Carola-Bibiane Schönlieb
on behalf of the AIX-COVNET collaboration
DiTASK: Multi-Task Fine-Tuning with Diffeomorphic Transformations
Feb 09, 2025Abstract:Pre-trained Vision Transformers now serve as powerful tools for computer vision. Yet, efficiently adapting them for multiple tasks remains a challenge that arises from the need to modify the rich hidden representations encoded by the learned weight matrices, without inducing interference between tasks. Current parameter-efficient methods like LoRA, which apply low-rank updates, force tasks to compete within constrained subspaces, ultimately degrading performance. We introduce DiTASK a novel Diffeomorphic Multi-Task Fine-Tuning approach that maintains pre-trained representations by preserving weight matrix singular vectors, while enabling task-specific adaptations through neural diffeomorphic transformations of the singular values. By following this approach, DiTASK enables both shared and task-specific feature modulations with minimal added parameters. Our theoretical analysis shows that DITASK achieves full-rank updates during optimization, preserving the geometric structure of pre-trained features, and establishing a new paradigm for efficient multi-task learning (MTL). Our experiments on PASCAL MTL and NYUD show that DiTASK achieves state-of-the-art performance across four dense prediction tasks, using 75% fewer parameters than existing methods.
Generative Unordered Flow for Set-Structured Data Generation
Jan 29, 2025Abstract:Flow-based generative models have demonstrated promising performance across a broad spectrum of data modalities (e.g., image and text). However, there are few works exploring their extension to unordered data (e.g., spatial point set), which is not trivial because previous models are mostly designed for vector data that are naturally ordered. In this paper, we present unordered flow, a type of flow-based generative model for set-structured data generation. Specifically, we convert unordered data into an appropriate function representation, and learn the probability measure of such representations through function-valued flow matching. For the inverse map from a function representation to unordered data, we propose a method similar to particle filtering, with Langevin dynamics to first warm-up the initial particles and gradient-based search to update them until convergence. We have conducted extensive experiments on multiple real-world datasets, showing that our unordered flow model is very effective in generating set-structured data and significantly outperforms previous baselines.
Inverse Evolution Data Augmentation for Neural PDE Solvers
Jan 24, 2025Abstract:Neural networks have emerged as promising tools for solving partial differential equations (PDEs), particularly through the application of neural operators. Training neural operators typically requires a large amount of training data to ensure accuracy and generalization. In this paper, we propose a novel data augmentation method specifically designed for training neural operators on evolution equations. Our approach utilizes insights from inverse processes of these equations to efficiently generate data from random initialization that are combined with original data. To further enhance the accuracy of the augmented data, we introduce high-order inverse evolution schemes. These schemes consist of only a few explicit computation steps, yet the resulting data pairs can be proven to satisfy the corresponding implicit numerical schemes. In contrast to traditional PDE solvers that require small time steps or implicit schemes to guarantee accuracy, our data augmentation method employs explicit schemes with relatively large time steps, thereby significantly reducing computational costs. Accuracy and efficacy experiments confirm the effectiveness of our approach. Additionally, we validate our approach through experiments with the Fourier Neural Operator and UNet on three common evolution equations that are Burgers' equation, the Allen-Cahn equation and the Navier-Stokes equation. The results demonstrate a significant improvement in the performance and robustness of the Fourier Neural Operator when coupled with our inverse evolution data augmentation method.
Towards Foundation Models on Graphs: An Analysis on Cross-Dataset Transfer of Pretrained GNNs
Dec 23, 2024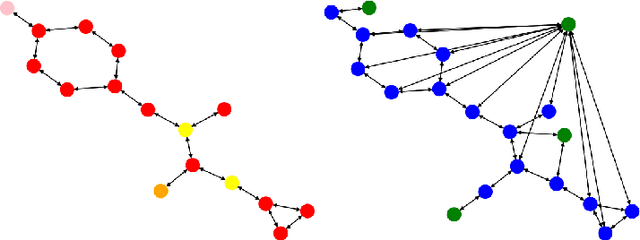
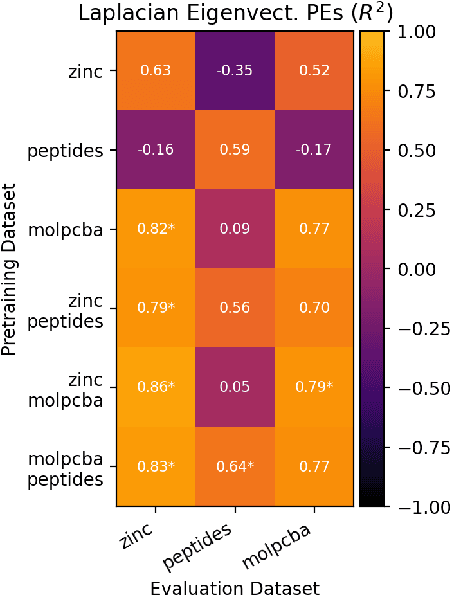
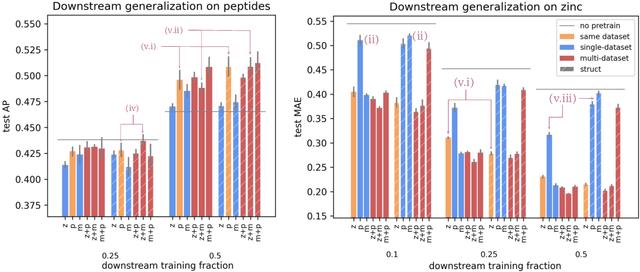
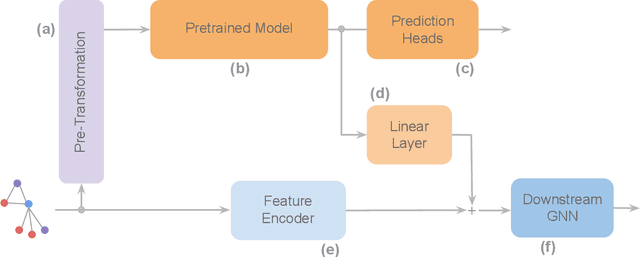
Abstract:To develop a preliminary understanding towards Graph Foundation Models, we study the extent to which pretrained Graph Neural Networks can be applied across datasets, an effort requiring to be agnostic to dataset-specific features and their encodings. We build upon a purely structural pretraining approach and propose an extension to capture feature information while still being feature-agnostic. We evaluate pretrained models on downstream tasks for varying amounts of training samples and choices of pretraining datasets. Our preliminary results indicate that embeddings from pretrained models improve generalization only with enough downstream data points and in a degree which depends on the quantity and properties of pretraining data. Feature information can lead to improvements, but currently requires some similarities between pretraining and downstream feature spaces.
Symplectic Neural Flows for Modeling and Discovery
Dec 21, 2024Abstract:Hamilton's equations are fundamental for modeling complex physical systems, where preserving key properties such as energy and momentum is crucial for reliable long-term simulations. Geometric integrators are widely used for this purpose, but neural network-based methods that incorporate these principles remain underexplored. This work introduces SympFlow, a time-dependent symplectic neural network designed using parameterized Hamiltonian flow maps. This design allows for backward error analysis and ensures the preservation of the symplectic structure. SympFlow allows for two key applications: (i) providing a time-continuous symplectic approximation of the exact flow of a Hamiltonian system--purely based on the differential equations it satisfies, and (ii) approximating the flow map of an unknown Hamiltonian system relying on trajectory data. We demonstrate the effectiveness of SympFlow on diverse problems, including chaotic and dissipative systems, showing improved energy conservation compared to general-purpose numerical methods and accurate
Cross-Modal Few-Shot Learning with Second-Order Neural Ordinary Differential Equations
Dec 20, 2024



Abstract:We introduce SONO, a novel method leveraging Second-Order Neural Ordinary Differential Equations (Second-Order NODEs) to enhance cross-modal few-shot learning. By employing a simple yet effective architecture consisting of a Second-Order NODEs model paired with a cross-modal classifier, SONO addresses the significant challenge of overfitting, which is common in few-shot scenarios due to limited training examples. Our second-order approach can approximate a broader class of functions, enhancing the model's expressive power and feature generalization capabilities. We initialize our cross-modal classifier with text embeddings derived from class-relevant prompts, streamlining training efficiency by avoiding the need for frequent text encoder processing. Additionally, we utilize text-based image augmentation, exploiting CLIP's robust image-text correlation to enrich training data significantly. Extensive experiments across multiple datasets demonstrate that SONO outperforms existing state-of-the-art methods in few-shot learning performance.
Training Data Reconstruction: Privacy due to Uncertainty?
Dec 11, 2024



Abstract:Being able to reconstruct training data from the parameters of a neural network is a major privacy concern. Previous works have shown that reconstructing training data, under certain circumstances, is possible. In this work, we analyse such reconstructions empirically and propose a new formulation of the reconstruction as a solution to a bilevel optimisation problem. We demonstrate that our formulation as well as previous approaches highly depend on the initialisation of the training images $x$ to reconstruct. In particular, we show that a random initialisation of $x$ can lead to reconstructions that resemble valid training samples while not being part of the actual training dataset. Thus, our experiments on affine and one-hidden layer networks suggest that when reconstructing natural images, yet an adversary cannot identify whether reconstructed images have indeed been part of the set of training samples.
Benchmarking learned algorithms for computed tomography image reconstruction tasks
Dec 11, 2024



Abstract:Computed tomography (CT) is a widely used non-invasive diagnostic method in various fields, and recent advances in deep learning have led to significant progress in CT image reconstruction. However, the lack of large-scale, open-access datasets has hindered the comparison of different types of learned methods. To address this gap, we use the 2DeteCT dataset, a real-world experimental computed tomography dataset, for benchmarking machine learning based CT image reconstruction algorithms. We categorize these methods into post-processing networks, learned/unrolled iterative methods, learned regularizer methods, and plug-and-play methods, and provide a pipeline for easy implementation and evaluation. Using key performance metrics, including SSIM and PSNR, our benchmarking results showcase the effectiveness of various algorithms on tasks such as full data reconstruction, limited-angle reconstruction, sparse-angle reconstruction, low-dose reconstruction, and beam-hardening corrected reconstruction. With this benchmarking study, we provide an evaluation of a range of algorithms representative for different categories of learned reconstruction methods on a recently published dataset of real-world experimental CT measurements. The reproducible setup of methods and CT image reconstruction tasks in an open-source toolbox enables straightforward addition and comparison of new methods later on. The toolbox also provides the option to load the 2DeteCT dataset differently for extensions to other problems and different CT reconstruction tasks.
Towards Graph Foundation Models: A Study on the Generalization of Positional and Structural Encodings
Dec 10, 2024Abstract:Recent advances in integrating positional and structural encodings (PSEs) into graph neural networks (GNNs) have significantly enhanced their performance across various graph learning tasks. However, the general applicability of these encodings and their potential to serve as foundational representations for graphs remain uncertain. This paper investigates the fine-tuning efficiency, scalability with sample size, and generalization capability of learnable PSEs across diverse graph datasets. Specifically, we evaluate their potential as universal pre-trained models that can be easily adapted to new tasks with minimal fine-tuning and limited data. Furthermore, we assess the expressivity of the learned representations, particularly, when used to augment downstream GNNs. We demonstrate through extensive benchmarking and empirical analysis that PSEs generally enhance downstream models. However, some datasets may require specific PSE-augmentations to achieve optimal performance. Nevertheless, our findings highlight their significant potential to become integral components of future graph foundation models. We provide new insights into the strengths and limitations of PSEs, contributing to the broader discourse on foundation models in graph learning.
You KAN Do It in a Single Shot: Plug-and-Play Methods with Single-Instance Priors
Dec 09, 2024



Abstract:The use of Plug-and-Play (PnP) methods has become a central approach for solving inverse problems, with denoisers serving as regularising priors that guide optimisation towards a clean solution. In this work, we introduce KAN-PnP, an optimisation framework that incorporates Kolmogorov-Arnold Networks (KANs) as denoisers within the Plug-and-Play (PnP) paradigm. KAN-PnP is specifically designed to solve inverse problems with single-instance priors, where only a single noisy observation is available, eliminating the need for large datasets typically required by traditional denoising methods. We show that KANs, based on the Kolmogorov-Arnold representation theorem, serve effectively as priors in such settings, providing a robust approach to denoising. We prove that the KAN denoiser is Lipschitz continuous, ensuring stability and convergence in optimisation algorithms like PnP-ADMM, even in the context of single-shot learning. Additionally, we provide theoretical guarantees for KAN-PnP, demonstrating its convergence under key conditions: the convexity of the data fidelity term, Lipschitz continuity of the denoiser, and boundedness of the regularisation functional. These conditions are crucial for stable and reliable optimisation. Our experimental results show, on super-resolution and joint optimisation, that KAN-PnP outperforms exiting methods, delivering superior performance in single-shot learning with minimal data. The method exhibits strong convergence properties, achieving high accuracy with fewer iterations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge