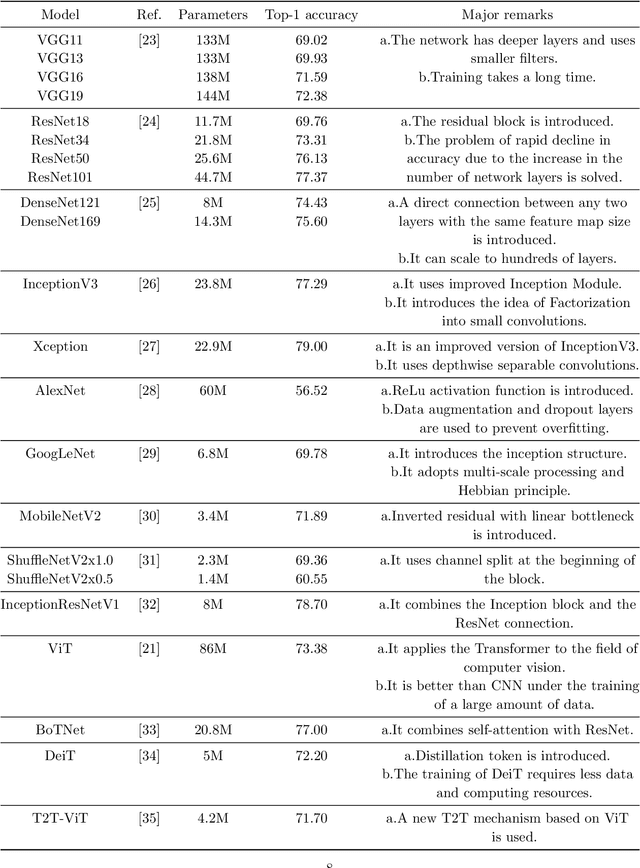
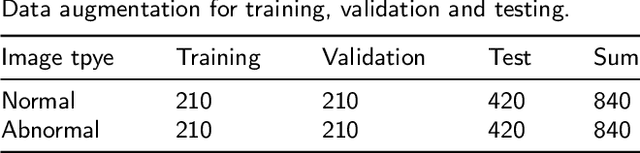
Abstract:Cervical cancer is a very common and fatal cancer in women, but it can be prevented through early examination and treatment. Cytopathology images are often used to screen for cancer. Then, because of the possibility of artificial errors due to the large number of this method, the computer-aided diagnosis system based on deep learning is developed. The image input required by the deep learning method is usually consistent, but the size of the clinical medical image is inconsistent. The internal information is lost after resizing the image directly, so it is unreasonable. A lot of research is to directly resize the image, and the results are still robust. In order to find a reasonable explanation, 22 deep learning models are used to process images of different scales, and experiments are conducted on the SIPaKMeD dataset. The conclusion is that the deep learning method is very robust to the size changes of images. This conclusion is also validated on the Herlev dataset.
A Hierarchical Conditional Random Field-based Attention Mechanism Approach for Gastric Histopathology Image Classification
Feb 21, 2021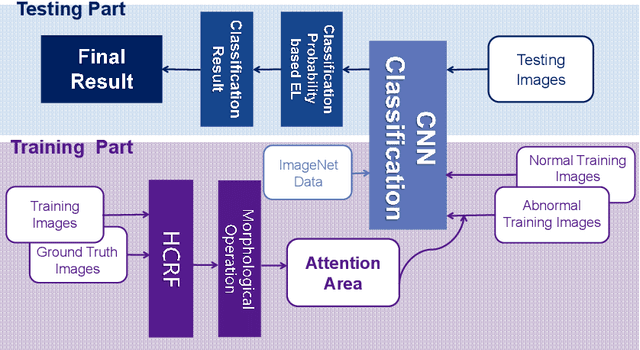

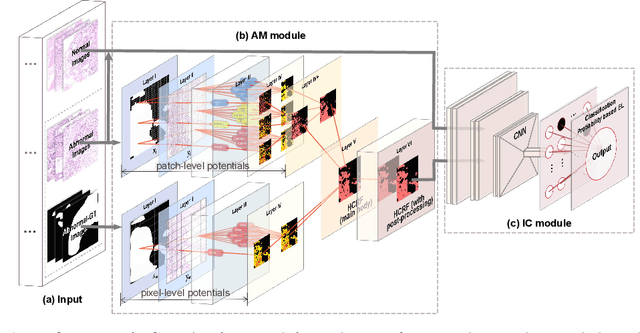
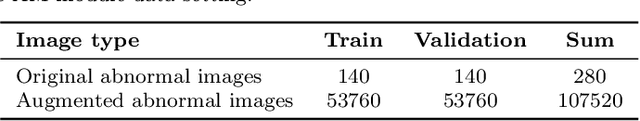
Abstract:In the Gastric Histopathology Image Classification (GHIC) tasks, which is usually weakly supervised learning missions, there is inevitably redundant information in the images. Therefore, designing networks that can focus on effective distinguishing features has become a popular research topic. In this paper, to accomplish the tasks of GHIC superiorly and to assist pathologists in clinical diagnosis, an intelligent Hierarchical Conditional Random Field based Attention Mechanism (HCRF-AM) model is proposed. The HCRF-AM model consists of an Attention Mechanism (AM) module and an Image Classification (IC) module. In the AM module, an HCRF model is built to extract attention regions. In the IC module, a Convolutional Neural Network (CNN) model is trained with the attention regions selected and then an algorithm called Classification Probability-based Ensemble Learning is applied to obtain the image-level results from patch-level output of the CNN. In the experiment, a classification specificity of 96.67% is achieved on a gastric histopathology dataset with 700 images. Our HCRF-AM model demonstrates high classification performance and shows its effectiveness and future potential in the GHIC field.
Study on MCS Selection and Spectrum Allocation for URLLC Traffic under Delay and Reliability Constraint in 5G Network
Jan 06, 2021
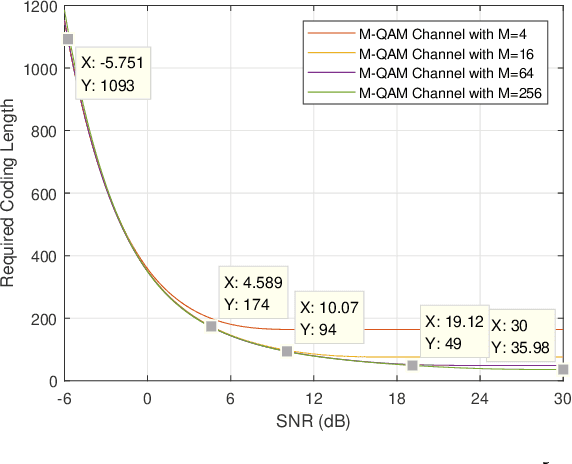
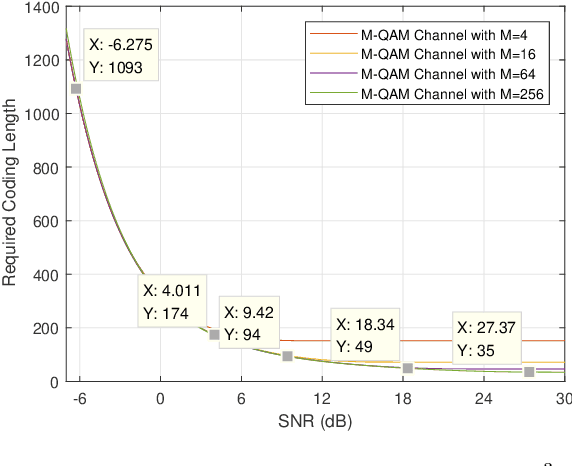
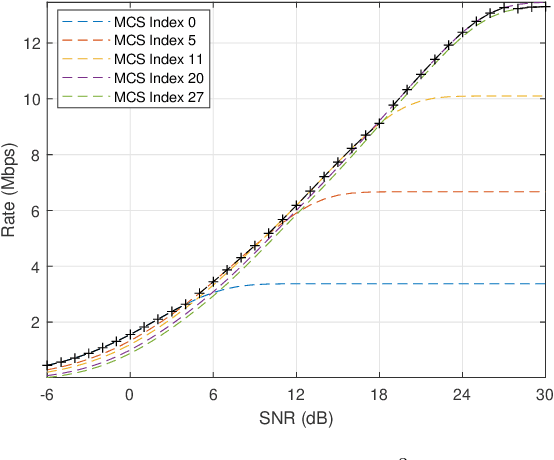
Abstract:To support Ultra-Reliable and Low Latency Communications (URLLC) is an essential character of the 5th Generation (5G) communication system. Unlike the other two use cases defined in 5G, e.g. enhanced Mobile Broadband (eMBB) and massive Machine Type Communications (mMTC), URLLC traffic has strict delay and reliability requirement. In this paper, an analysis model for URLLC traffic is proposed from the generation of a URLLC traffic until its transmission over a wireless channel, where channel quality, coding scheme with finite coding length, modulation scheme and allocated spectrum resource are taken into consideration. Then, network calculus analysis is applied to derive the delay guarantee for periodical URLLC traffic. Based on the delay analysis, the admission region is found under certain delay and reliability requirement, which gives a lower bound on required spectrum resource. Theoretical results in the scenario of a 5G New Radio system are presented, where the SNR thresholds for adaptive modulation and coding scheme selection, transmission rate and delay, as well as admission region under different configurations are discussed. In addition, simulation results are obtained and compared with theoretical results, which validates that the admission region derived in this work provides a lower spectrum allocation bound.
A Comprehensive Review for MRF and CRF Approaches in Pathology Image Analysis
Sep 29, 2020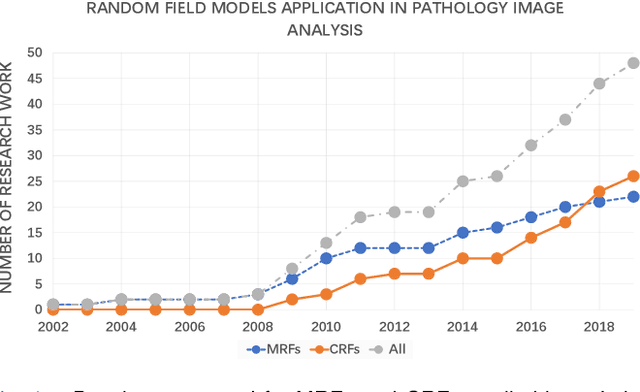
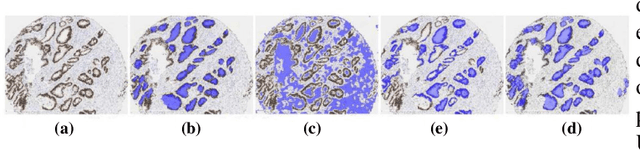
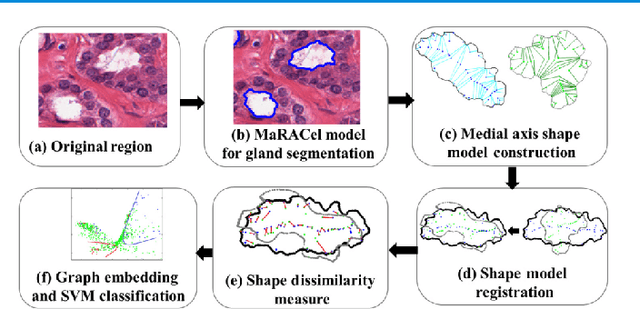
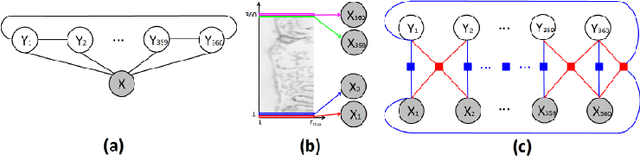

Abstract:Pathology image analysis is an essential procedure for clinical diagnosis of many diseases. To boost the accuracy and objectivity of detection, nowadays, an increasing number of computer-aided diagnosis (CAD) system is proposed. Among these methods, random field models play an indispensable role in improving the analysis performance. In this review, we present a comprehensive overview of pathology image analysis based on the markov random fields (MRFs) and conditional random fields (CRFs), which are two popular random field models. Firstly, we introduce the background of two random fields and pathology images. Secondly, we summarize the basic mathematical knowledge of MRFs and CRFs from modelling to optimization. Then, a thorough review of the recent research on the MRFs and CRFs of pathology images analysis is presented. Finally, we investigate the popular methodologies in the related works and discuss the method migration among CAD field.
A Comprehensive Review for Breast Histopathology Image Analysis Using Classical and Deep Neural Networks
Mar 27, 2020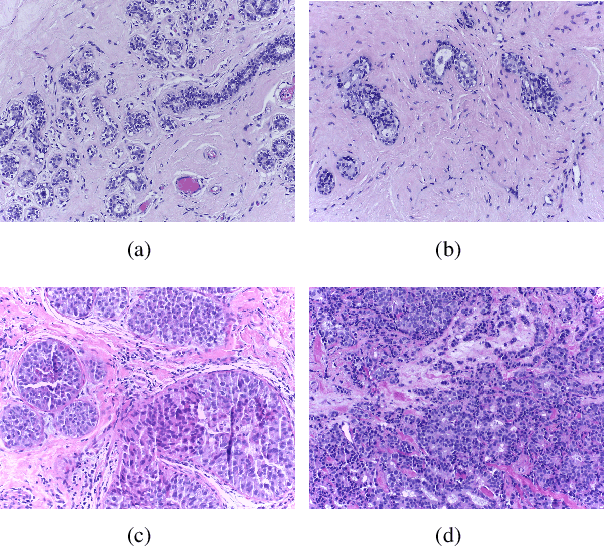
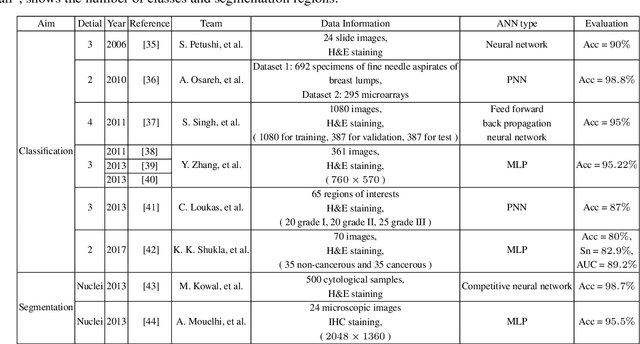
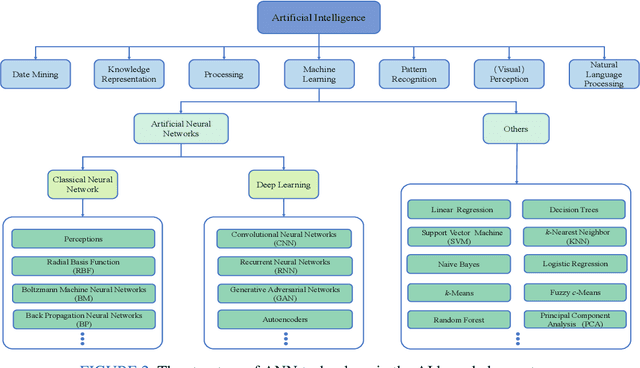
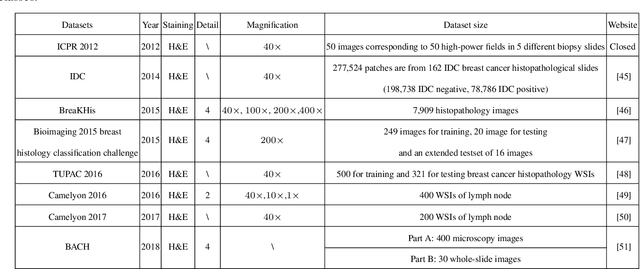
Abstract:Breast cancer is one of the most common and deadliest cancers among women. Since histopathological images contain sufficient phenotypic information, they play an indispensable role in the diagnosis and treatment of breast cancers. To improve the accuracy and objectivity of Breast Histopathological Image Analysis (BHIA), Artificial Neural Network (ANN) approaches are widely used in the segmentation and classification tasks of breast histopathological images. In this review, we present a comprehensive overview of the BHIA techniques based on ANNs. First of all, we categorize the BHIA systems into classical and deep neural networks for in-depth investigation. Then, the relevant studies based on BHIA systems are presented. After that, we analyze the existing models to discover the most suitable algorithms. Finally, publicly accessible datasets, along with their download links, are provided for the convenience of future researchers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge