Byron C. Wallace
Evaluating Factuality in Text Simplification
Apr 15, 2022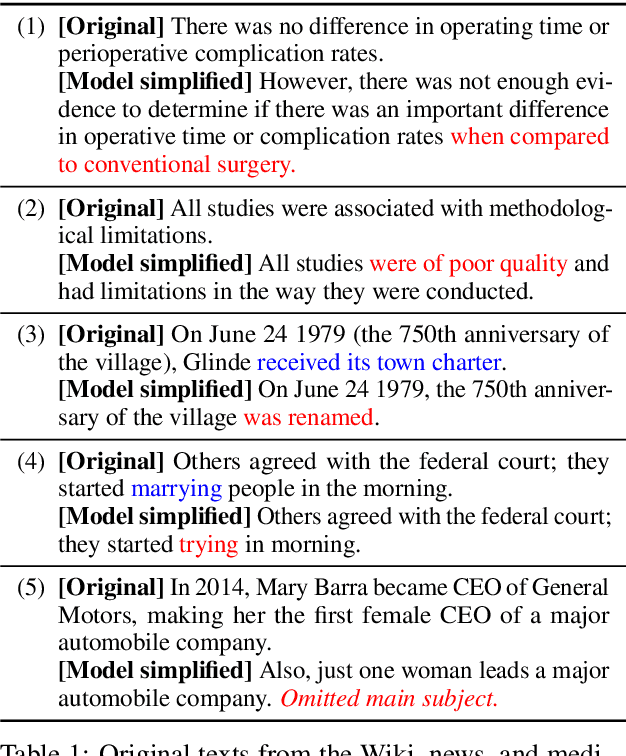


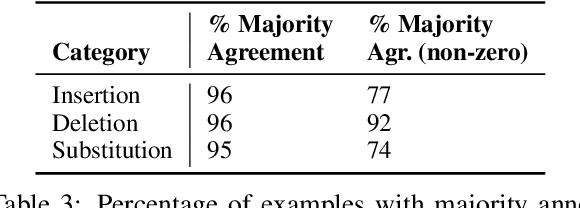
Abstract:Automated simplification models aim to make input texts more readable. Such methods have the potential to make complex information accessible to a wider audience, e.g., providing access to recent medical literature which might otherwise be impenetrable for a lay reader. However, such models risk introducing errors into automatically simplified texts, for instance by inserting statements unsupported by the corresponding original text, or by omitting key information. Providing more readable but inaccurate versions of texts may in many cases be worse than providing no such access at all. The problem of factual accuracy (and the lack thereof) has received heightened attention in the context of summarization models, but the factuality of automatically simplified texts has not been investigated. We introduce a taxonomy of errors that we use to analyze both references drawn from standard simplification datasets and state-of-the-art model outputs. We find that errors often appear in both that are not captured by existing evaluation metrics, motivating a need for research into ensuring the factual accuracy of automated simplification models.
What Would it Take to get Biomedical QA Systems into Practice?
Sep 21, 2021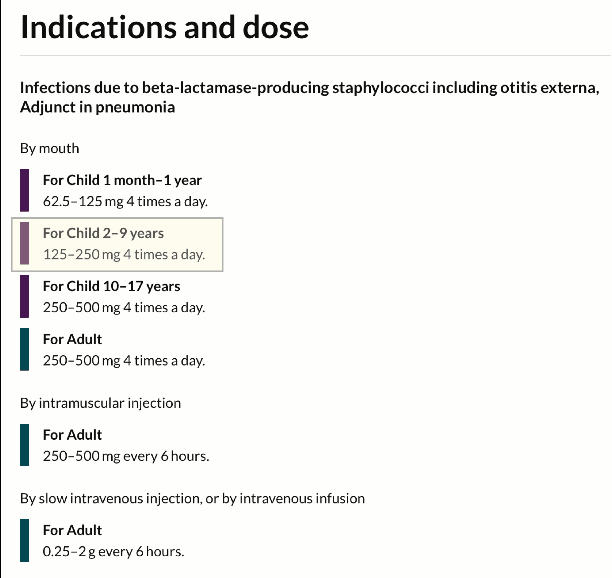
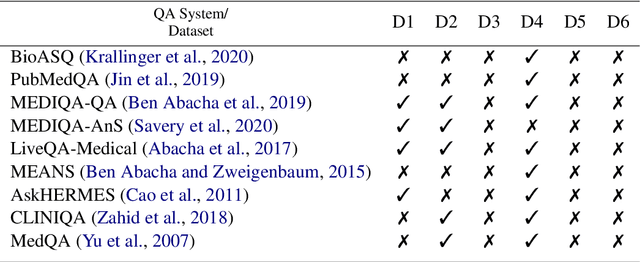


Abstract:Medical question answering (QA) systems have the potential to answer clinicians uncertainties about treatment and diagnosis on demand, informed by the latest evidence. However, despite the significant progress in general QA made by the NLP community, medical QA systems are still not widely used in clinical environments. One likely reason for this is that clinicians may not readily trust QA system outputs, in part because transparency, trustworthiness, and provenance have not been key considerations in the design of such models. In this paper we discuss a set of criteria that, if met, we argue would likely increase the utility of biomedical QA systems, which may in turn lead to adoption of such systems in practice. We assess existing models, tasks, and datasets with respect to these criteria, highlighting shortcomings of previously proposed approaches and pointing toward what might be more usable QA systems.
Combining Feature and Instance Attribution to Detect Artifacts
Jul 01, 2021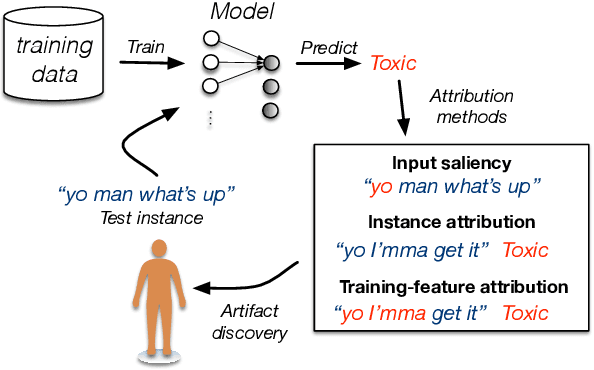
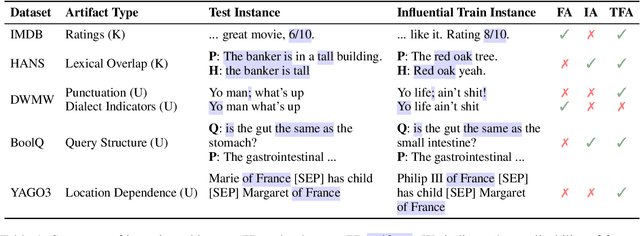

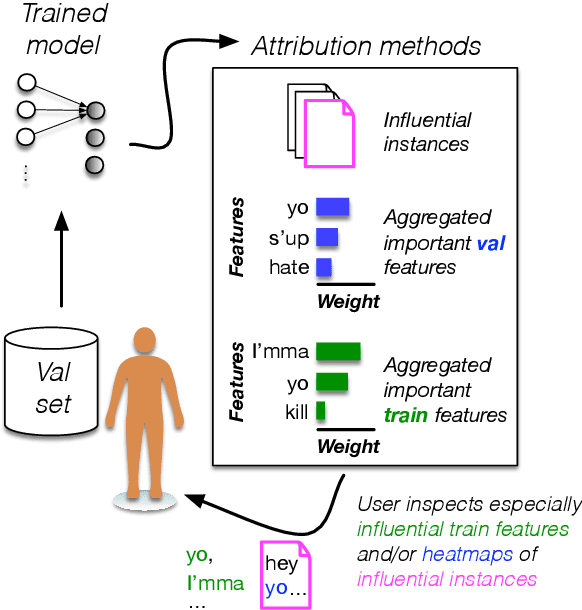
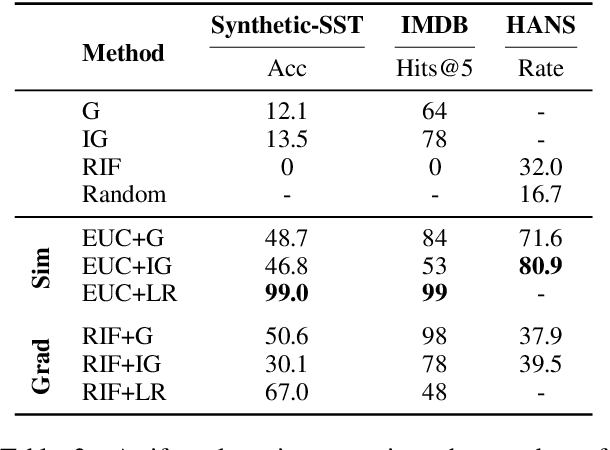
Abstract:Training the large deep neural networks that dominate NLP requires large datasets. Many of these are collected automatically or via crowdsourcing, and may exhibit systematic biases or annotation artifacts. By the latter, we mean correlations between inputs and outputs that are spurious, insofar as they do not represent a generally held causal relationship between features and classes; models that exploit such correlations may appear to perform a given task well, but fail on out of sample data. In this paper we propose methods to facilitate identification of training data artifacts, using new hybrid approaches that combine saliency maps (which highlight important input features) with instance attribution methods (which retrieve training samples influential to a given prediction). We show that this proposed training-feature attribution approach can be used to uncover artifacts in training data, and use it to identify previously unreported artifacts in a few standard NLP datasets. We execute a small user study to evaluate whether these methods are useful to NLP researchers in practice, with promising results. We make code for all methods and experiments in this paper available.
Biomedical Interpretable Entity Representations
Jun 17, 2021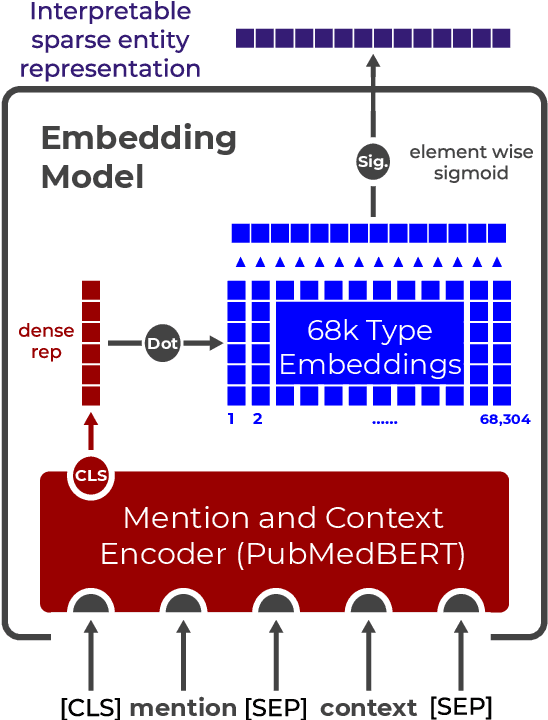
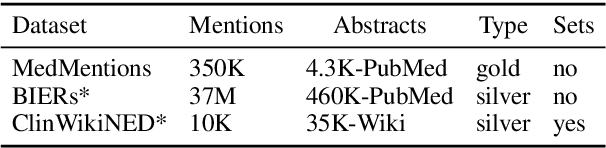
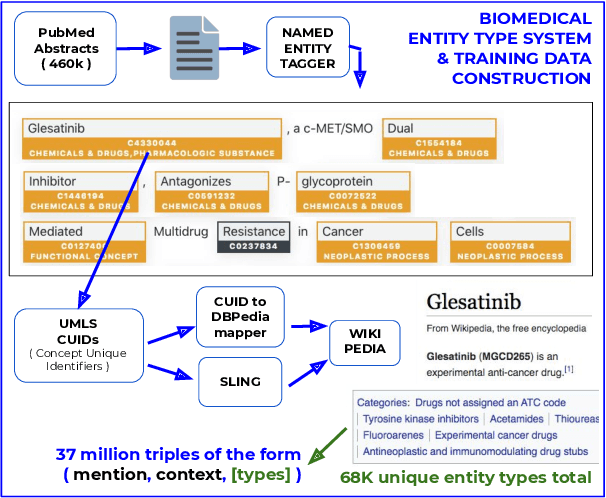
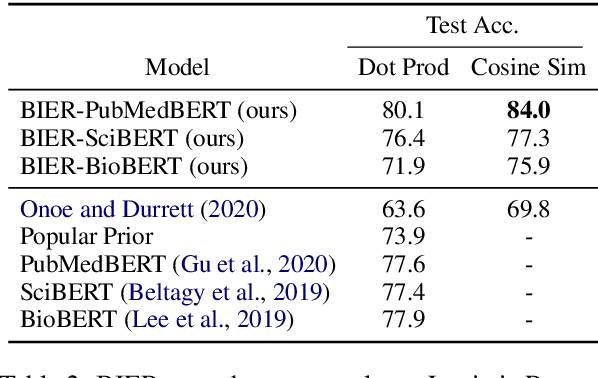
Abstract:Pre-trained language models induce dense entity representations that offer strong performance on entity-centric NLP tasks, but such representations are not immediately interpretable. This can be a barrier to model uptake in important domains such as biomedicine. There has been recent work on general interpretable representation learning (Onoe and Durrett, 2020), but these domain-agnostic representations do not readily transfer to the important domain of biomedicine. In this paper, we create a new entity type system and training set from a large corpus of biomedical texts by mapping entities to concepts in a medical ontology, and from these to Wikipedia pages whose categories are our types. From this mapping we derive Biomedical Interpretable Entity Representations(BIERs), in which dimensions correspond to fine-grained entity types, and values are predicted probabilities that a given entity is of the corresponding type. We propose a novel method that exploits BIER's final sparse and intermediate dense representations to facilitate model and entity type debugging. We show that BIERs achieve strong performance in biomedical tasks including named entity disambiguation and entity label classification, and we provide error analysis to highlight the utility of their interpretability, particularly in low-supervision settings. Finally, we provide our induced 68K biomedical type system, the corresponding 37 million triples of derived data used to train BIER models and our best performing model.
Does BERT Pretrained on Clinical Notes Reveal Sensitive Data?
Apr 22, 2021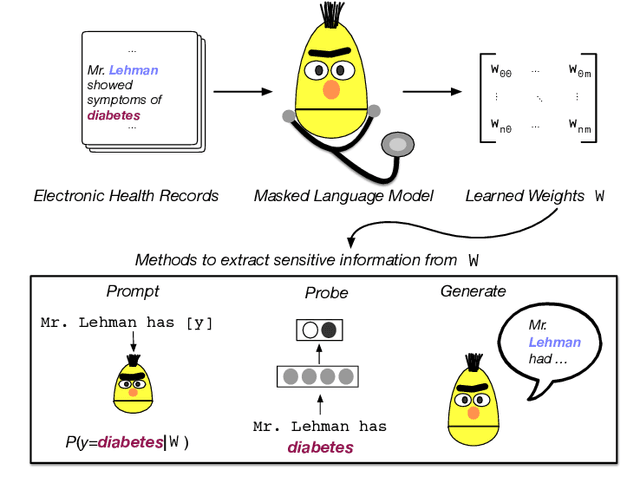

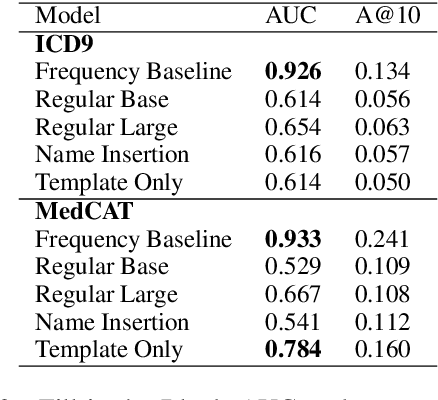
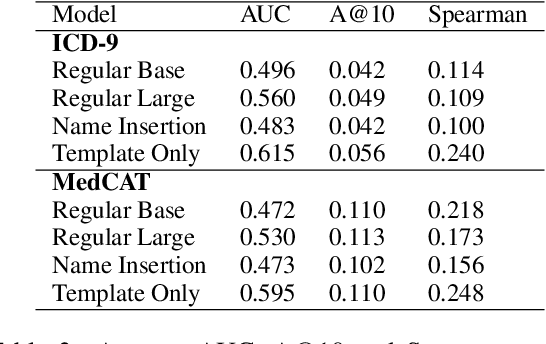
Abstract:Large Transformers pretrained over clinical notes from Electronic Health Records (EHR) have afforded substantial gains in performance on predictive clinical tasks. The cost of training such models (and the necessity of data access to do so) coupled with their utility motivates parameter sharing, i.e., the release of pretrained models such as ClinicalBERT. While most efforts have used deidentified EHR, many researchers have access to large sets of sensitive, non-deidentified EHR with which they might train a BERT model (or similar). Would it be safe to release the weights of such a model if they did? In this work, we design a battery of approaches intended to recover Personal Health Information (PHI) from a trained BERT. Specifically, we attempt to recover patient names and conditions with which they are associated. We find that simple probing methods are not able to meaningfully extract sensitive information from BERT trained over the MIMIC-III corpus of EHR. However, more sophisticated "attacks" may succeed in doing so: To facilitate such research, we make our experimental setup and baseline probing models available at https://github.com/elehman16/exposing_patient_data_release
Disentangling Representations of Text by Masking Transformers
Apr 14, 2021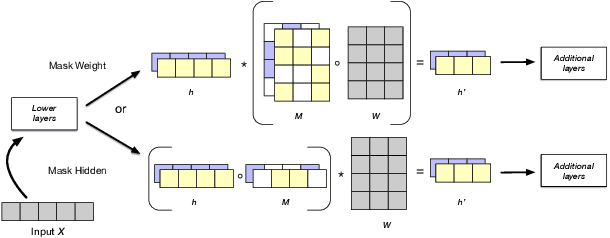
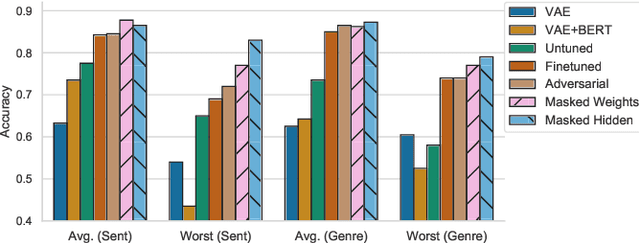
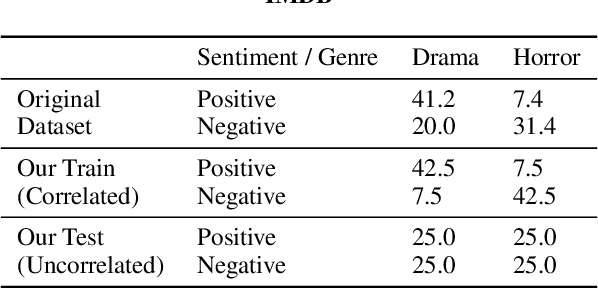
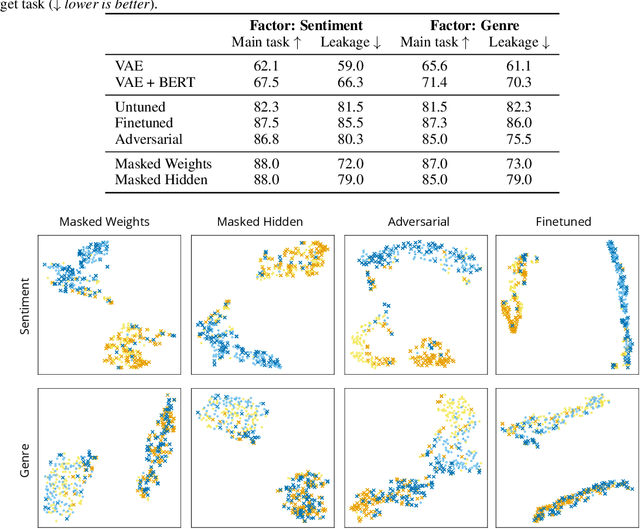
Abstract:Representations from large pretrained models such as BERT encode a range of features into monolithic vectors, affording strong predictive accuracy across a multitude of downstream tasks. In this paper we explore whether it is possible to learn disentangled representations by identifying existing subnetworks within pretrained models that encode distinct, complementary aspect representations. Concretely, we learn binary masks over transformer weights or hidden units to uncover subsets of features that correlate with a specific factor of variation; this eliminates the need to train a disentangled model from scratch for a particular task. We evaluate this method with respect to its ability to disentangle representations of sentiment from genre in movie reviews, "toxicity" from dialect in Tweets, and syntax from semantics. By combining masking with magnitude pruning we find that we can identify sparse subnetworks within BERT that strongly encode particular aspects (e.g., toxicity) while only weakly encoding others (e.g., race). Moreover, despite only learning masks, we find that disentanglement-via-masking performs as well as -- and often better than -- previously proposed methods based on variational autoencoders and adversarial training.
On the Impact of Random Seeds on the Fairness of Clinical Classifiers
Apr 13, 2021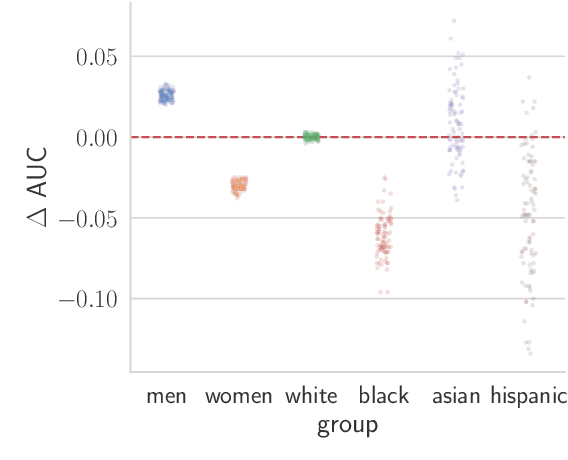
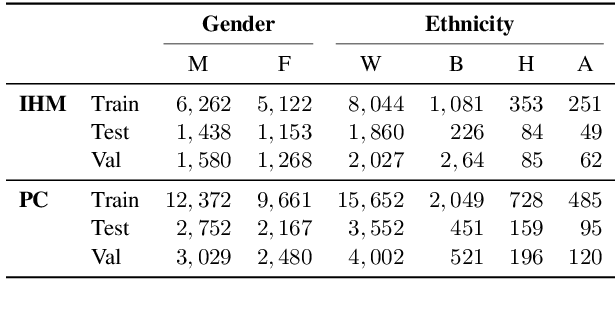
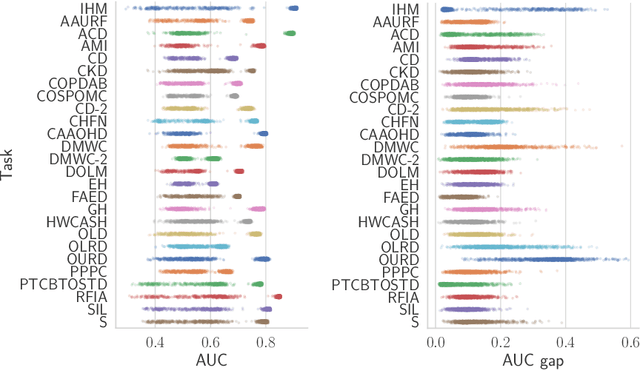
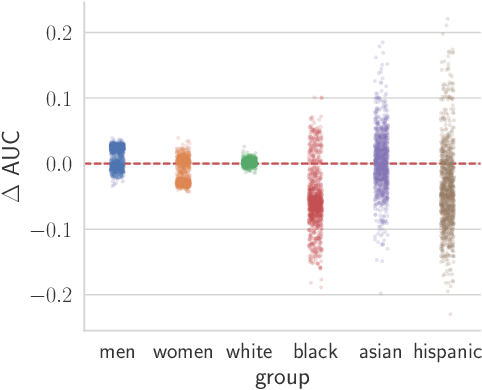
Abstract:Recent work has shown that fine-tuning large networks is surprisingly sensitive to changes in random seed(s). We explore the implications of this phenomenon for model fairness across demographic groups in clinical prediction tasks over electronic health records (EHR) in MIMIC-III -- the standard dataset in clinical NLP research. Apparent subgroup performance varies substantially for seeds that yield similar overall performance, although there is no evidence of a trade-off between overall and subgroup performance. However, we also find that the small sample sizes inherent to looking at intersections of minority groups and somewhat rare conditions limit our ability to accurately estimate disparities. Further, we find that jointly optimizing for high overall performance and low disparities does not yield statistically significant improvements. Our results suggest that fairness work using MIMIC-III should carefully account for variations in apparent differences that may arise from stochasticity and small sample sizes.
Paragraph-level Simplification of Medical Texts
Apr 12, 2021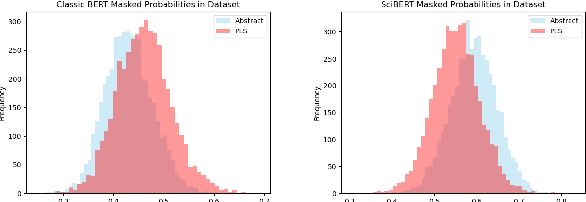
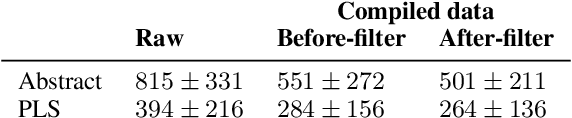
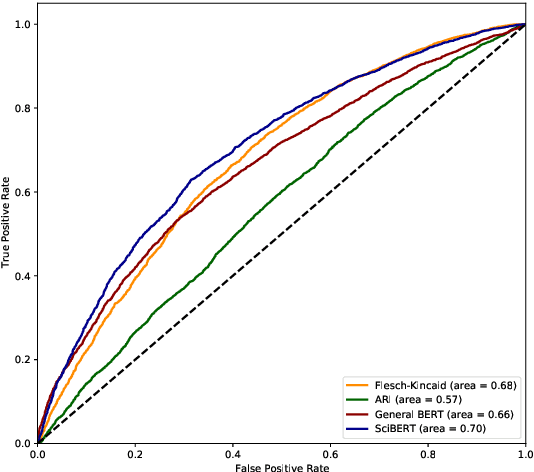

Abstract:We consider the problem of learning to simplify medical texts. This is important because most reliable, up-to-date information in biomedicine is dense with jargon and thus practically inaccessible to the lay audience. Furthermore, manual simplification does not scale to the rapidly growing body of biomedical literature, motivating the need for automated approaches. Unfortunately, there are no large-scale resources available for this task. In this work we introduce a new corpus of parallel texts in English comprising technical and lay summaries of all published evidence pertaining to different clinical topics. We then propose a new metric based on likelihood scores from a masked language model pretrained on scientific texts. We show that this automated measure better differentiates between technical and lay summaries than existing heuristics. We introduce and evaluate baseline encoder-decoder Transformer models for simplification and propose a novel augmentation to these in which we explicitly penalize the decoder for producing "jargon" terms; we find that this yields improvements over baselines in terms of readability.
An Empirical Comparison of Instance Attribution Methods for NLP
Apr 09, 2021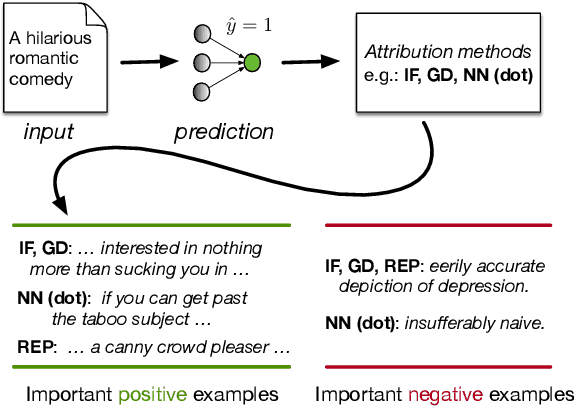
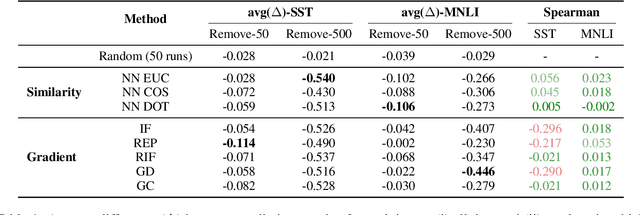
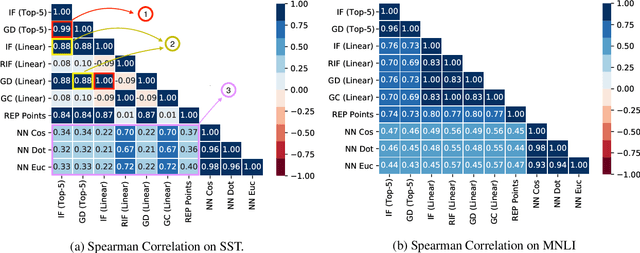
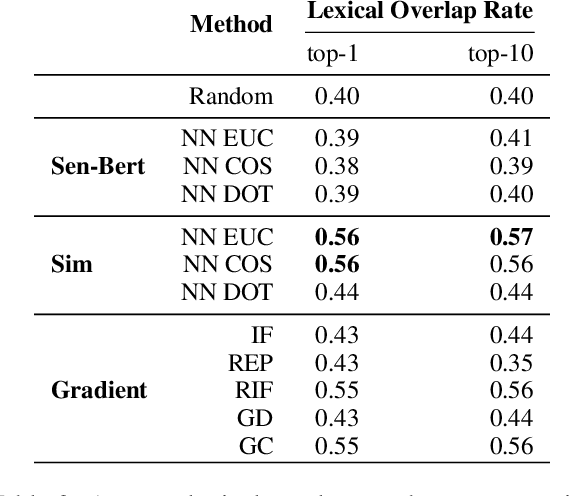
Abstract:Widespread adoption of deep models has motivated a pressing need for approaches to interpret network outputs and to facilitate model debugging. Instance attribution methods constitute one means of accomplishing these goals by retrieving training instances that (may have) led to a particular prediction. Influence functions (IF; Koh and Liang 2017) provide machinery for doing this by quantifying the effect that perturbing individual train instances would have on a specific test prediction. However, even approximating the IF is computationally expensive, to the degree that may be prohibitive in many cases. Might simpler approaches (e.g., retrieving train examples most similar to a given test point) perform comparably? In this work, we evaluate the degree to which different potential instance attribution agree with respect to the importance of training samples. We find that simple retrieval methods yield training instances that differ from those identified via gradient-based methods (such as IFs), but that nonetheless exhibit desirable characteristics similar to more complex attribution methods. Code for all methods and experiments in this paper is available at: https://github.com/successar/instance_attributions_NLP.
Unsupervised Data Augmentation with Naive Augmentation and without Unlabeled Data
Oct 22, 2020
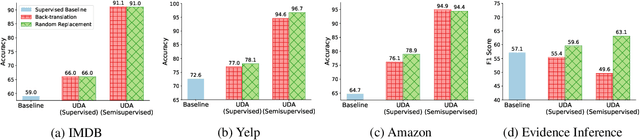
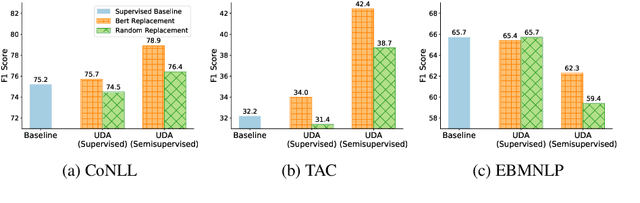

Abstract:Unsupervised Data Augmentation (UDA) is a semi-supervised technique that applies a consistency loss to penalize differences between a model's predictions on (a) observed (unlabeled) examples; and (b) corresponding 'noised' examples produced via data augmentation. While UDA has gained popularity for text classification, open questions linger over which design decisions are necessary and over how to extend the method to sequence labeling tasks. This method has recently gained traction for text classification. In this paper, we re-examine UDA and demonstrate its efficacy on several sequential tasks. Our main contribution is an empirical study of UDA to establish which components of the algorithm confer benefits in NLP. Notably, although prior work has emphasized the use of clever augmentation techniques including back-translation, we find that enforcing consistency between predictions assigned to observed and randomly substituted words often yields comparable (or greater) benefits compared to these complex perturbation models. Furthermore, we find that applying its consistency loss affords meaningful gains without any unlabeled data at all, i.e., in a standard supervised setting. In short: UDA need not be unsupervised, and does not require complex data augmentation to be effective.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge