Brooks Paige
Moment Matching Denoising Gibbs Sampling
May 19, 2023Abstract:Energy-Based Models (EBMs) offer a versatile framework for modeling complex data distributions. However, training and sampling from EBMs continue to pose significant challenges. The widely-used Denoising Score Matching (DSM) method for scalable EBM training suffers from inconsistency issues, causing the energy model to learn a `noisy' data distribution. In this work, we propose an efficient sampling framework: (pseudo)-Gibbs sampling with moment matching, which enables effective sampling from the underlying clean model when given a `noisy' model that has been well-trained via DSM. We explore the benefits of our approach compared to related methods and demonstrate how to scale the method to high-dimensional datasets.
Towards Healing the Blindness of Score Matching
Sep 15, 2022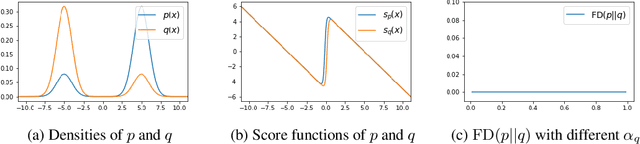
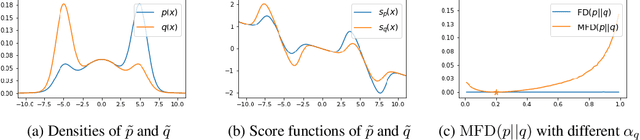
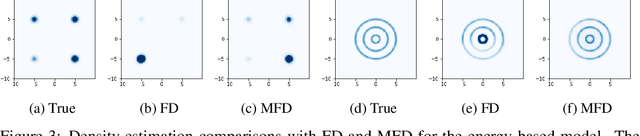
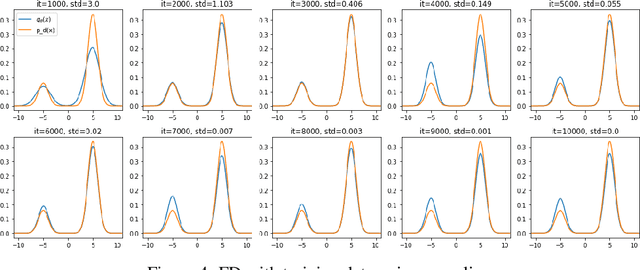
Abstract:Score-based divergences have been widely used in machine learning and statistics applications. Despite their empirical success, a blindness problem has been observed when using these for multi-modal distributions. In this work, we discuss the blindness problem and propose a new family of divergences that can mitigate the blindness problem. We illustrate our proposed divergence in the context of density estimation and report improved performance compared to traditional approaches.
Improving VAE-based Representation Learning
May 28, 2022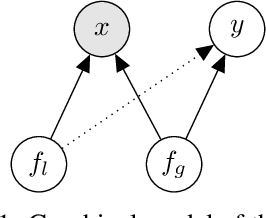
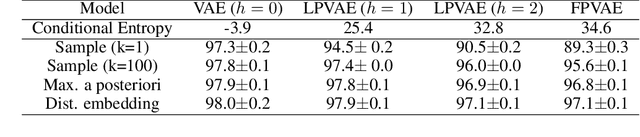

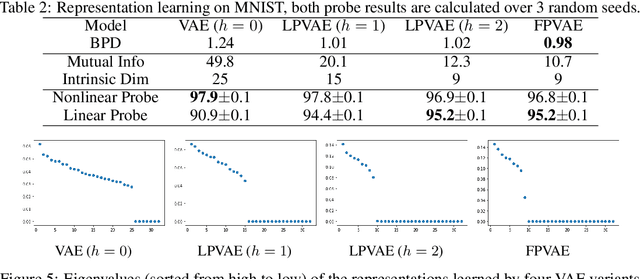
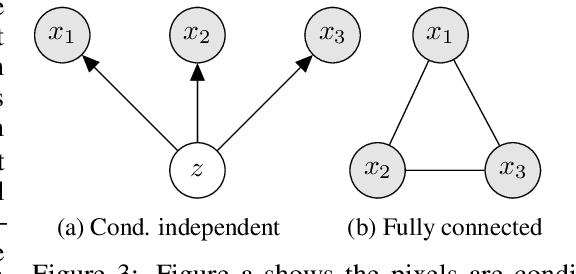
Abstract:Latent variable models like the Variational Auto-Encoder (VAE) are commonly used to learn representations of images. However, for downstream tasks like semantic classification, the representations learned by VAE are less competitive than other non-latent variable models. This has led to some speculations that latent variable models may be fundamentally unsuitable for representation learning. In this work, we study what properties are required for good representations and how different VAE structure choices could affect the learned properties. We show that by using a decoder that prefers to learn local features, the remaining global features can be well captured by the latent, which significantly improves performance of a downstream classification task. We further apply the proposed model to semi-supervised learning tasks and demonstrate improvements in data efficiency.
Simulation Intelligence: Towards a New Generation of Scientific Methods
Dec 06, 2021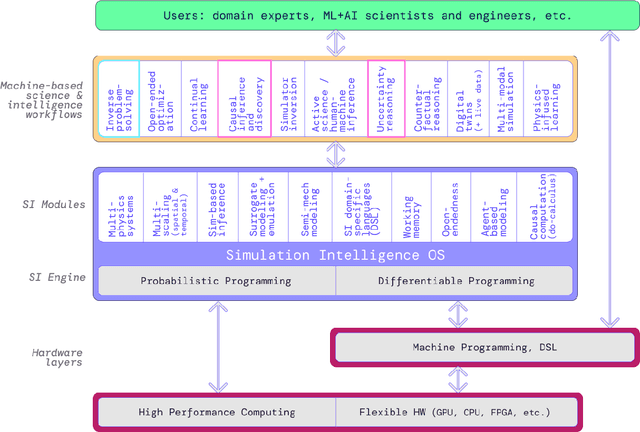
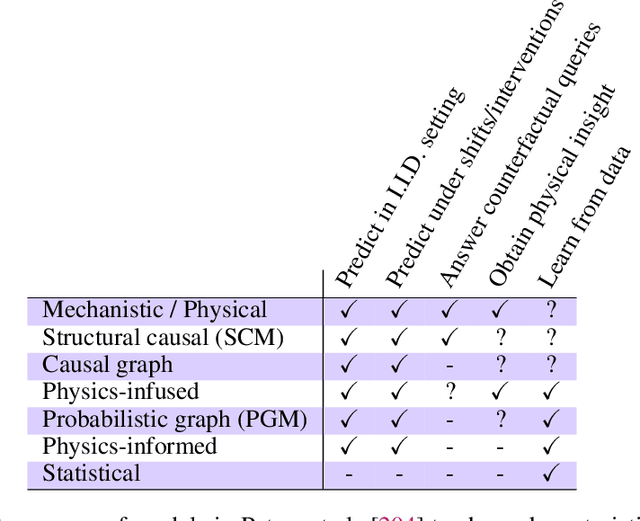
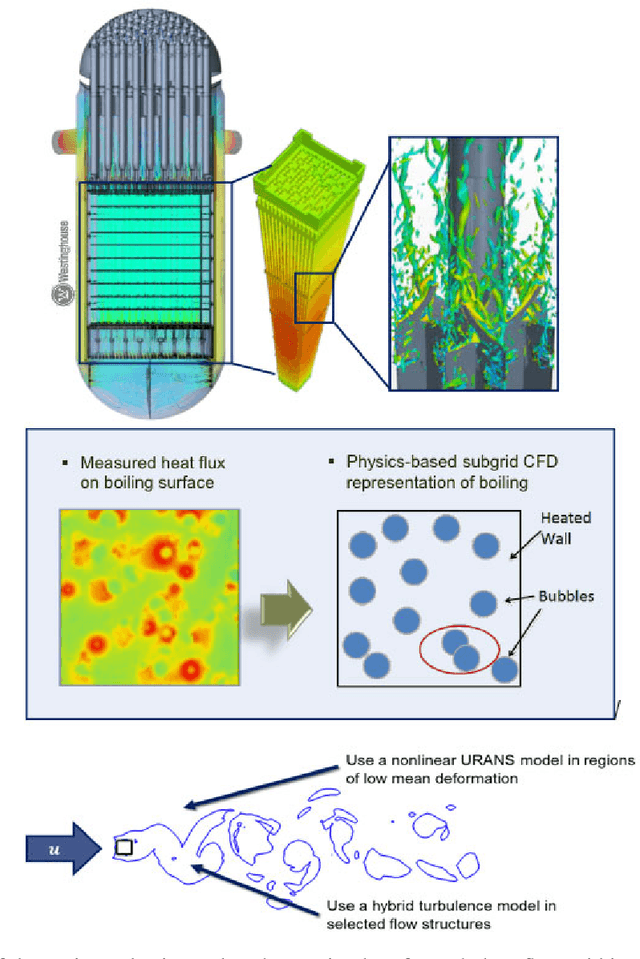
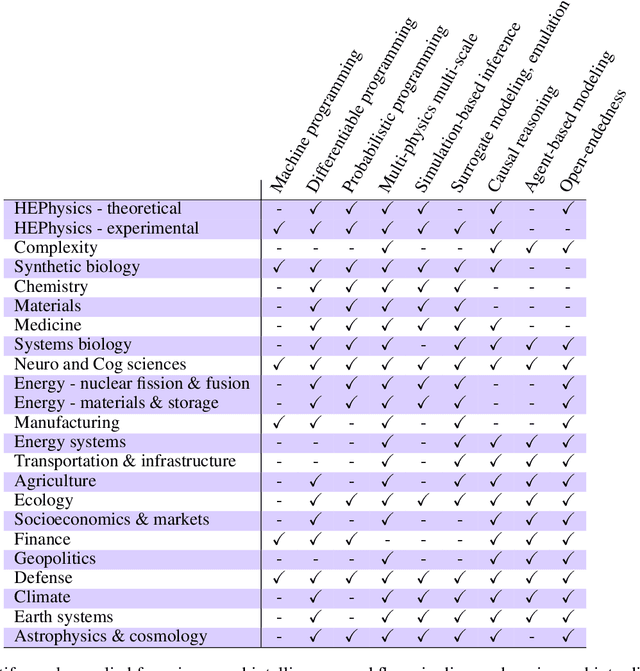
Abstract:The original "Seven Motifs" set forth a roadmap of essential methods for the field of scientific computing, where a motif is an algorithmic method that captures a pattern of computation and data movement. We present the "Nine Motifs of Simulation Intelligence", a roadmap for the development and integration of the essential algorithms necessary for a merger of scientific computing, scientific simulation, and artificial intelligence. We call this merger simulation intelligence (SI), for short. We argue the motifs of simulation intelligence are interconnected and interdependent, much like the components within the layers of an operating system. Using this metaphor, we explore the nature of each layer of the simulation intelligence operating system stack (SI-stack) and the motifs therein: (1) Multi-physics and multi-scale modeling; (2) Surrogate modeling and emulation; (3) Simulation-based inference; (4) Causal modeling and inference; (5) Agent-based modeling; (6) Probabilistic programming; (7) Differentiable programming; (8) Open-ended optimization; (9) Machine programming. We believe coordinated efforts between motifs offers immense opportunity to accelerate scientific discovery, from solving inverse problems in synthetic biology and climate science, to directing nuclear energy experiments and predicting emergent behavior in socioeconomic settings. We elaborate on each layer of the SI-stack, detailing the state-of-art methods, presenting examples to highlight challenges and opportunities, and advocating for specific ways to advance the motifs and the synergies from their combinations. Advancing and integrating these technologies can enable a robust and efficient hypothesis-simulation-analysis type of scientific method, which we introduce with several use-cases for human-machine teaming and automated science.
Fast and Scalable Spike and Slab Variable Selection in High-Dimensional Gaussian Processes
Nov 08, 2021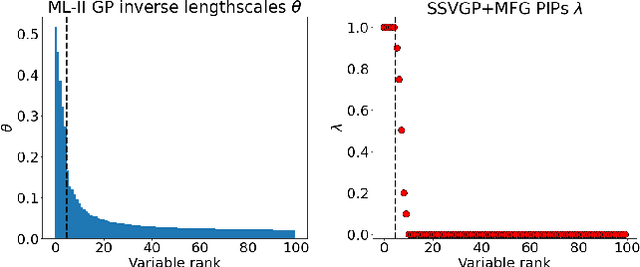
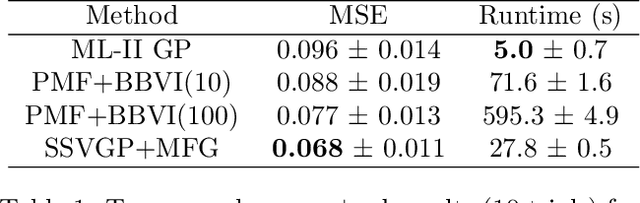
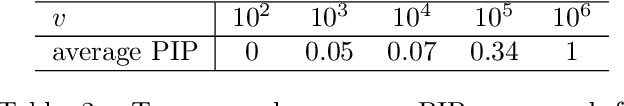
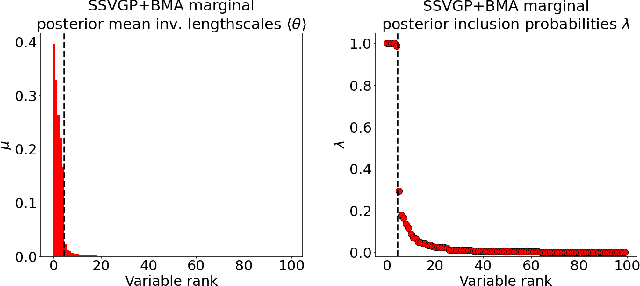
Abstract:Variable selection in Gaussian processes (GPs) is typically undertaken by thresholding the inverse lengthscales of `automatic relevance determination' kernels, but in high-dimensional datasets this approach can be unreliable. A more probabilistically principled alternative is to use spike and slab priors and infer a posterior probability of variable inclusion. However, existing implementations in GPs are extremely costly to run in both high-dimensional and large-$n$ datasets, or are intractable for most kernels. As such, we develop a fast and scalable variational inference algorithm for the spike and slab GP that is tractable with arbitrary differentiable kernels. We improve our algorithm's ability to adapt to the sparsity of relevant variables by Bayesian model averaging over hyperparameters, and achieve substantial speed ups using zero temperature posterior restrictions, dropout pruning and nearest neighbour minibatching. In experiments our method consistently outperforms vanilla and sparse variational GPs whilst retaining similar runtimes (even when $n=10^6$) and performs competitively with a spike and slab GP using MCMC but runs up to $1000$ times faster.
I Don't Need $\mathbf{u}$: Identifiable Non-Linear ICA Without Side Information
Jun 09, 2021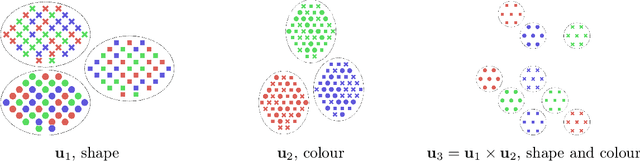
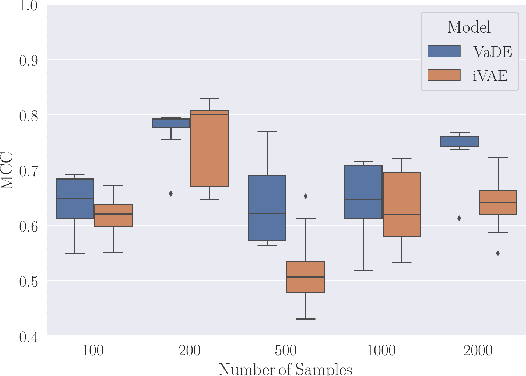
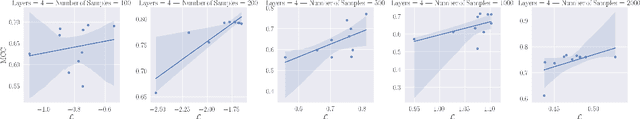
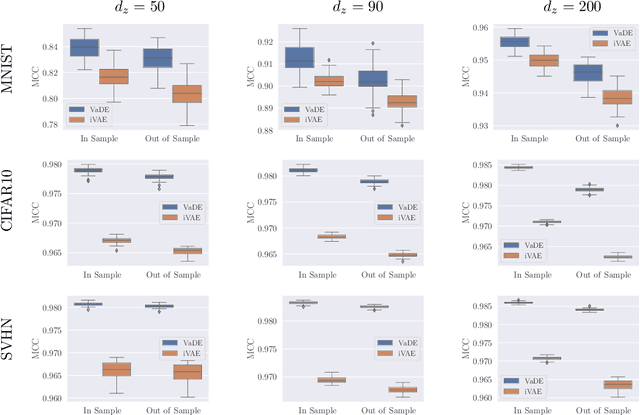
Abstract:In this work we introduce a new approach for identifiable non-linear ICA models. Recently there has been a renaissance in identifiability results in deep generative models, not least for non-linear ICA. These prior works, however, have assumed access to a sufficiently-informative auxiliary set of observations, denoted $\mathbf{u}$. We show here how identifiability can be obtained in the absence of this side-information, rendering possible fully-unsupervised identifiable non-linear ICA. While previous theoretical results have established the impossibility of identifiable non-linear ICA in the presence of infinitely-flexible universal function approximators, here we rely on the intrinsically-finite modelling capacity of any particular chosen parameterisation of a deep generative model. In particular, we focus on generative models which perform clustering in their latent space -- a model structure which matches previous identifiable models, but with the learnt clustering providing a synthetic form of auxiliary information. We evaluate our proposals using VAEs, on synthetic and image datasets, and find that the learned clusterings function effectively: deep generative models with latent clusterings are empirically identifiable, to the same degree as models which rely on side information.
Barking up the right tree: an approach to search over molecule synthesis DAGs
Dec 21, 2020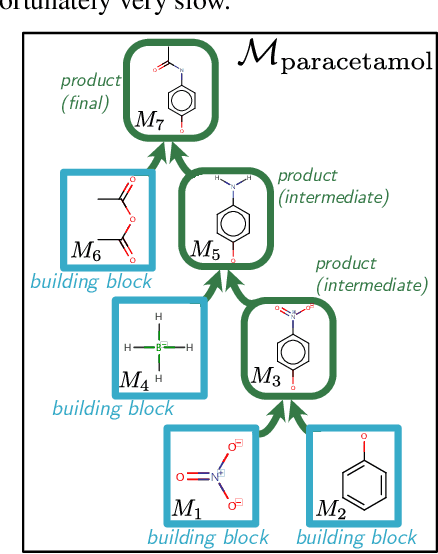
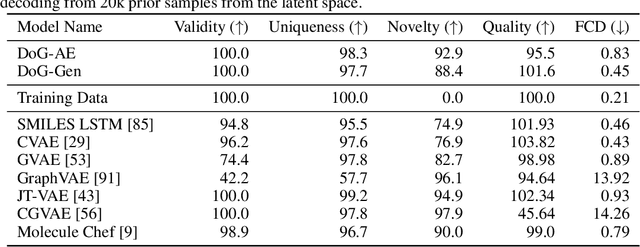
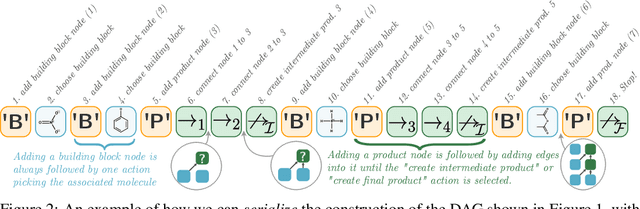
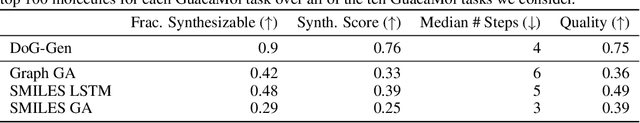
Abstract:When designing new molecules with particular properties, it is not only important what to make but crucially how to make it. These instructions form a synthesis directed acyclic graph (DAG), describing how a large vocabulary of simple building blocks can be recursively combined through chemical reactions to create more complicated molecules of interest. In contrast, many current deep generative models for molecules ignore synthesizability. We therefore propose a deep generative model that better represents the real world process, by directly outputting molecule synthesis DAGs. We argue that this provides sensible inductive biases, ensuring that our model searches over the same chemical space that chemists would also have access to, as well as interpretability. We show that our approach is able to model chemical space well, producing a wide range of diverse molecules, and allows for unconstrained optimization of an inherently constrained problem: maximize certain chemical properties such that discovered molecules are synthesizable.
Bayesian Graph Neural Networks for Molecular Property Prediction
Nov 25, 2020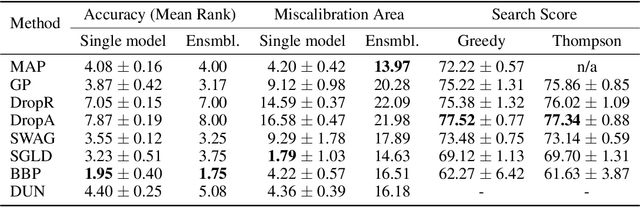
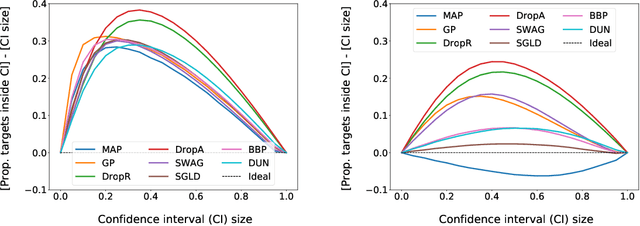
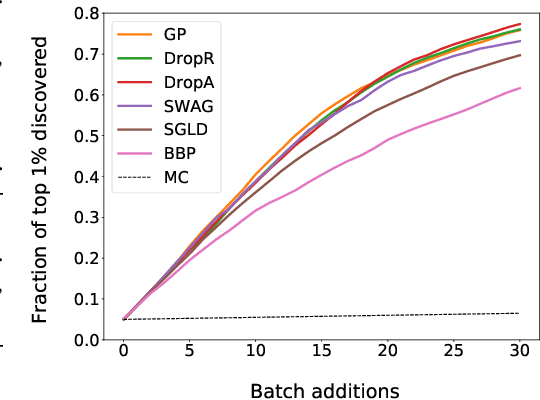
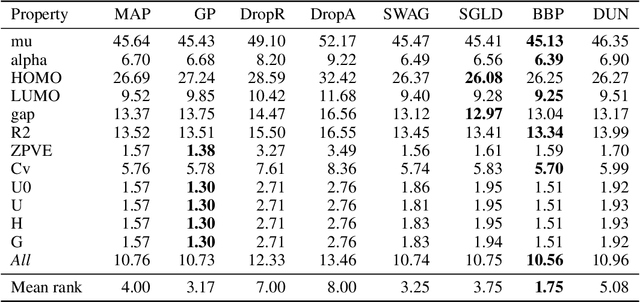
Abstract:Graph neural networks for molecular property prediction are frequently underspecified by data and fail to generalise to new scaffolds at test time. A potential solution is Bayesian learning, which can capture our uncertainty in the model parameters. This study benchmarks a set of Bayesian methods applied to a directed MPNN, using the QM9 regression dataset. We find that capturing uncertainty in both readout and message passing parameters yields enhanced predictive accuracy, calibration, and performance on a downstream molecular search task.
Making Graph Neural Networks Worth It for Low-Data Molecular Machine Learning
Nov 24, 2020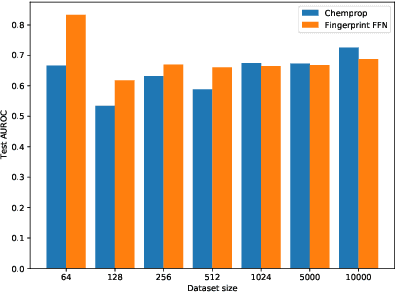
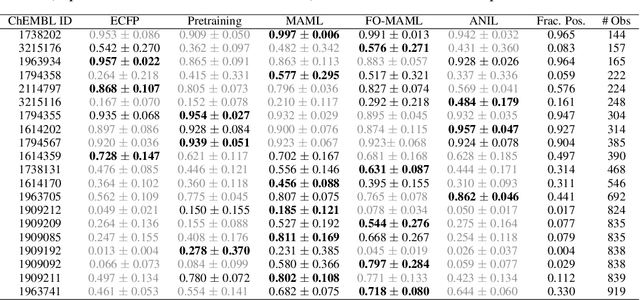
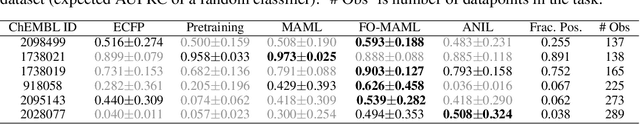
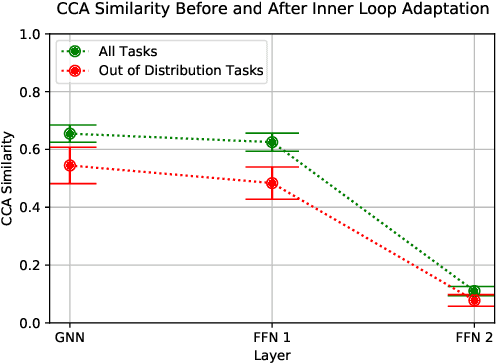
Abstract:Graph neural networks have become very popular for machine learning on molecules due to the expressive power of their learnt representations. However, molecular machine learning is a classically low-data regime and it isn't clear that graph neural networks can avoid overfitting in low-resource settings. In contrast, fingerprint methods are the traditional standard for low-data environments due to their reduced number of parameters and manually engineered features. In this work, we investigate whether graph neural networks are competitive in small data settings compared to the parametrically 'cheaper' alternative of fingerprint methods. When we find that they are not, we explore pretraining and the meta-learning method MAML (and variants FO-MAML and ANIL) for improving graph neural network performance by transfer learning from related tasks. We find that MAML and FO-MAML do enable the graph neural network to outperform models based on fingerprints, providing a path to using graph neural networks even in settings with severely restricted data availability. In contrast to previous work, we find ANIL performs worse that other meta-learning approaches in this molecule setting. Our results suggest two reasons: molecular machine learning tasks may require significant task-specific adaptation, and distribution shifts in test tasks relative to train tasks may contribute to worse ANIL performance.
Goal-directed Generation of Discrete Structures with Conditional Generative Models
Oct 23, 2020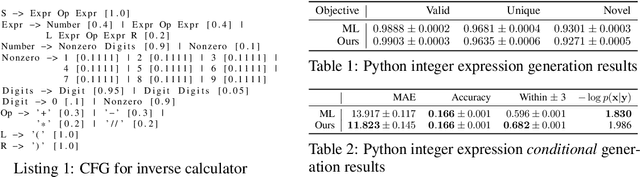
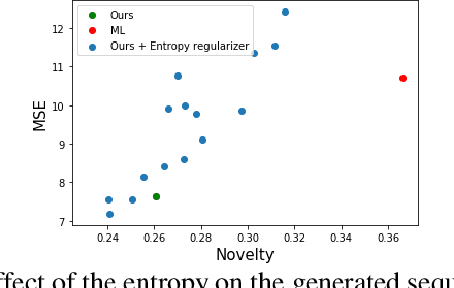

Abstract:Despite recent advances, goal-directed generation of structured discrete data remains challenging. For problems such as program synthesis (generating source code) and materials design (generating molecules), finding examples which satisfy desired constraints or exhibit desired properties is difficult. In practice, expensive heuristic search or reinforcement learning algorithms are often employed. In this paper we investigate the use of conditional generative models which directly attack this inverse problem, by modeling the distribution of discrete structures given properties of interest. Unfortunately, maximum likelihood training of such models often fails with the samples from the generative model inadequately respecting the input properties. To address this, we introduce a novel approach to directly optimize a reinforcement learning objective, maximizing an expected reward. We avoid high-variance score-function estimators that would otherwise be required by sampling from an approximation to the normalized rewards, allowing simple Monte Carlo estimation of model gradients. We test our methodology on two tasks: generating molecules with user-defined properties and identifying short python expressions which evaluate to a given target value. In both cases, we find improvements over maximum likelihood estimation and other baselines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge