Alexandre Gramfort
MIND - INRIA
MultiView Independent Component Analysis with Delays
Dec 01, 2023Abstract:Linear Independent Component Analysis (ICA) is a blind source separation technique that has been used in various domains to identify independent latent sources from observed signals. In order to obtain a higher signal-to-noise ratio, the presence of multiple views of the same sources can be used. In this work, we present MultiView Independent Component Analysis with Delays (MVICAD). This algorithm builds on the MultiView ICA model by allowing sources to be delayed versions of some shared sources: sources are shared across views up to some unknown latencies that are view- and source-specific. Using simulations, we demonstrate that MVICAD leads to better unmixing of the sources. Moreover, as ICA is often used in neuroscience, we show that latencies are age-related when applied to Cam-CAN, a large-scale magnetoencephalography (MEG) dataset. These results demonstrate that the MVICAD model can reveal rich effects on neural signals without human supervision.
Evaluating the structure of cognitive tasks with transfer learning
Jul 28, 2023



Abstract:Electroencephalography (EEG) decoding is a challenging task due to the limited availability of labelled data. While transfer learning is a promising technique to address this challenge, it assumes that transferable data domains and task are known, which is not the case in this setting. This study investigates the transferability of deep learning representations between different EEG decoding tasks. We conduct extensive experiments using state-of-the-art decoding models on two recently released EEG datasets, ERP CORE and M$^3$CV, containing over 140 subjects and 11 distinct cognitive tasks. We measure the transferability of learned representations by pre-training deep neural networks on one task and assessing their ability to decode subsequent tasks. Our experiments demonstrate that, even with linear probing transfer, significant improvements in decoding performance can be obtained, with gains of up to 28% compare with the pure supervised approach. Additionally, we discover evidence that certain decoding paradigms elicit specific and narrow brain activities, while others benefit from pre-training on a broad range of representations. By revealing which tasks transfer well and demonstrating the benefits of transfer learning for EEG decoding, our findings have practical implications for mitigating data scarcity in this setting. The transfer maps generated also provide insights into the hierarchical relations between cognitive tasks, hence enhancing our understanding of how these tasks are connected from a neuroscientific standpoint.
L-C2ST: Local Diagnostics for Posterior Approximations in Simulation-Based Inference
Jun 06, 2023Abstract:Many recent works in simulation-based inference (SBI) rely on deep generative models to approximate complex, high-dimensional posterior distributions. However, evaluating whether or not these approximations can be trusted remains a challenge. Most approaches evaluate the posterior estimator only in expectation over the observation space. This limits their interpretability and is not sufficient to identify for which observations the approximation can be trusted or should be improved. Building upon the well-known classifier two-sample test (C2ST), we introduce L-C2ST, a new method that allows for a local evaluation of the posterior estimator at any given observation. It offers theoretically grounded and easy to interpret - e.g. graphical - diagnostics, and unlike C2ST, does not require access to samples from the true posterior. In the case of normalizing flow-based posterior estimators, L-C2ST can be specialized to offer better statistical power, while being computationally more efficient. On standard SBI benchmarks, L-C2ST provides comparable results to C2ST and outperforms alternative local approaches such as coverage tests based on highest predictive density (HPD). We further highlight the importance of local evaluation and the benefit of interpretability of L-C2ST on a challenging application from computational neuroscience.
Convolutional Monge Mapping Normalization for learning on biosignals
May 31, 2023Abstract:In many machine learning applications on signals and biomedical data, especially electroencephalogram (EEG), one major challenge is the variability of the data across subjects, sessions, and hardware devices. In this work, we propose a new method called Convolutional Monge Mapping Normalization (CMMN), which consists in filtering the signals in order to adapt their power spectrum density (PSD) to a Wasserstein barycenter estimated on training data. CMMN relies on novel closed-form solutions for optimal transport mappings and barycenters and provides individual test time adaptation to new data without needing to retrain a prediction model. Numerical experiments on sleep EEG data show that CMMN leads to significant and consistent performance gains independent from the neural network architecture when adapting between subjects, sessions, and even datasets collected with different hardware. Notably our performance gain is on par with much more numerically intensive Domain Adaptation (DA) methods and can be used in conjunction with those for even better performances.
Optimizing the Noise in Self-Supervised Learning: from Importance Sampling to Noise-Contrastive Estimation
Jan 23, 2023Abstract:Self-supervised learning is an increasingly popular approach to unsupervised learning, achieving state-of-the-art results. A prevalent approach consists in contrasting data points and noise points within a classification task: this requires a good noise distribution which is notoriously hard to specify. While a comprehensive theory is missing, it is widely assumed that the optimal noise distribution should in practice be made equal to the data distribution, as in Generative Adversarial Networks (GANs). We here empirically and theoretically challenge this assumption. We turn to Noise-Contrastive Estimation (NCE) which grounds this self-supervised task as an estimation problem of an energy-based model of the data. This ties the optimality of the noise distribution to the sample efficiency of the estimator, which is rigorously defined as its asymptotic variance, or mean-squared error. In the special case where the normalization constant only is unknown, we show that NCE recovers a family of Importance Sampling estimators for which the optimal noise is indeed equal to the data distribution. However, in the general case where the energy is also unknown, we prove that the optimal noise density is the data density multiplied by a correction term based on the Fisher score. In particular, the optimal noise distribution is different from the data distribution, and is even from a different family. Nevertheless, we soberly conclude that the optimal noise may be hard to sample from, and the gain in efficiency can be modest compared to choosing the noise distribution equal to the data's.
Validation Diagnostics for SBI algorithms based on Normalizing Flows
Nov 24, 2022Abstract:Building on the recent trend of new deep generative models known as Normalizing Flows (NF), simulation-based inference (SBI) algorithms can now efficiently accommodate arbitrary complex and high-dimensional data distributions. The development of appropriate validation methods however has fallen behind. Indeed, most of the existing metrics either require access to the true posterior distribution, or fail to provide theoretical guarantees on the consistency of the inferred approximation beyond the one-dimensional setting. This work proposes easy to interpret validation diagnostics for multi-dimensional conditional (posterior) density estimators based on NF. It also offers theoretical guarantees based on results of local consistency. The proposed workflow can be used to check, analyse and guarantee consistent behavior of the estimator. The method is illustrated with a challenging example that involves tightly coupled parameters in the context of computational neuroscience. This work should help the design of better specified models or drive the development of novel SBI-algorithms, hence allowing to build up trust on their ability to address important questions in experimental science.
FaDIn: Fast Discretized Inference for Hawkes Processes with General Parametric Kernels
Oct 10, 2022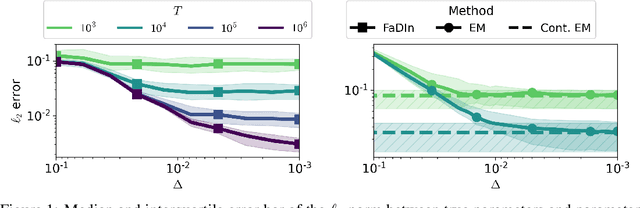
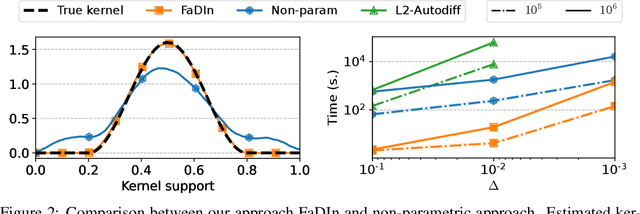
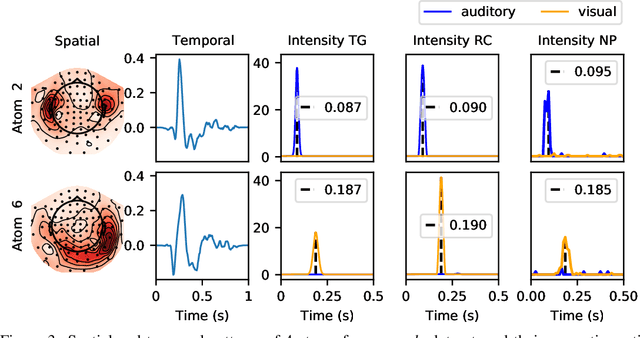
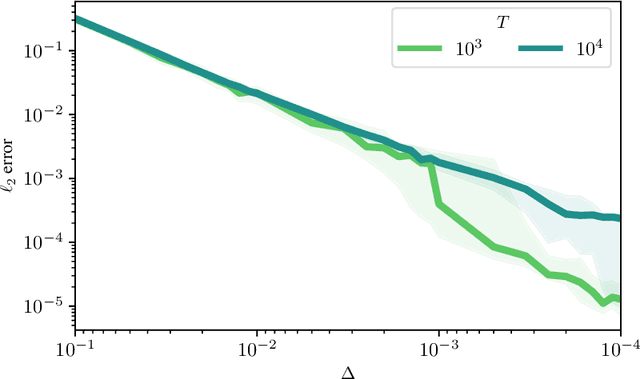
Abstract:Temporal point processes (TPP) are a natural tool for modeling event-based data. Among all TPP models, Hawkes processes have proven to be the most widely used, mainly due to their simplicity and computational ease when considering exponential or non-parametric kernels. Although non-parametric kernels are an option, such models require large datasets. While exponential kernels are more data efficient and relevant for certain applications where events immediately trigger more events, they are ill-suited for applications where latencies need to be estimated, such as in neuroscience. This work aims to offer an efficient solution to TPP inference using general parametric kernels with finite support. The developed solution consists of a fast L2 gradient-based solver leveraging a discretized version of the events. After supporting the use of discretization theoretically, the statistical and computational efficiency of the novel approach is demonstrated through various numerical experiments. Finally, the effectiveness of the method is evaluated by modeling the occurrence of stimuli-induced patterns from brain signals recorded with magnetoencephalography (MEG). Given the use of general parametric kernels, results show that the proposed approach leads to a more plausible estimation of pattern latency compared to the state-of-the-art.
Data augmentation for learning predictive models on EEG: a systematic comparison
Jun 29, 2022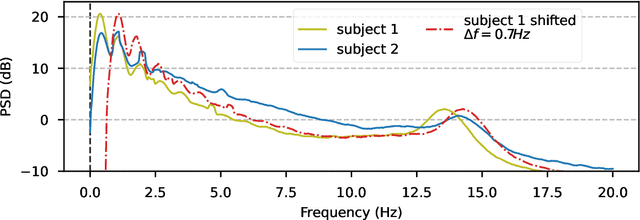

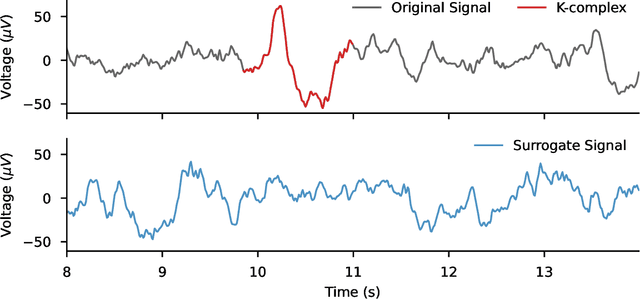

Abstract:The use of deep learning for electroencephalography (EEG) classification tasks has been rapidly growing in the last years, yet its application has been limited by the relatively small size of EEG datasets. Data augmentation, which consists in artificially increasing the size of the dataset during training, has been a key ingredient to obtain state-of-the-art performances across applications such as computer vision or speech. While a few augmentation transformations for EEG data have been proposed in the literature, their positive impact on performance across tasks remains elusive. In this work, we propose a unified and exhaustive analysis of the main existing EEG augmentations, which are compared in a common experimental setting. Our results highlight the best data augmentations to consider for sleep stage classification and motor imagery brain computer interfaces, showing predictive power improvements greater than 10% in some cases.
Benchopt: Reproducible, efficient and collaborative optimization benchmarks
Jun 28, 2022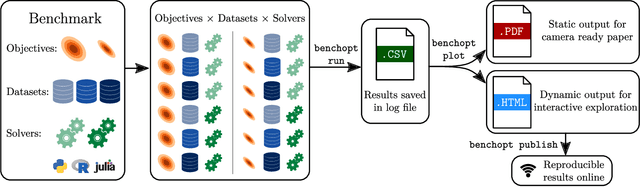
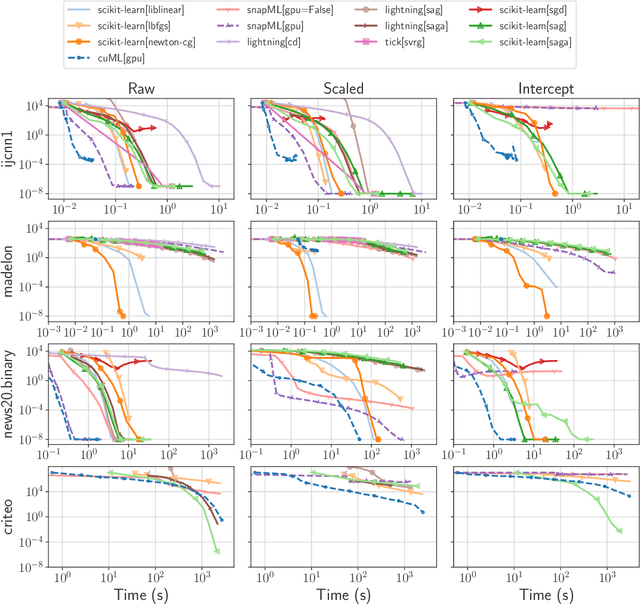
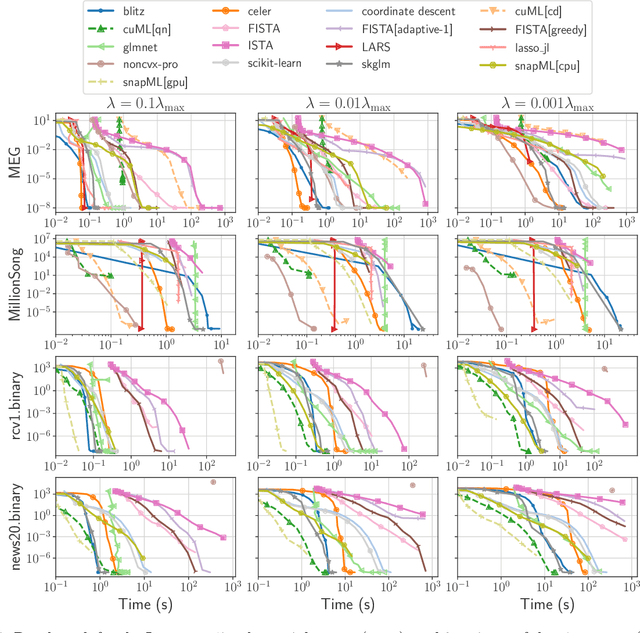
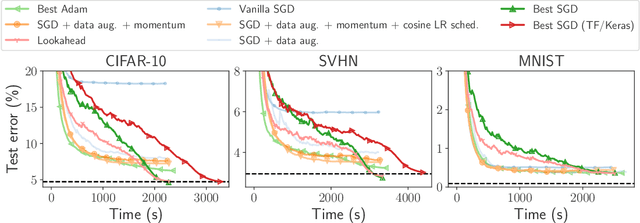
Abstract:Numerical validation is at the core of machine learning research as it allows to assess the actual impact of new methods, and to confirm the agreement between theory and practice. Yet, the rapid development of the field poses several challenges: researchers are confronted with a profusion of methods to compare, limited transparency and consensus on best practices, as well as tedious re-implementation work. As a result, validation is often very partial, which can lead to wrong conclusions that slow down the progress of research. We propose Benchopt, a collaborative framework to automate, reproduce and publish optimization benchmarks in machine learning across programming languages and hardware architectures. Benchopt simplifies benchmarking for the community by providing an off-the-shelf tool for running, sharing and extending experiments. To demonstrate its broad usability, we showcase benchmarks on three standard learning tasks: $\ell_2$-regularized logistic regression, Lasso, and ResNet18 training for image classification. These benchmarks highlight key practical findings that give a more nuanced view of the state-of-the-art for these problems, showing that for practical evaluation, the devil is in the details. We hope that Benchopt will foster collaborative work in the community hence improving the reproducibility of research findings.
Toward a realistic model of speech processing in the brain with self-supervised learning
Jun 03, 2022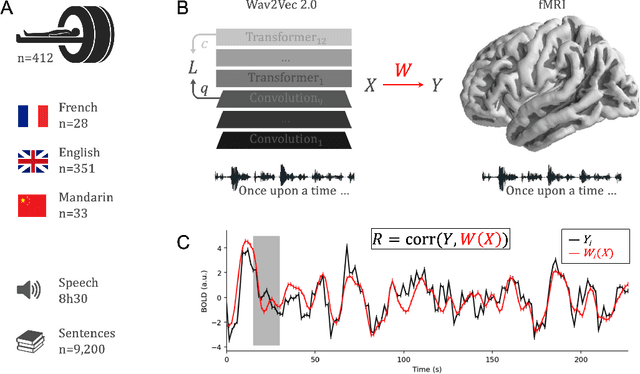
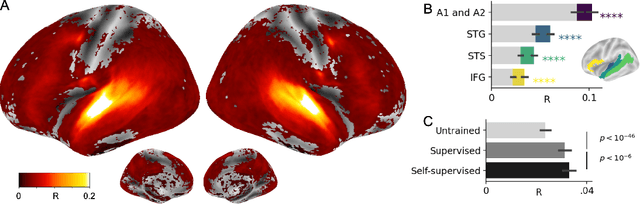
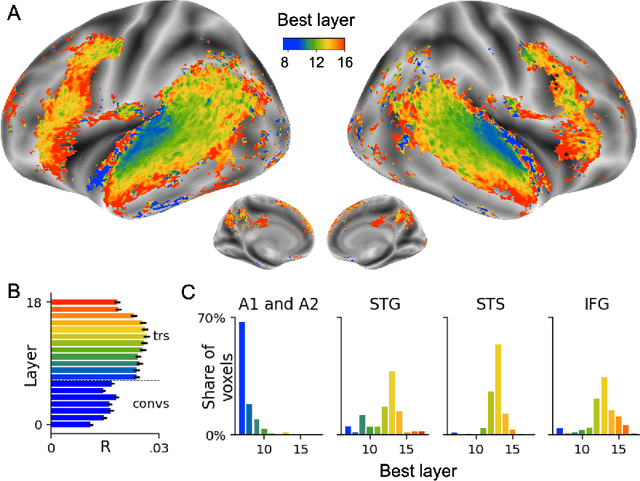
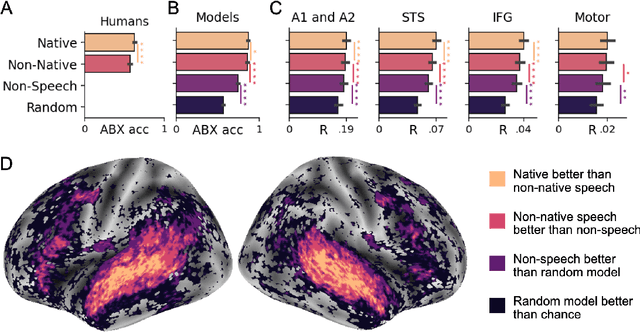
Abstract:Several deep neural networks have recently been shown to generate activations similar to those of the brain in response to the same input. These algorithms, however, remain largely implausible: they require (1) extraordinarily large amounts of data, (2) unobtainable supervised labels, (3) textual rather than raw sensory input, and / or (4) implausibly large memory (e.g. thousands of contextual words). These elements highlight the need to identify algorithms that, under these limitations, would suffice to account for both behavioral and brain responses. Focusing on the issue of speech processing, we here hypothesize that self-supervised algorithms trained on the raw waveform constitute a promising candidate. Specifically, we compare a recent self-supervised architecture, Wav2Vec 2.0, to the brain activity of 412 English, French, and Mandarin individuals recorded with functional Magnetic Resonance Imaging (fMRI), while they listened to ~1h of audio books. Our results are four-fold. First, we show that this algorithm learns brain-like representations with as little as 600 hours of unlabelled speech -- a quantity comparable to what infants can be exposed to during language acquisition. Second, its functional hierarchy aligns with the cortical hierarchy of speech processing. Third, different training regimes reveal a functional specialization akin to the cortex: Wav2Vec 2.0 learns sound-generic, speech-specific and language-specific representations similar to those of the prefrontal and temporal cortices. Fourth, we confirm the similarity of this specialization with the behavior of 386 additional participants. These elements, resulting from the largest neuroimaging benchmark to date, show how self-supervised learning can account for a rich organization of speech processing in the brain, and thus delineate a path to identify the laws of language acquisition which shape the human brain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge