Antonino Miceli
Unsupervised Semantic Segmentation in Synchrotron Computed Tomography with Self-Correcting Pseudo Labels
Feb 27, 2026Abstract:X-ray computed tomography (CT) is a widely used imaging technique that provides detailed examinations into the internal structure of an object with synchrotron CT (SR-CT) enabling improved data quality by using higher energy, monochromatic X-rays. While SR-CT allows for improved resolution, time-resolved experimentation, and reduced imaging artifacts, it also produces significantly larger datasets than conventional CT. Accurate and efficient evaluation of these datasets is a critical component of these workflows; yet is often done manually representing a major bottleneck in the analysis phase. While deep learning has emerged as a powerful tool capable of providing a wide range of purely data-driven solutions, it requires a substantial amount of labeled data for training and manual annotation of SR-CT datasets is impractical in practice. In this paper, we introduce a novel framework that enables automatic segmentation of large, high-resolution SR-CT datasets by eliminating the need to hand label images for deep learning training. First, we generate pseudo labels by clustering on the voxel values identifying regions in the volume with similar attenuation coefficients producing an initial semantic map. Afterwards, we train a segmentation model on the pseudo labels before utilizing the Unbiased Teacher approach to self-correct them ensuring accurate final segmentations. We find our approach improves pixel-wise accuracy and mIoU by 13.31% and 15.94%, respectively, over the baseline pseudo labels when using a magnesium crystal SR-CT sample. Additionally, we extensively evaluate the different components of our workflow including segmentation model, loss function, pseudo labeling strategy, and input type. Finally, we evaluate our approach on to two additional samples highlighting our frameworks ability to produce segmentations that are considerably better than the original pseudo labels.
Rapid detection of rare events from in situ X-ray diffraction data using machine learning
Dec 07, 2023



Abstract:High-energy X-ray diffraction methods can non-destructively map the 3D microstructure and associated attributes of metallic polycrystalline engineering materials in their bulk form. These methods are often combined with external stimuli such as thermo-mechanical loading to take snapshots over time of the evolving microstructure and attributes. However, the extreme data volumes and the high costs of traditional data acquisition and reduction approaches pose a barrier to quickly extracting actionable insights and improving the temporal resolution of these snapshots. Here we present a fully automated technique capable of rapidly detecting the onset of plasticity in high-energy X-ray microscopy data. Our technique is computationally faster by at least 50 times than the traditional approaches and works for data sets that are up to 9 times sparser than a full data set. This new technique leverages self-supervised image representation learning and clustering to transform massive data into compact, semantic-rich representations of visually salient characteristics (e.g., peak shapes). These characteristics can be a rapid indicator of anomalous events such as changes in diffraction peak shapes. We anticipate that this technique will provide just-in-time actionable information to drive smarter experiments that effectively deploy multi-modal X-ray diffraction methods that span many decades of length scales.
AI-assisted Automated Workflow for Real-time X-ray Ptychography Data Analysis via Federated Resources
Apr 09, 2023
Abstract:We present an end-to-end automated workflow that uses large-scale remote compute resources and an embedded GPU platform at the edge to enable AI/ML-accelerated real-time analysis of data collected for x-ray ptychography. Ptychography is a lensless method that is being used to image samples through a simultaneous numerical inversion of a large number of diffraction patterns from adjacent overlapping scan positions. This acquisition method can enable nanoscale imaging with x-rays and electrons, but this often requires very large experimental datasets and commensurately high turnaround times, which can limit experimental capabilities such as real-time experimental steering and low-latency monitoring. In this work, we introduce a software system that can automate ptychography data analysis tasks. We accelerate the data analysis pipeline by using a modified version of PtychoNN -- an ML-based approach to solve phase retrieval problem that shows two orders of magnitude speedup compared to traditional iterative methods. Further, our system coordinates and overlaps different data analysis tasks to minimize synchronization overhead between different stages of the workflow. We evaluate our workflow system with real-world experimental workloads from the 26ID beamline at Advanced Photon Source and ThetaGPU cluster at Argonne Leadership Computing Resources.
Deep learning at the edge enables real-time streaming ptychographic imaging
Sep 20, 2022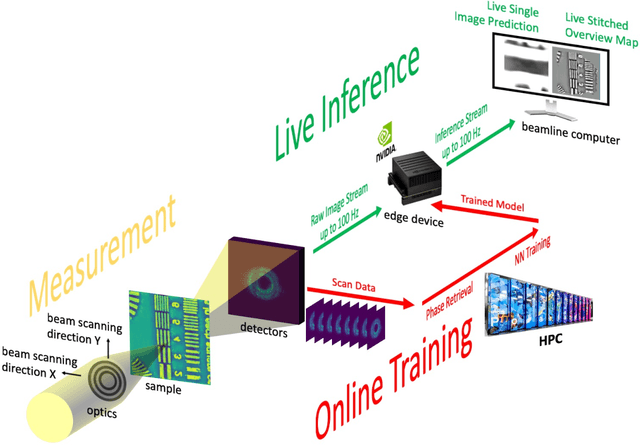
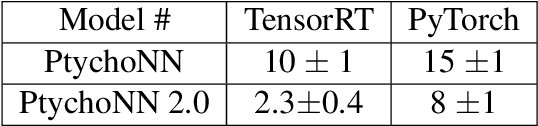
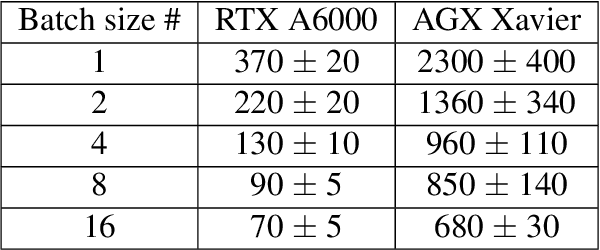
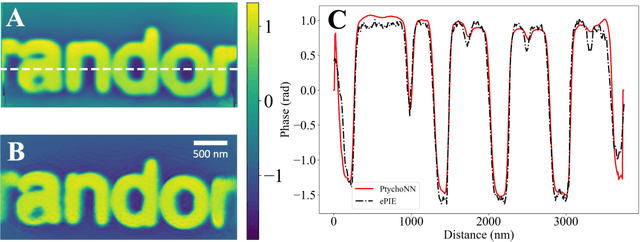

Abstract:Coherent microscopy techniques provide an unparalleled multi-scale view of materials across scientific and technological fields, from structural materials to quantum devices, from integrated circuits to biological cells. Driven by the construction of brighter sources and high-rate detectors, coherent X-ray microscopy methods like ptychography are poised to revolutionize nanoscale materials characterization. However, associated significant increases in data and compute needs mean that conventional approaches no longer suffice for recovering sample images in real-time from high-speed coherent imaging experiments. Here, we demonstrate a workflow that leverages artificial intelligence at the edge and high-performance computing to enable real-time inversion on X-ray ptychography data streamed directly from a detector at up to 2 kHz. The proposed AI-enabled workflow eliminates the sampling constraints imposed by traditional ptychography, allowing low dose imaging using orders of magnitude less data than required by traditional methods.
fairDMS: Rapid Model Training by Data and Model Reuse
Apr 20, 2022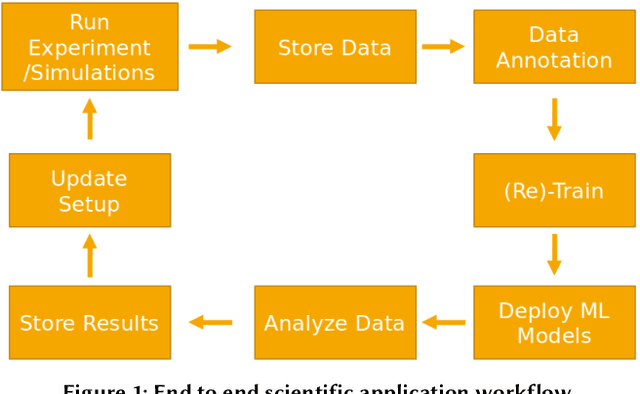
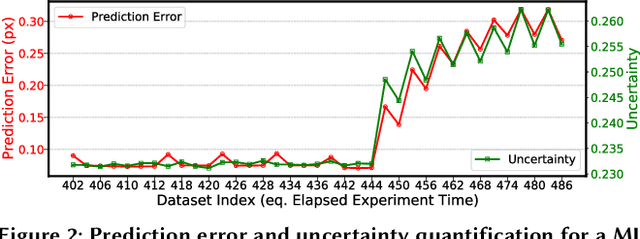
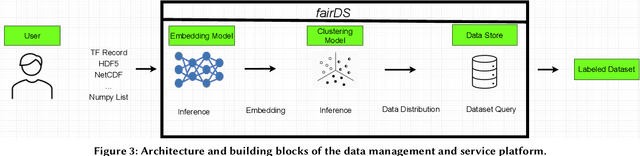
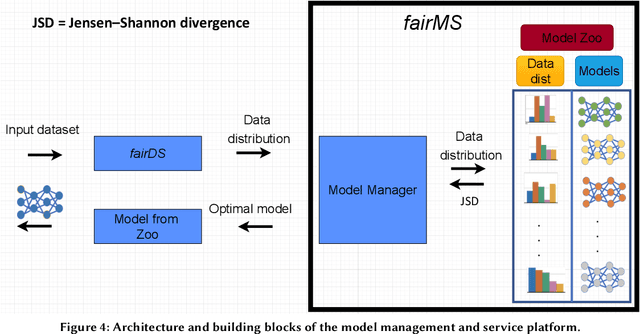
Abstract:Extracting actionable information from data sources such as the Linac Coherent Light Source (LCLS-II) and Advanced Photon Source Upgrade (APS-U) is becoming more challenging due to the fast-growing data generation rate. The rapid analysis possible with ML methods can enable fast feedback loops that can be used to adjust experimental setups in real-time, for example when errors occur or interesting events are detected. However, to avoid degradation in ML performance over time due to changes in an instrument or sample, we need a way to update ML models rapidly while an experiment is running. We present here a data service and model service to accelerate deep neural network training with a focus on ML-based scientific applications. Our proposed data service achieves 100x speedup in terms of data labeling compare to the current state-of-the-art. Further, our model service achieves up to 200x improvement in training speed. Overall, fairDMS achieves up to 92x speedup in terms of end-to-end model updating time.
Bridge Data Center AI Systems with Edge Computing for Actionable Information Retrieval
May 28, 2021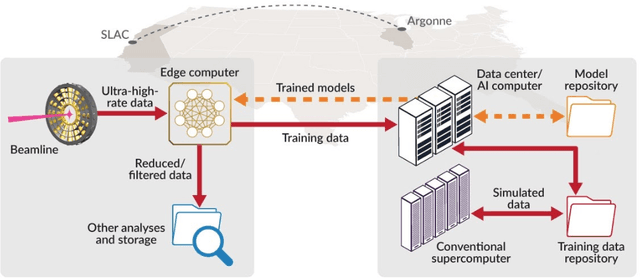

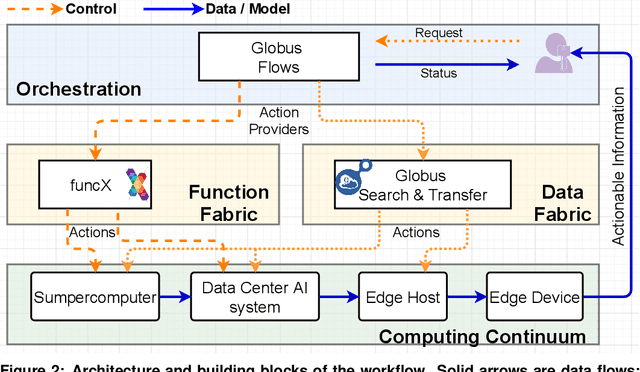
Abstract:Extremely high data rates at modern synchrotron and X-ray free-electron lasers (XFELs) light source beamlines motivate the use of machine learning methods for data reduction, feature detection, and other purposes. Regardless of the application, the basic concept is the same: data collected in early stages of an experiment, data from past similar experiments, and/or data simulated for the upcoming experiment are used to train machine learning models that, in effect, learn specific characteristics of those data; these models are then used to process subsequent data more efficiently than would general-purpose models that lack knowledge of the specific dataset or data class. Thus, a key challenge is to be able to train models with sufficient rapidity that they can be deployed and used within useful timescales. We describe here how specialized data center AI systems can be used for this purpose.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge