Aydogan Ozcan
Scalable, Energy-Efficient Optical-Neural Architecture for Multiplexed Deepfake Video Detection
May 19, 2026Abstract:The rapid proliferation of AI-generated visual media has created an urgent need for efficient, trustworthy deepfake detection systems. However, existing deep learning-based detection methods rely on computationally intensive and energy-demanding inference algorithms, limiting their scalability. Here, we present a hybrid digital-analog deepfake video detection framework that combines a lightweight digital front-end with a spatially multiplexed optical decoding back-end for massively parallel analog inference through a programmable spatial light modulator. By simultaneously processing 15 or more video streams within a single optical propagation pass, the system enables high-throughput and accurate video-level authenticity prediction at reduced computational cost compared with purely digital methods. We validated this hybrid deepfake video processor using different datasets spanning classical face-swapping, real-world deepfake recordings, and fully AI-generated videos. Using a spatially multiplexed experimental set-up operating in the visible spectrum, we achieved average deepfake detection accuracy, sensitivity and specificity of 97.79%, 99.86% and 95.72%, respectively, on the Celeb-DF video dataset with 15 videos tested in parallel in a single optical pass per inference. The multiplexed optical decoder also demonstrates resilience against various types of video degradation, noise, compression, experimental misalignments and black-box adversarial attacks. Our results show that integrating optical computation into AI inference enables simultaneous gains in throughput, energy efficiency, and adversarial robustness - three properties that are difficult to achieve together in purely digital systems.
Continuous quantification of viral plaque dynamics using ultra-large-area label-free imaging enables rapid antiviral susceptibility testing
May 03, 2026Abstract:The plaque reduction assay (PRA) remains the gold standard for antiviral susceptibility testing, evaluating drug potency by measuring reductions in plaque-forming units (PFUs). However, the traditional PRA is time-consuming, labor-intensive, prone to manual counting errors, and offers limited scalability. Moreover, its reliance on destructive fixation and chemical staining reduces the assay to a static, endpoint observation, obscuring the dynamic, time-resolved kinetics of dose-dependent viral inhibition. Here, we introduce a label-free, time-resolved PRA platform that transforms the conventional assay into a continuous, high-dimensional measurement of viral infection dynamics. Our system integrates a compact lens-free imaging setup with a custom-designed ultra-large-area (100 cm^2) thin-film transistor (TFT) image sensor and deep learning-based algorithms to autonomously quantify PFU dynamics within an incubator. Validated using herpes simplex virus type-1 (HSV-1) treated with acyclovir, the platform matched chemically-stained ground truth measurements with zero false positives while accelerating readout by ~26 hours. Crucially, our system revealed that increasing drug concentrations induce temporally distinct delays and suppress new PFU formation, enabling conclusive drug efficacy evaluations within ~60 hours post-infection. This scalable, label-free framework redefines antiviral susceptibility testing as a rapid, time-resolved and information-rich measurement framework, providing a generalizable platform for virology research, high-throughput drug screening, and clinical diagnostics.
Wavelength-multiplexed massively parallel diffractive optical information storage and image projection
Apr 03, 2026Abstract:We introduce a wavelength-multiplexed massively parallel diffractive information storage platform composed of dielectric surfaces that are structurally optimized at the wavelength scale using deep learning to store and project thousands of distinct image patterns, each assigned to a unique wavelength. Through numerical simulations in the visible spectrum, we demonstrated that our wavelength-multiplexed diffractive system can store and project over 4,000 independent desired images/patterns within its output field-of-view, with high image quality and minimal crosstalk between spectral channels. Furthermore, in a proof-of-concept experiment, we demonstrated a two-layer diffractive design that stored six distinct patterns and projected them onto the same output field of view at six different wavelengths (500, 548, 596, 644, 692, and 740 nm). This diffractive architecture is scalable and can operate at various parts of the electromagnetic spectrum without the need for material dispersion engineering or redesigning its optimized diffractive layers. The demonstrated storage capacity, reconstruction image fidelity, and wavelength-encoded massively parallel read-out of our diffractive platform offer a compact and fast-access solution for large-scale optical information storage, image projection applications.
Large-scale nonlinear optical computing with incoherent light via linear diffractive systems
Mar 31, 2026Abstract:Nonlinear computation is essential for various information processing tasks. Optical implementations are attractive because passive light propagation can manipulate high-dimensional signals with extreme throughput and parallelism; yet realizing nonlinear mappings in optical hardware remains challenging due to the weak nonlinearity of optical materials and the large intensities required to induce nonlinear interactions. This challenge is further amplified in many systems that operate with incoherent illumination, motivating a coherence-aware framework for scalable optical nonlinear processing. Here, we show that linear optical systems, in particular, optimized diffractive processors comprising passive surfaces, can perform large-scale nonlinear function approximation under spatially incoherent or partially coherent illumination, when preceded by intensity-only input encoding. We quantify how the accuracy of the nonlinear function approximation varies with the degree of parallelism, the number of diffractive layers, and the number of trainable diffractive features. Numerical results demonstrate snapshot computation of up to one million distinct nonlinear functions in a single forward pass through a diffractive processor, with the function outputs spatially multiplexed and read out using densely packed detectors at the output. We further provide a proof-of-concept experimental demonstration under incoherent illumination from a liquid crystal display (LCD), enabled by a model-free in situ learning strategy that jointly optimizes the diffractive profile and detector readout geometry in the presence of hardware imperfections and misalignments. Our findings establish diffractive processors as a massively parallel universal function approximator for both spatially incoherent and partially coherent illumination.
Compressive single-pixel imaging via a wavelength-multiplexed spatially incoherent diffractive optical processor
Mar 23, 2026Abstract:Despite offering high sensitivity, a high signal-to-noise ratio, and a broad spectral range, single-pixel imaging (SPI) is limited by low measurement efficiency and long data-acquisition times. To address this, we propose a wavelength-multiplexed, spatially incoherent diffractive optical processor combined with a compact/shallow digital artificial neural network (ANN) to implement compressive SPI. Specifically, we model the bucket detection process in conventional SPI as a linear intensity transformation with spatially and spectrally varying point-spread functions. This transformation matrix is treated as a learnable parameter and jointly optimized with a shallow digital ANN composed of 2 hidden nonlinear layers. The wavelength-multiplexed diffractive processor is then configured via data-free optimization to approximate this pre-trained transformation matrix; after this optimization, the diffractive processor remains static/fixed. Upon multi-wavelength illumination and diffractive modulation, the target spatial information of the input object is spectrally encoded. A single-pixel detector captures the output spectral power at each illumination band, which is then rapidly decoded by the jointly trained digital ANN to reconstruct the input image. In addition to our numerical analyses demonstrating the feasibility of this approach, we experimentally validated its proof-of-concept using an array of light-emitting diodes (LEDs). Overall, this work demonstrates a computational imaging framework for compressive SPI that can be useful in applications such as biomedical imaging, autonomous devices, and remote sensing.
Automated HER2 scoring with uncertainty quantification using lensfree holography and deep learning
Jan 26, 2026Abstract:Accurate assessment of human epidermal growth factor receptor 2 (HER2) expression is critical for breast cancer diagnosis, prognosis, and therapy selection; yet, most existing digital HER2 scoring methods rely on bulky and expensive optical systems. Here, we present a compact and cost-effective lensfree holography platform integrated with deep learning for automated HER2 scoring of immunohistochemically stained breast tissue sections. The system captures lensfree diffraction patterns of stained HER2 tissue sections under RGB laser illumination and acquires complex field information over a sample area of ~1,250 mm^2 at an effective throughput of ~84 mm^2 per minute. To enhance diagnostic reliability, we incorporated an uncertainty quantification strategy based on Bayesian Monte Carlo dropout, which provides autonomous uncertainty estimates for each prediction and supports reliable, robust HER2 scoring, with an overall correction rate of 30.4%. Using a blinded test set of 412 unique tissue samples, our approach achieved a testing accuracy of 84.9% for 4-class (0, 1+, 2+, 3+) HER2 classification and 94.8% for binary (0/1+ vs. 2+/3+) HER2 scoring with uncertainty quantification. Overall, this lensfree holography approach provides a practical pathway toward portable, high-throughput, and cost-effective HER2 scoring, particularly suited for resource-limited settings, where traditional digital pathology infrastructure is unavailable.
Autonomous Uncertainty Quantification for Computational Point-of-care Sensors
Dec 24, 2025Abstract:Computational point-of-care (POC) sensors enable rapid, low-cost, and accessible diagnostics in emergency, remote and resource-limited areas that lack access to centralized medical facilities. These systems can utilize neural network-based algorithms to accurately infer a diagnosis from the signals generated by rapid diagnostic tests or sensors. However, neural network-based diagnostic models are subject to hallucinations and can produce erroneous predictions, posing a risk of misdiagnosis and inaccurate clinical decisions. To address this challenge, here we present an autonomous uncertainty quantification technique developed for POC diagnostics. As our testbed, we used a paper-based, computational vertical flow assay (xVFA) platform developed for rapid POC diagnosis of Lyme disease, the most prevalent tick-borne disease globally. The xVFA platform integrates a disposable paper-based assay, a handheld optical reader and a neural network-based inference algorithm, providing rapid and cost-effective Lyme disease diagnostics in under 20 min using only 20 uL of patient serum. By incorporating a Monte Carlo dropout (MCDO)-based uncertainty quantification approach into the diagnostics pipeline, we identified and excluded erroneous predictions with high uncertainty, significantly improving the sensitivity and reliability of the xVFA in an autonomous manner, without access to the ground truth diagnostic information of patients. Blinded testing using new patient samples demonstrated an increase in diagnostic sensitivity from 88.2% to 95.7%, indicating the effectiveness of MCDO-based uncertainty quantification in enhancing the robustness of neural network-driven computational POC sensing systems.
Deep learning-enhanced dual-mode multiplexed optical sensor for point-of-care diagnostics of cardiovascular diseases
Dec 24, 2025Abstract:Rapid and accessible cardiac biomarker testing is essential for the timely diagnosis and risk assessment of myocardial infarction (MI) and heart failure (HF), two interrelated conditions that frequently coexist and drive recurrent hospitalizations with high mortality. However, current laboratory and point-of-care testing systems are limited by long turnaround times, narrow dynamic ranges for the tested biomarkers, and single-analyte formats that fail to capture the complexity of cardiovascular disease. Here, we present a deep learning-enhanced dual-mode multiplexed vertical flow assay (xVFA) with a portable optical reader and a neural network-based quantification pipeline. This optical sensor integrates colorimetric and chemiluminescent detection within a single paper-based cartridge to complementarily cover a large dynamic range (spanning ~6 orders of magnitude) for both low- and high-abundance biomarkers, while maintaining quantitative accuracy. Using 50 uL of serum, the optical sensor simultaneously quantifies cardiac troponin I (cTnI), creatine kinase-MB (CK-MB), and N-terminal pro-B-type natriuretic peptide (NT-proBNP) within 23 min. The xVFA achieves sub-pg/mL sensitivity for cTnI and sub-ng/mL sensitivity for CK-MB and NT-proBNP, spanning the clinically relevant ranges for these biomarkers. Neural network models trained and blindly tested on 92 patient serum samples yielded a robust quantification performance (Pearson's r > 0.96 vs. reference assays). By combining high sensitivity, multiplexing, and automation in a compact and cost-effective optical sensor format, the dual-mode xVFA enables rapid and quantitative cardiovascular diagnostics at the point of care.
Snapshot 3D image projection using a diffractive decoder
Dec 23, 2025Abstract:3D image display is essential for next-generation volumetric imaging; however, dense depth multiplexing for 3D image projection remains challenging because diffraction-induced cross-talk rapidly increases as the axial image planes get closer. Here, we introduce a 3D display system comprising a digital encoder and a diffractive optical decoder, which simultaneously projects different images onto multiple target axial planes with high axial resolution. By leveraging multi-layer diffractive wavefront decoding and deep learning-based end-to-end optimization, the system achieves high-fidelity depth-resolved 3D image projection in a snapshot, enabling axial plane separations on the order of a wavelength. The digital encoder leverages a Fourier encoder network to capture multi-scale spatial and frequency-domain features from input images, integrates axial position encoding, and generates a unified phase representation that simultaneously encodes all images to be axially projected in a single snapshot through a jointly-optimized diffractive decoder. We characterized the impact of diffractive decoder depth, output diffraction efficiency, spatial light modulator resolution, and axial encoding density, revealing trade-offs that govern axial separation and 3D image projection quality. We further demonstrated the capability to display volumetric images containing 28 axial slices, as well as the ability to dynamically reconfigure the axial locations of the image planes, performed on demand. Finally, we experimentally validated the presented approach, demonstrating close agreement between the measured results and the target images. These results establish the diffractive 3D display system as a compact and scalable framework for depth-resolved snapshot 3D image projection, with potential applications in holographic displays, AR/VR interfaces, and volumetric optical computing.
Pixel Super-Resolved Fluorescence Lifetime Imaging Using Deep Learning
Dec 18, 2025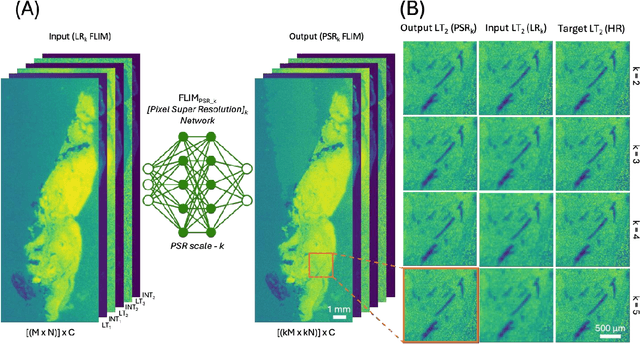
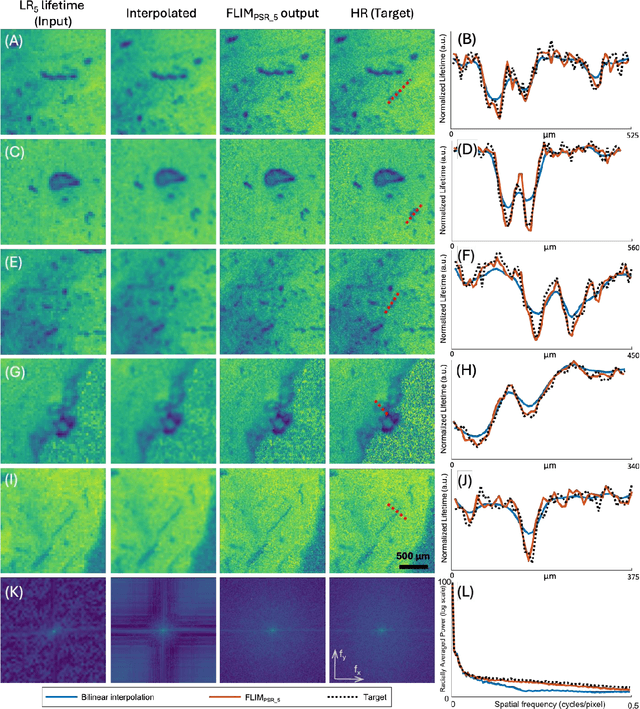
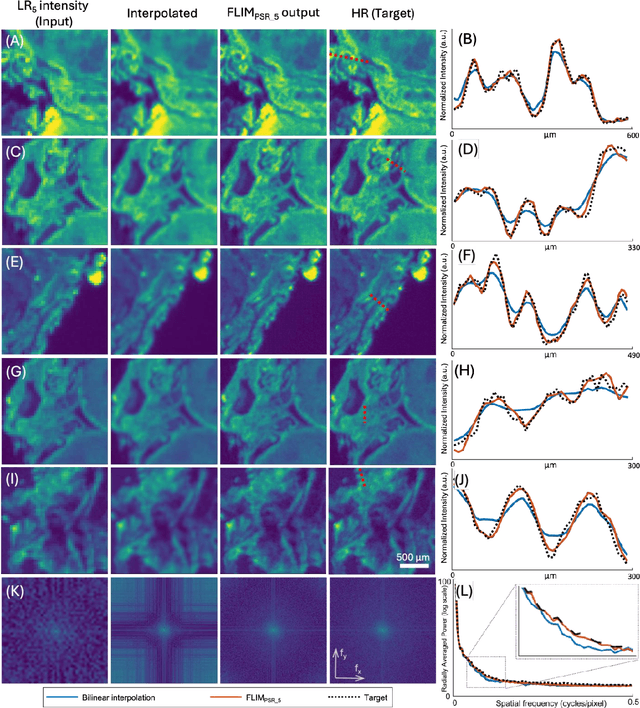
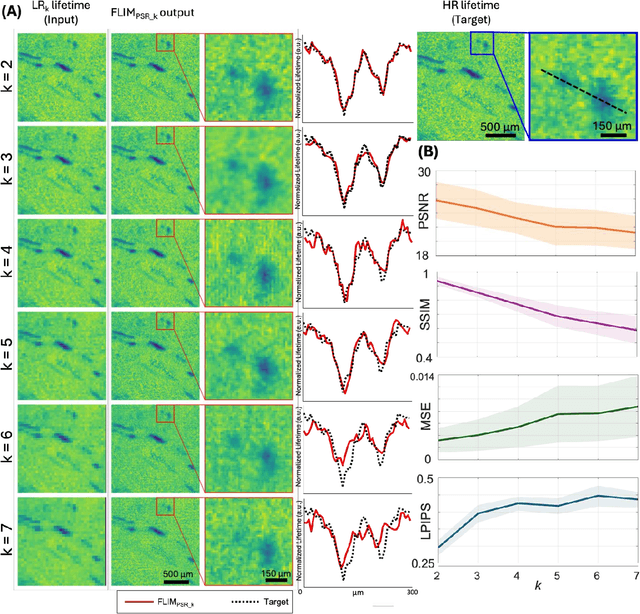
Abstract:Fluorescence lifetime imaging microscopy (FLIM) is a powerful quantitative technique that provides metabolic and molecular contrast, offering strong translational potential for label-free, real-time diagnostics. However, its clinical adoption remains limited by long pixel dwell times and low signal-to-noise ratio (SNR), which impose a stricter resolution-speed trade-off than conventional optical imaging approaches. Here, we introduce FLIM_PSR_k, a deep learning-based multi-channel pixel super-resolution (PSR) framework that reconstructs high-resolution FLIM images from data acquired with up to a 5-fold increased pixel size. The model is trained using the conditional generative adversarial network (cGAN) framework, which, compared to diffusion model-based alternatives, delivers a more robust PSR reconstruction with substantially shorter inference times, a crucial advantage for practical deployment. FLIM_PSR_k not only enables faster image acquisition but can also alleviate SNR limitations in autofluorescence-based FLIM. Blind testing on held-out patient-derived tumor tissue samples demonstrates that FLIM_PSR_k reliably achieves a super-resolution factor of k = 5, resulting in a 25-fold increase in the space-bandwidth product of the output images and revealing fine architectural features lost in lower-resolution inputs, with statistically significant improvements across various image quality metrics. By increasing FLIM's effective spatial resolution, FLIM_PSR_k advances lifetime imaging toward faster, higher-resolution, and hardware-flexible implementations compatible with low-numerical-aperture and miniaturized platforms, better positioning FLIM for translational applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge