Andrew Zhang
A multimodal and temporal foundation model for virtual patient representations at healthcare system scale
Apr 21, 2026Abstract:Modern medicine generates vast multimodal data across siloed systems, yet no existing model integrates the full breadth and temporal depth of the clinical record into a unified patient representation. We introduce Apollo, a multimodal temporal foundation model trained and evaluated on over three decades of longitudinal hospital records from a major US hospital system, composed of 25 billion records from 7.2 million patients, representing 28 distinct medical modalities and 12 major medical specialties. Apollo learns a unified representation space integrating over 100 thousand unique medical events in our clinical vocabulary as well as images and clinical text. This "atlas of medical concepts" forms a computational substrate for modeling entire patient care journeys comprised of sequences of structured and unstructured events, which are compressed by Apollo into virtual patient representations. To assess the potential of these whole-patient representations, we created 322 prognosis and retrieval tasks from a held-out test set of 1.4 million patients. We demonstrate the generalized clinical forecasting potential of Apollo embeddings, including predicting new disease onset risk up to five years in advance (95 tasks), disease progression (78 tasks), treatment response (59 tasks), risk of treatment-related adverse events (17 tasks), and hospital operations endpoints (12 tasks). Using feature attribution techniques, we show that model predictions align with clinically-interpretable multimodal biomarkers. We evaluate semantic similarity search on 61 retrieval tasks, and moreover demonstrate the potential of Apollo as a multimodal medical search engine using text and image queries. Together, these modeling capabilities establish the foundation for computable medicine, where the full context of patient care becomes accessible to computational reasoning.
Persona Vectors in Games: Measuring and Steering Strategies via Activation Vectors
Mar 22, 2026Abstract:Large language models (LLMs) are increasingly deployed as autonomous decision-makers in strategic settings, yet we have limited tools for understanding their high-level behavioral traits. We use activation steering methods in game-theoretic settings, constructing persona vectors for altruism, forgiveness, and expectations of others by contrastive activation addition. Evaluating on canonical games, we find that activation steering systematically shifts both quantitative strategic choices and natural-language justifications. However, we also observe that rhetoric and strategy can diverge under steering. In addition, vectors for self-behavior and expectations of others are partially distinct. Our results suggest that persona vectors offer a promising mechanistic handle on high-level traits in strategic environments.
SteerVLM: Robust Model Control through Lightweight Activation Steering for Vision Language Models
Oct 30, 2025Abstract:This work introduces SteerVLM, a lightweight steering module designed to guide Vision-Language Models (VLMs) towards outputs that better adhere to desired instructions. Our approach learns from the latent embeddings of paired prompts encoding target and converse behaviors to dynamically adjust activations connecting the language modality with image context. This allows for fine-grained, inference-time control over complex output semantics without modifying model weights while preserving performance on off-target tasks. Our steering module requires learning parameters equal to 0.14% of the original VLM's size. Our steering module gains model control through dimension-wise activation modulation and adaptive steering across layers without requiring pre-extracted static vectors or manual tuning of intervention points. Furthermore, we introduce VNIA (Visual Narrative Intent Alignment), a multimodal dataset specifically created to facilitate the development and evaluation of VLM steering techniques. Our method outperforms existing intervention techniques on steering and hallucination mitigation benchmarks for VLMs and proposes a robust solution for multimodal model control through activation engineering.
Maximal Matching Matters: Preventing Representation Collapse for Robust Cross-Modal Retrieval
Jun 26, 2025



Abstract:Cross-modal image-text retrieval is challenging because of the diverse possible associations between content from different modalities. Traditional methods learn a single-vector embedding to represent semantics of each sample, but struggle to capture nuanced and diverse relationships that can exist across modalities. Set-based approaches, which represent each sample with multiple embeddings, offer a promising alternative, as they can capture richer and more diverse relationships. In this paper, we show that, despite their promise, these set-based representations continue to face issues including sparse supervision and set collapse, which limits their effectiveness. To address these challenges, we propose Maximal Pair Assignment Similarity to optimize one-to-one matching between embedding sets which preserve semantic diversity within the set. We also introduce two loss functions to further enhance the representations: Global Discriminative Loss to enhance distinction among embeddings, and Intra-Set Divergence Loss to prevent collapse within each set. Our method achieves state-of-the-art performance on MS-COCO and Flickr30k without relying on external data.
Flexible-length Text Infilling for Discrete Diffusion Models
Jun 16, 2025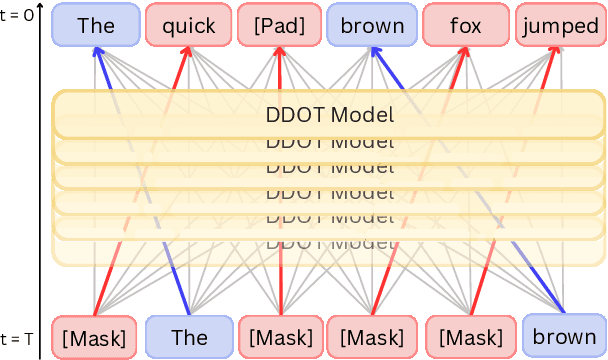

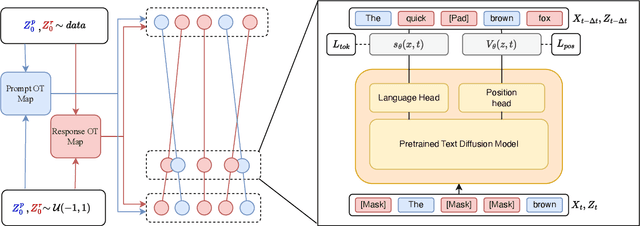

Abstract:Discrete diffusion models are a new class of text generators that offer advantages such as bidirectional context use, parallelizable generation, and flexible prompting compared to autoregressive models. However, a critical limitation of discrete diffusion models is their inability to perform flexible-length or flexible-position text infilling without access to ground-truth positional data. We introduce \textbf{DDOT} (\textbf{D}iscrete \textbf{D}iffusion with \textbf{O}ptimal \textbf{T}ransport Position Coupling), the first discrete diffusion model to overcome this challenge. DDOT jointly denoises token values and token positions, employing a novel sample-level Optimal Transport (OT) coupling. This coupling preserves relative token ordering while dynamically adjusting the positions and length of infilled segments, a capability previously missing in text diffusion. Our method is orthogonal to existing discrete text diffusion methods and is compatible with various pretrained text denoisers. Extensive experiments on text infilling benchmarks such as One-Billion-Word and Yelp demonstrate that DDOT outperforms naive diffusion baselines. Furthermore, DDOT achieves performance on par with state-of-the-art non-autoregressive models and enables significant improvements in training efficiency and flexibility.
MAISY: Motion-Aware Image SYnthesis for Medical Image Motion Correction
May 08, 2025Abstract:Patient motion during medical image acquisition causes blurring, ghosting, and distorts organs, which makes image interpretation challenging. Current state-of-the-art algorithms using Generative Adversarial Network (GAN)-based methods with their ability to learn the mappings between corrupted images and their ground truth via Structural Similarity Index Measure (SSIM) loss effectively generate motion-free images. However, we identified the following limitations: (i) they mainly focus on global structural characteristics and therefore overlook localized features that often carry critical pathological information, and (ii) the SSIM loss function struggles to handle images with varying pixel intensities, luminance factors, and variance. In this study, we propose Motion-Aware Image SYnthesis (MAISY) which initially characterize motion and then uses it for correction by: (a) leveraging the foundation model Segment Anything Model (SAM), to dynamically learn spatial patterns along anatomical boundaries where motion artifacts are most pronounced and, (b) introducing the Variance-Selective SSIM (VS-SSIM) loss which adaptively emphasizes spatial regions with high pixel variance to preserve essential anatomical details during artifact correction. Experiments on chest and head CT datasets demonstrate that our model outperformed the state-of-the-art counterparts, with Peak Signal-to-Noise Ratio (PSNR) increasing by 40%, SSIM by 10%, and Dice by 16%.
Accelerating Data Processing and Benchmarking of AI Models for Pathology
Feb 10, 2025Abstract:Advances in foundation modeling have reshaped computational pathology. However, the increasing number of available models and lack of standardized benchmarks make it increasingly complex to assess their strengths, limitations, and potential for further development. To address these challenges, we introduce a new suite of software tools for whole-slide image processing, foundation model benchmarking, and curated publicly available tasks. We anticipate that these resources will promote transparency, reproducibility, and continued progress in the field.
Molecular-driven Foundation Model for Oncologic Pathology
Jan 28, 2025Abstract:Foundation models are reshaping computational pathology by enabling transfer learning, where models pre-trained on vast datasets can be adapted for downstream diagnostic, prognostic, and therapeutic response tasks. Despite these advances, foundation models are still limited in their ability to encode the entire gigapixel whole-slide images without additional training and often lack complementary multimodal data. Here, we introduce Threads, a slide-level foundation model capable of generating universal representations of whole-slide images of any size. Threads was pre-trained using a multimodal learning approach on a diverse cohort of 47,171 hematoxylin and eosin (H&E)-stained tissue sections, paired with corresponding genomic and transcriptomic profiles - the largest such paired dataset to be used for foundation model development to date. This unique training paradigm enables Threads to capture the tissue's underlying molecular composition, yielding powerful representations applicable to a wide array of downstream tasks. In extensive benchmarking across 54 oncology tasks, including clinical subtyping, grading, mutation prediction, immunohistochemistry status determination, treatment response prediction, and survival prediction, Threads outperformed all baselines while demonstrating remarkable generalizability and label efficiency. It is particularly well suited for predicting rare events, further emphasizing its clinical utility. We intend to make the model publicly available for the broader community.
Multimodal Whole Slide Foundation Model for Pathology
Nov 29, 2024



Abstract:The field of computational pathology has been transformed with recent advances in foundation models that encode histopathology region-of-interests (ROIs) into versatile and transferable feature representations via self-supervised learning (SSL). However, translating these advancements to address complex clinical challenges at the patient and slide level remains constrained by limited clinical data in disease-specific cohorts, especially for rare clinical conditions. We propose TITAN, a multimodal whole slide foundation model pretrained using 335,645 WSIs via visual self-supervised learning and vision-language alignment with corresponding pathology reports and 423,122 synthetic captions generated from a multimodal generative AI copilot for pathology. Without any finetuning or requiring clinical labels, TITAN can extract general-purpose slide representations and generate pathology reports that generalize to resource-limited clinical scenarios such as rare disease retrieval and cancer prognosis. We evaluate TITAN on diverse clinical tasks and find that TITAN outperforms both ROI and slide foundation models across machine learning settings such as linear probing, few-shot and zero-shot classification, rare cancer retrieval and cross-modal retrieval, and pathology report generation.
Multistain Pretraining for Slide Representation Learning in Pathology
Aug 05, 2024Abstract:Developing self-supervised learning (SSL) models that can learn universal and transferable representations of H&E gigapixel whole-slide images (WSIs) is becoming increasingly valuable in computational pathology. These models hold the potential to advance critical tasks such as few-shot classification, slide retrieval, and patient stratification. Existing approaches for slide representation learning extend the principles of SSL from small images (e.g., 224 x 224 patches) to entire slides, usually by aligning two different augmentations (or views) of the slide. Yet the resulting representation remains constrained by the limited clinical and biological diversity of the views. Instead, we postulate that slides stained with multiple markers, such as immunohistochemistry, can be used as different views to form a rich task-agnostic training signal. To this end, we introduce Madeleine, a multimodal pretraining strategy for slide representation learning. Madeleine is trained with a dual global-local cross-stain alignment objective on large cohorts of breast cancer samples (N=4,211 WSIs across five stains) and kidney transplant samples (N=12,070 WSIs across four stains). We demonstrate the quality of slide representations learned by Madeleine on various downstream evaluations, ranging from morphological and molecular classification to prognostic prediction, comprising 21 tasks using 7,299 WSIs from multiple medical centers. Code is available at https://github.com/mahmoodlab/MADELEINE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge