Jun-Sang Park
Unsupervised Semantic Segmentation in Synchrotron Computed Tomography with Self-Correcting Pseudo Labels
Feb 27, 2026Abstract:X-ray computed tomography (CT) is a widely used imaging technique that provides detailed examinations into the internal structure of an object with synchrotron CT (SR-CT) enabling improved data quality by using higher energy, monochromatic X-rays. While SR-CT allows for improved resolution, time-resolved experimentation, and reduced imaging artifacts, it also produces significantly larger datasets than conventional CT. Accurate and efficient evaluation of these datasets is a critical component of these workflows; yet is often done manually representing a major bottleneck in the analysis phase. While deep learning has emerged as a powerful tool capable of providing a wide range of purely data-driven solutions, it requires a substantial amount of labeled data for training and manual annotation of SR-CT datasets is impractical in practice. In this paper, we introduce a novel framework that enables automatic segmentation of large, high-resolution SR-CT datasets by eliminating the need to hand label images for deep learning training. First, we generate pseudo labels by clustering on the voxel values identifying regions in the volume with similar attenuation coefficients producing an initial semantic map. Afterwards, we train a segmentation model on the pseudo labels before utilizing the Unbiased Teacher approach to self-correct them ensuring accurate final segmentations. We find our approach improves pixel-wise accuracy and mIoU by 13.31% and 15.94%, respectively, over the baseline pseudo labels when using a magnesium crystal SR-CT sample. Additionally, we extensively evaluate the different components of our workflow including segmentation model, loss function, pseudo labeling strategy, and input type. Finally, we evaluate our approach on to two additional samples highlighting our frameworks ability to produce segmentations that are considerably better than the original pseudo labels.
Rapid detection of rare events from in situ X-ray diffraction data using machine learning
Dec 07, 2023



Abstract:High-energy X-ray diffraction methods can non-destructively map the 3D microstructure and associated attributes of metallic polycrystalline engineering materials in their bulk form. These methods are often combined with external stimuli such as thermo-mechanical loading to take snapshots over time of the evolving microstructure and attributes. However, the extreme data volumes and the high costs of traditional data acquisition and reduction approaches pose a barrier to quickly extracting actionable insights and improving the temporal resolution of these snapshots. Here we present a fully automated technique capable of rapidly detecting the onset of plasticity in high-energy X-ray microscopy data. Our technique is computationally faster by at least 50 times than the traditional approaches and works for data sets that are up to 9 times sparser than a full data set. This new technique leverages self-supervised image representation learning and clustering to transform massive data into compact, semantic-rich representations of visually salient characteristics (e.g., peak shapes). These characteristics can be a rapid indicator of anomalous events such as changes in diffraction peak shapes. We anticipate that this technique will provide just-in-time actionable information to drive smarter experiments that effectively deploy multi-modal X-ray diffraction methods that span many decades of length scales.
BraggNN: Fast X-ray Bragg Peak Analysis Using Deep Learning
Aug 18, 2020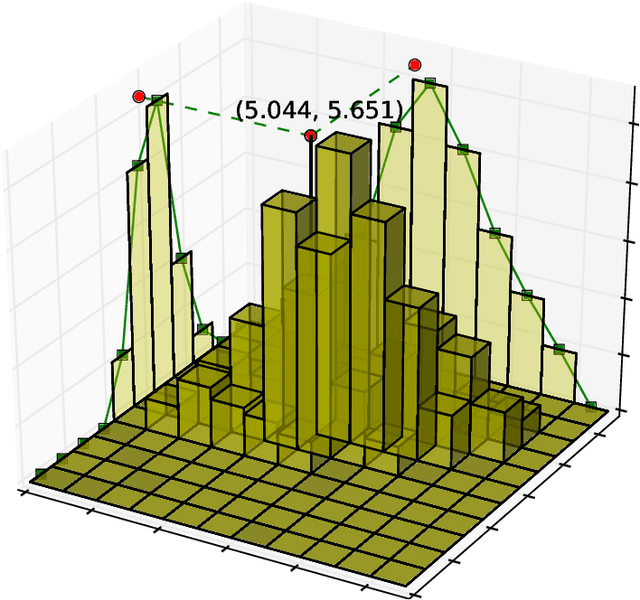
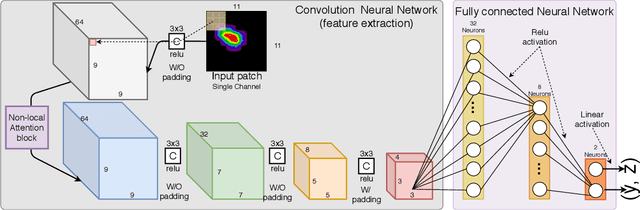
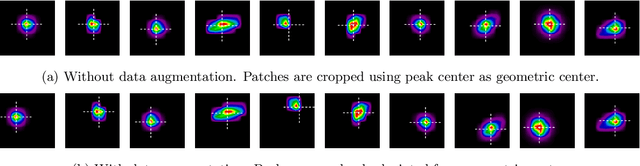
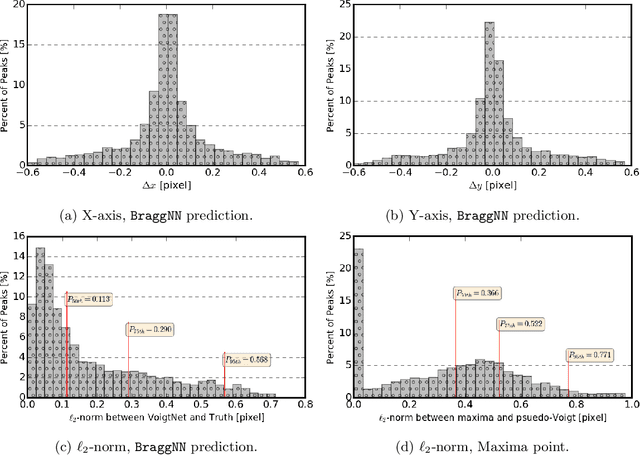
Abstract:X-ray diffraction based microscopy techniques such as high energy diffraction microscopy rely on knowledge of position of diffraction peaks with high resolution. These positions are typically computed by fitting the observed intensities in detector data to a theoretical peak shape such as pseudo-Voigt. As experiments become more complex and detector technologies evolve, the computational cost of such peak shape fitting becomes the biggest hurdle to the rapid analysis required for real-time feedback for experiments. To this end, this paper proposes BraggNN, a machine learning-based method that can localize Bragg peak much more rapidly than conventional pseudo-Voigt peak fitting. When applied to our test dataset, BraggNN gives errors of less than 0.29 and 0.57 voxels, relative to conventional method, for 75% and 95% of the peaks, respectively. When applied to a real experiment dataset, a 3D reconstruction using peak positions located by BraggNN yields an average grain position difference of 17 micrometer and size difference of 1.3 micrometer as compared to the results obtained when the reconstruction used peaks from conventional 2D pseudo-Voigt fitting. Recent advances in deep learning method implementations and special-purpose model inference accelerators allow BraggNN to deliver enormous performance improvements relative to the conventional method, running, for example, more than 200 times faster than a conventional method when using a GPU card with out-of-the-box software.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge