"Image": models, code, and papers
An open-source software package for on-the-fly deskewing and live viewing of volumetric lightsheet microscopy data
Oct 31, 2022
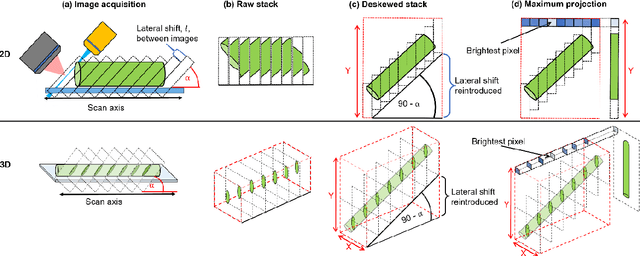
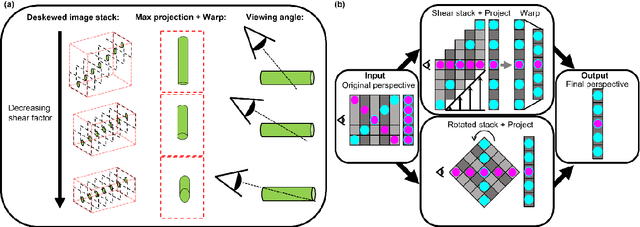
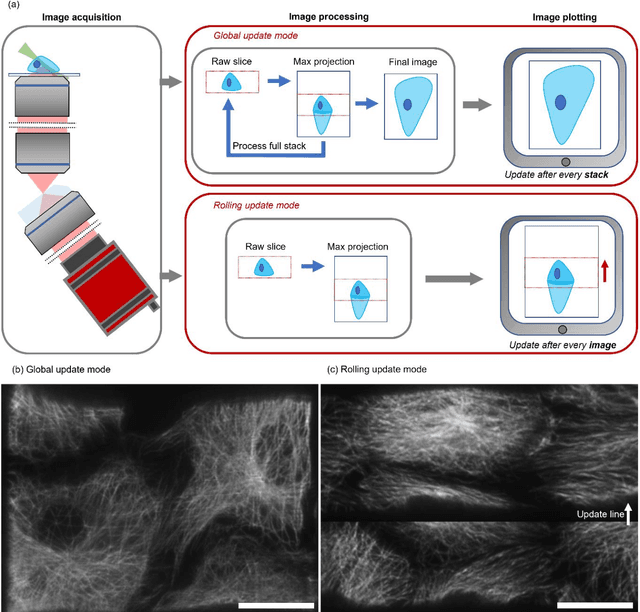
Oblique plane microscopy, OPM, is a form of lightsheet microscopy that permits volumetric imaging of biological samples at high temporal and spatial resolution. However, the imaging geometry of OPM, and related variants of light sheet microscopy, distorts the coordinate frame of the presented image sections with respect to real space coordinate frame in which the sample is moved to navigate to regions of interest. This makes live viewing and practical operation of such microscopes difficult. We present an open-source software package that utilises GPU acceleration and multiprocessing to transform the display of OPM imaging data in real time to produce live views that mimic that produced by standard widefield microscopes. Image stacks can be acquired, processed and plotted at rates of several Hz, making live operation of OPMs, and similar microscopes, more user friendly and intuitive.
A Feasibility Study on Image Inpainting for Non-cleft Lip Generation from Patients with Cleft Lip
Aug 01, 2022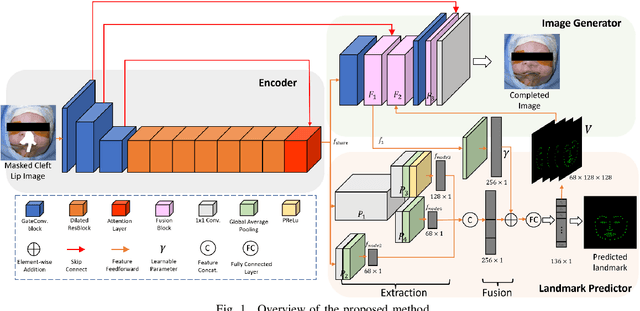
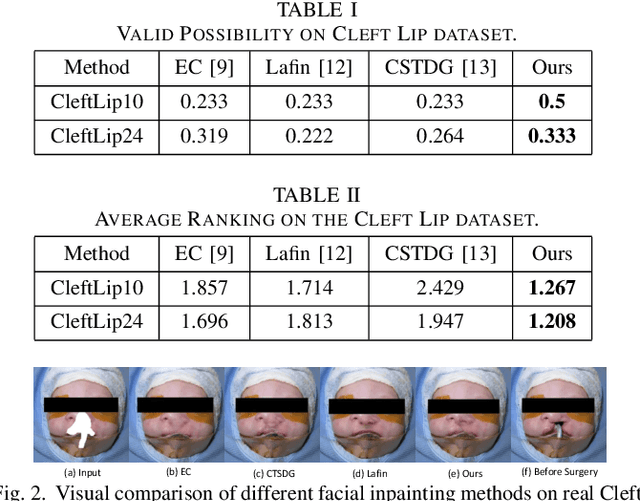
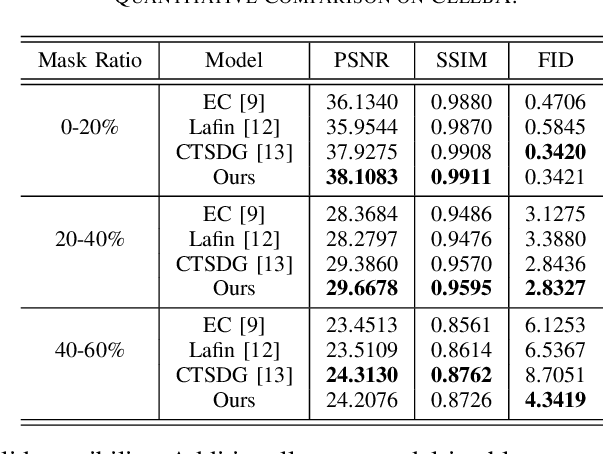
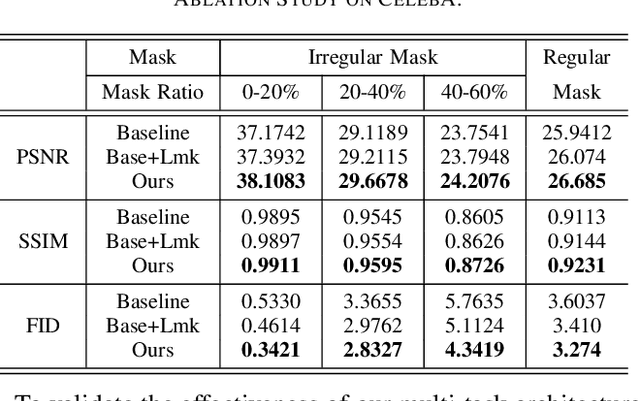
A Cleft lip is a congenital abnormality requiring surgical repair by a specialist. The surgeon must have extensive experience and theoretical knowledge to perform surgery, and Artificial Intelligence (AI) method has been proposed to guide surgeons in improving surgical outcomes. If AI can be used to predict what a repaired cleft lip would look like, surgeons could use it as an adjunct to adjust their surgical technique and improve results. To explore the feasibility of this idea while protecting patient privacy, we propose a deep learning-based image inpainting method that is capable of covering a cleft lip and generating a lip and nose without a cleft. Our experiments are conducted on two real-world cleft lip datasets and are assessed by expert cleft lip surgeons to demonstrate the feasibility of the proposed method.
Deep neural network techniques for monaural speech enhancement: state of the art analysis
Dec 01, 2022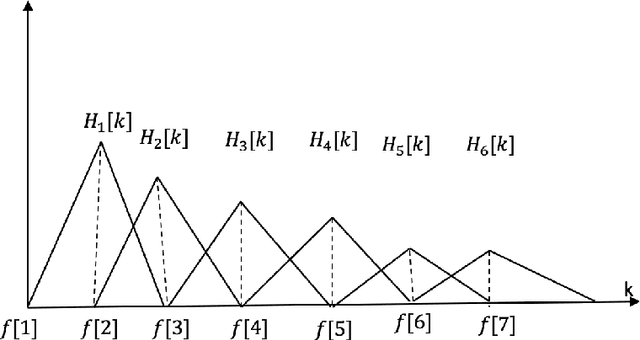
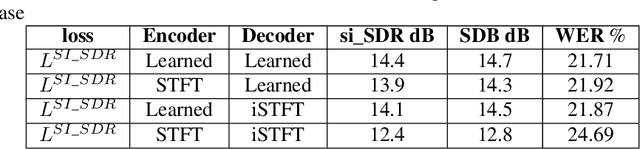
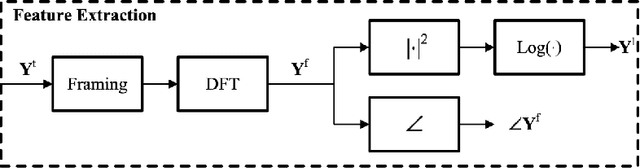

Deep neural networks (DNN) techniques have become pervasive in domains such as natural language processing and computer vision. They have achieved great success in these domains in task such as machine translation and image generation. Due to their success, these data driven techniques have been applied in audio domain. More specifically, DNN models have been applied in speech enhancement domain to achieve denosing, dereverberation and multi-speaker separation in monaural speech enhancement. In this paper, we review some dominant DNN techniques being employed to achieve speech separation. The review looks at the whole pipeline of speech enhancement from feature extraction, how DNN based tools are modelling both global and local features of speech and model training (supervised and unsupervised). We also review the use of speech-enhancement pre-trained models to boost speech enhancement process. The review is geared towards covering the dominant trends with regards to DNN application in speech enhancement in speech obtained via a single speaker.
Neural Representations Reveal Distinct Modes of Class Fitting in Residual Convolutional Networks
Dec 01, 2022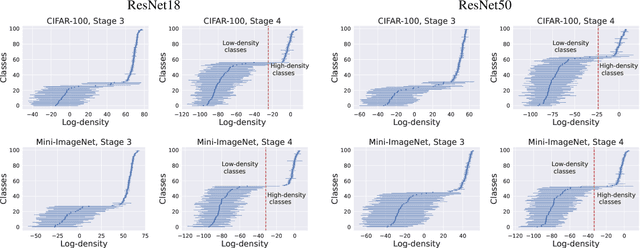
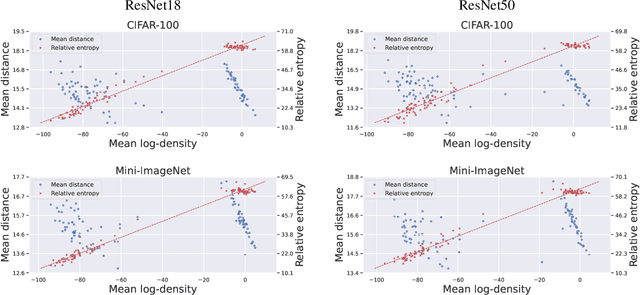
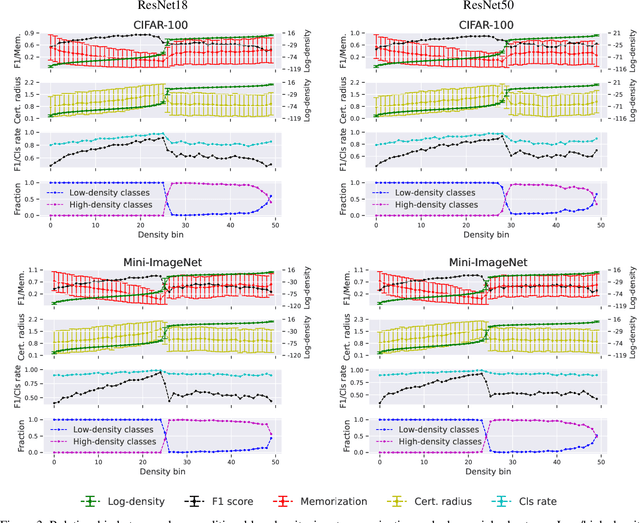
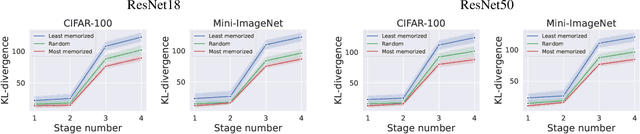
We leverage probabilistic models of neural representations to investigate how residual networks fit classes. To this end, we estimate class-conditional density models for representations learned by deep ResNets. We then use these models to characterize distributions of representations across learned classes. Surprisingly, we find that classes in the investigated models are not fitted in an uniform way. On the contrary: we uncover two groups of classes that are fitted with markedly different distributions of representations. These distinct modes of class-fitting are evident only in the deeper layers of the investigated models, indicating that they are not related to low-level image features. We show that the uncovered structure in neural representations correlate with memorization of training examples and adversarial robustness. Finally, we compare class-conditional distributions of neural representations between memorized and typical examples. This allows us to uncover where in the network structure class labels arise for memorized and standard inputs.
Low-complexity Three-dimensional Discrete Hartley Transform Approximations for Medical Image Compression
May 31, 2022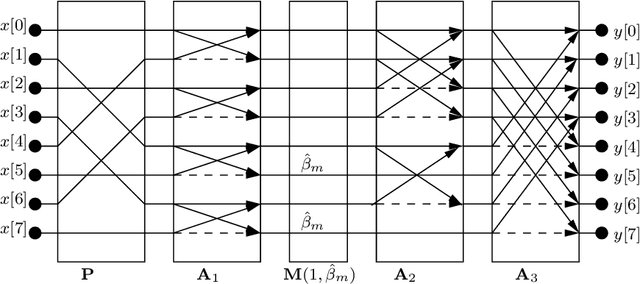
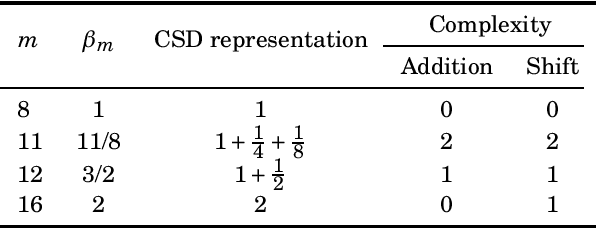
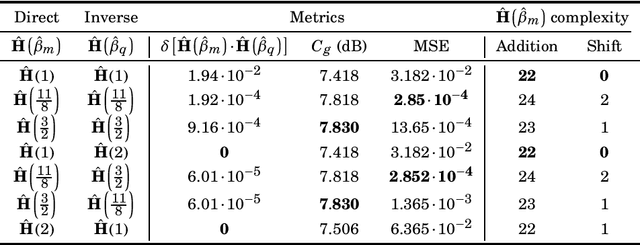
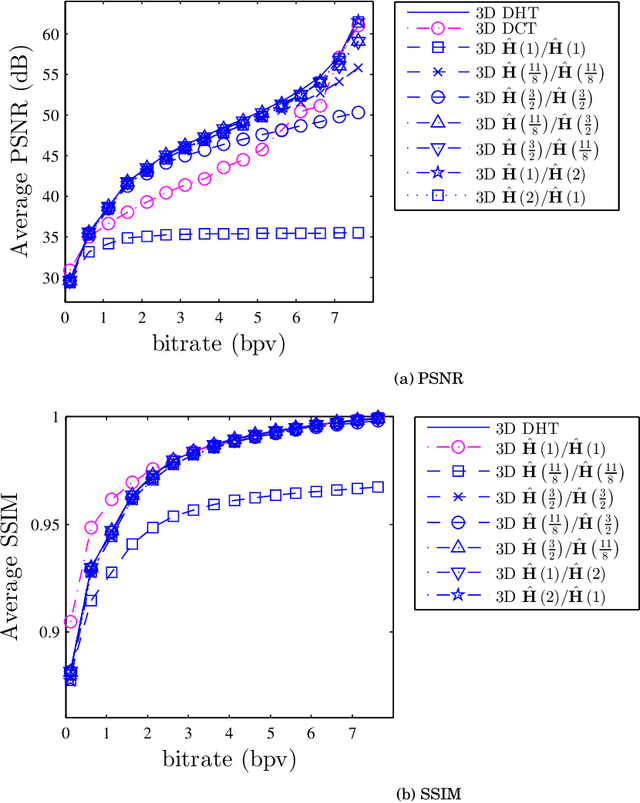
The discrete Hartley transform (DHT) is a useful tool for medical image coding. The three-dimensional DHT (3D DHT) can be employed to compress medical image data, such as magnetic resonance and X-ray angiography. However, the computation of the 3D DHT involves several multiplications by irrational quantities, which require floating-point arithmetic and inherent truncation errors. In recent years, a significant progress in wireless and implantable biomedical devices has been achieved. Such devices present critical power and hardware limitations. The multiplication operation demands higher hardware, power, and time consumption than other arithmetic operations, such as addition and bit-shifts. In this work, we present a set of multiplierless DHT approximations, which can be implemented with fixed-point arithmetic. We derive 3D DHT approximations by employing tensor formalism. Such proposed methods present prominent computational savings compared to the usual 3D DHT approach, being appropriate for devices with limited resources. The proposed transforms are applied in a lossy 3D DHT-based medical image compression algorithm, presenting practically the same level of visual quality ($>98\%$ in terms of SSIM) at a considerable reduction in computational effort ($100 \%$ multiplicative complexity reduction). Furthermore, we implemented the proposed 3D transforms in an ARM Cortex-M0+ processor employing the low-cost Raspberry Pi Pico board. The execution time was reduced by $\sim$70% compared to the usual 3D DHT and $\sim$90% compared to 3D DCT.
* 22 pages, 7 tables, 6 figures
Analysis of convolutional neural network image classifiers in a rotationally symmetric model
May 11, 2022
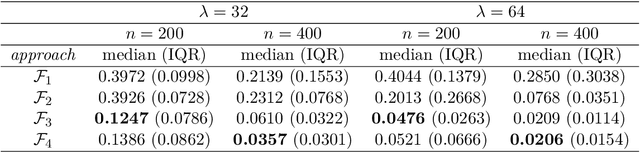
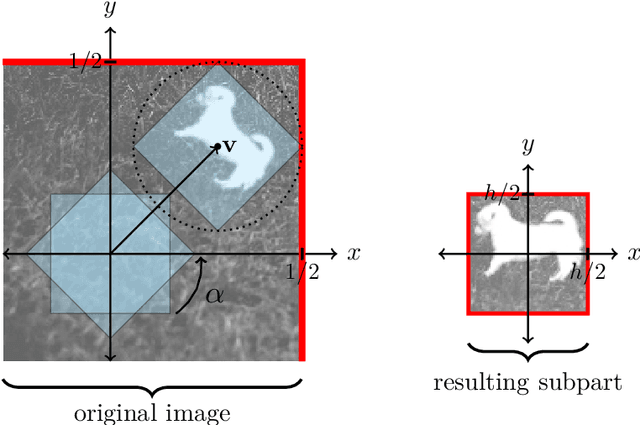
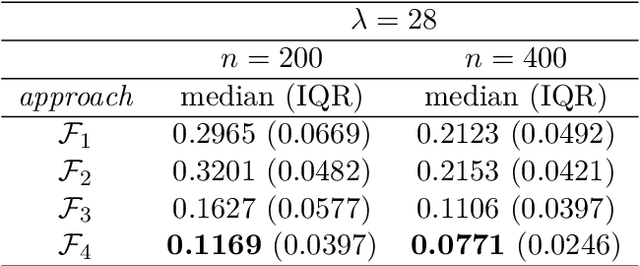
Convolutional neural network image classifiers are defined and the rate of convergence of the misclassification risk of the estimates towards the optimal misclassification risk is analyzed. Here we consider images as random variables with values in some functional space, where we only observe discrete samples as function values on some finite grid. Under suitable structural and smoothness assumptions on the functional a posteriori probability, which includes some kind of symmetry against rotation of subparts of the input image, it is shown that least squares plug-in classifiers based on convolutional neural networks are able to circumvent the curse of dimensionality in binary image classification if we neglect a resolution-dependent error term. The finite sample size behavior of the classifier is analyzed by applying it to simulated and real data.
The Cross Density Kernel Function: A Novel Framework to Quantify Statistical Dependence for Random Processes
Dec 09, 2022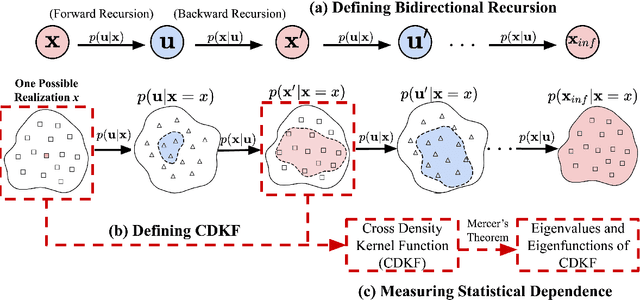
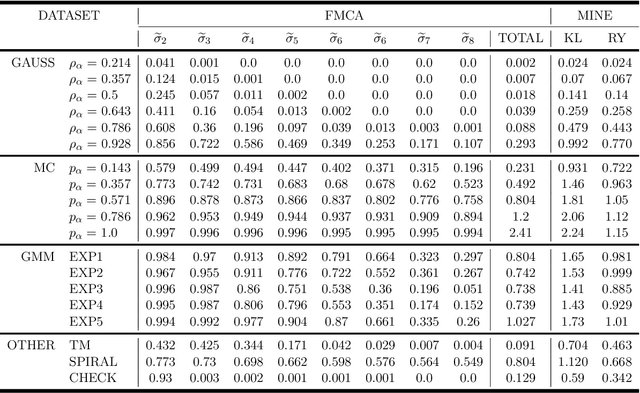
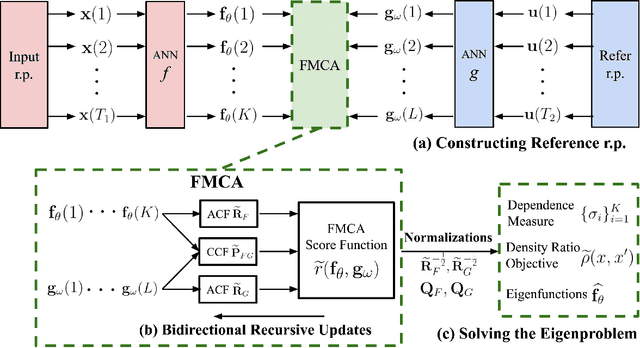
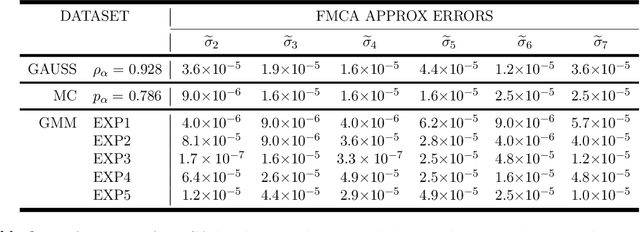
This paper proposes a novel multivariate definition of statistical dependence using a functional methodology inspired by Alfred R\'enyi. We define a new symmetric and self-adjoint cross density kernel through a recursive bidirectional statistical mapping between conditional densities of continuous random processes, which estimates their statistical dependence. Therefore, the kernel eigenspectrum is proposed as a new multivariate statistical dependence measure, and the formulation requires fewer assumptions about the data generation model than current methods. The measure can also be estimated from realizations. The proposed functional maximum correlation algorithm (FMCA) is applied to a learning architecture with two multivariate neural networks. The FMCA optimal solution is an equilibrium point that estimates the eigenspectrum of the cross density kernel. Preliminary results with synthetic data and medium size image datasets corroborate the theory. Four different strategies of applying the cross density kernel are thoroughly discussed and implemented to show the versatility and stability of the methodology, and it transcends supervised learning. When two random processes are high-dimensional real-world images and white uniform noise, respectively, the algorithm learns a factorial code i.e., the occurrence of a code guarantees that a certain input in the training set was present, which is quite important for feature learning.
Frugal Reinforcement-based Active Learning
Dec 09, 2022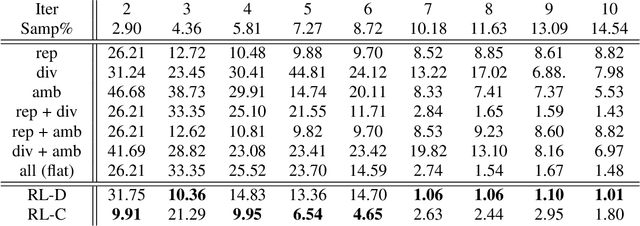
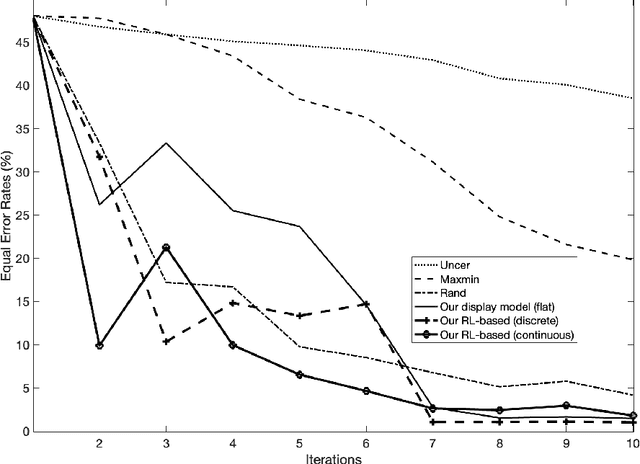
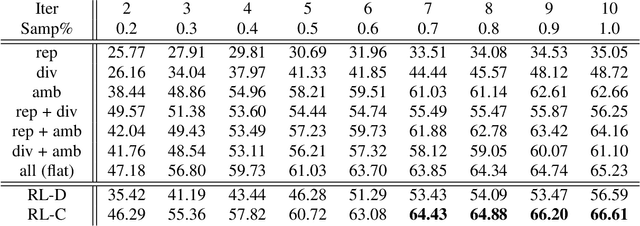
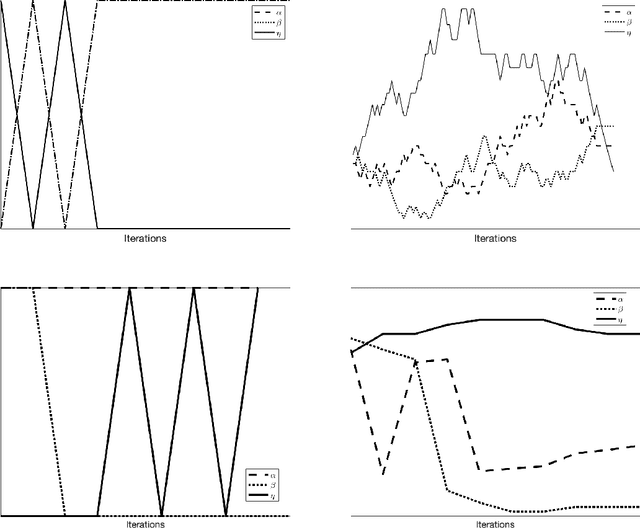
Most of the existing learning models, particularly deep neural networks, are reliant on large datasets whose hand-labeling is expensive and time demanding. A current trend is to make the learning of these models frugal and less dependent on large collections of labeled data. Among the existing solutions, deep active learning is currently witnessing a major interest and its purpose is to train deep networks using as few labeled samples as possible. However, the success of active learning is highly dependent on how critical are these samples when training models. In this paper, we devise a novel active learning approach for label-efficient training. The proposed method is iterative and aims at minimizing a constrained objective function that mixes diversity, representativity and uncertainty criteria. The proposed approach is probabilistic and unifies all these criteria in a single objective function whose solution models the probability of relevance of samples (i.e., how critical) when learning a decision function. We also introduce a novel weighting mechanism based on reinforcement learning, which adaptively balances these criteria at each training iteration, using a particular stateless Q-learning model. Extensive experiments conducted on staple image classification data, including Object-DOTA, show the effectiveness of our proposed model w.r.t. several baselines including random, uncertainty and flat as well as other work.
Dual adaptive training of photonic neural networks
Dec 09, 2022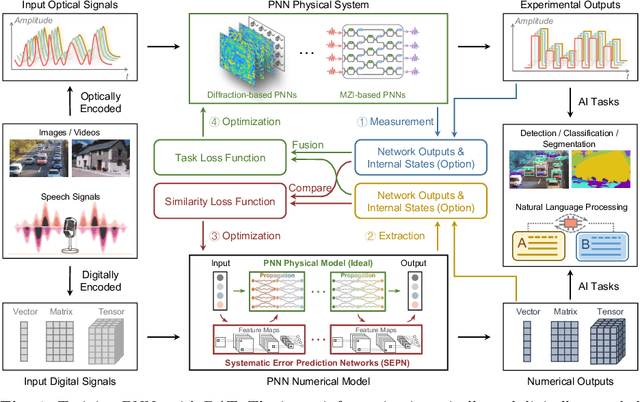
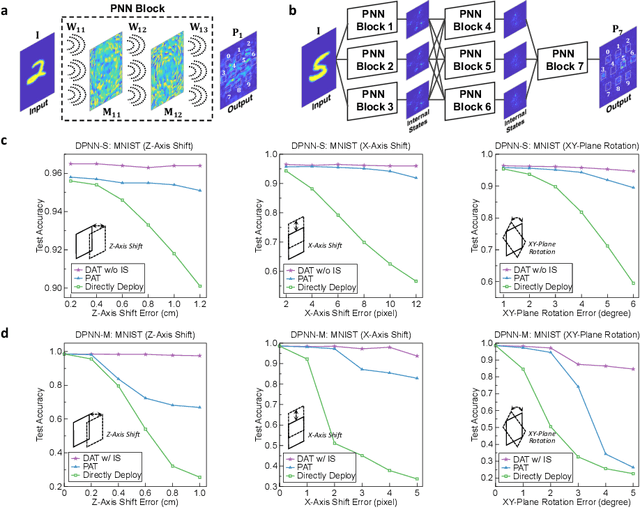
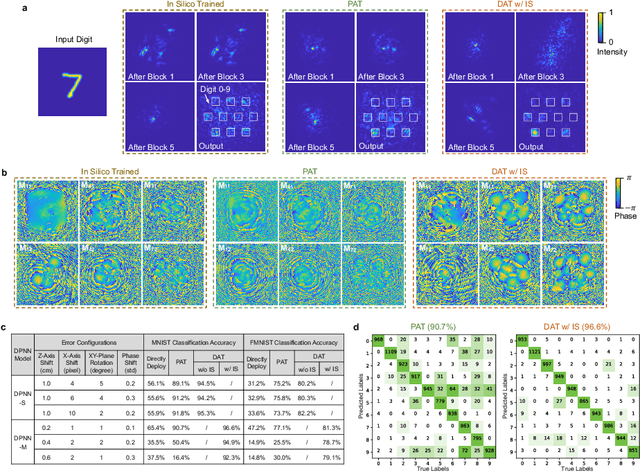
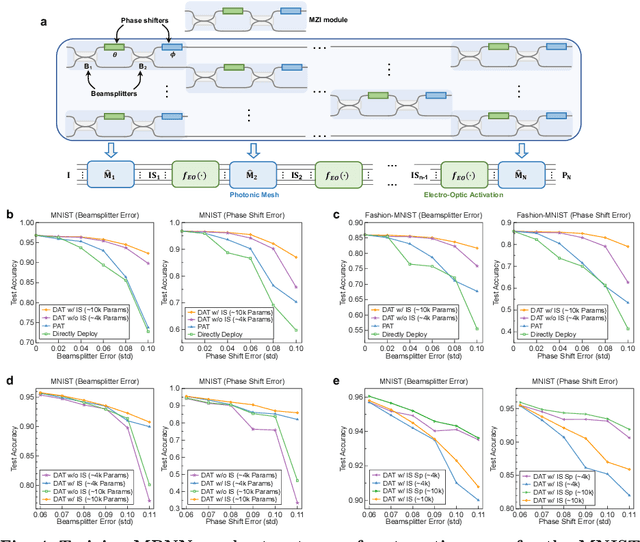
Photonic neural network (PNN) is a remarkable analog artificial intelligence (AI) accelerator that computes with photons instead of electrons to feature low latency, high energy efficiency, and high parallelism. However, the existing training approaches cannot address the extensive accumulation of systematic errors in large-scale PNNs, resulting in a significant decrease in model performance in physical systems. Here, we propose dual adaptive training (DAT) that allows the PNN model to adapt to substantial systematic errors and preserves its performance during the deployment. By introducing the systematic error prediction networks with task-similarity joint optimization, DAT achieves the high similarity mapping between the PNN numerical models and physical systems and high-accurate gradient calculations during the dual backpropagation training. We validated the effectiveness of DAT by using diffractive PNNs and interference-based PNNs on image classification tasks. DAT successfully trained large-scale PNNs under major systematic errors and preserved the model classification accuracies comparable to error-free systems. The results further demonstrated its superior performance over the state-of-the-art in situ training approaches. DAT provides critical support for constructing large-scale PNNs to achieve advanced architectures and can be generalized to other types of AI systems with analog computing errors.
Rethinking Surgical Instrument Segmentation: A Background Image Can Be All You Need
Jun 30, 2022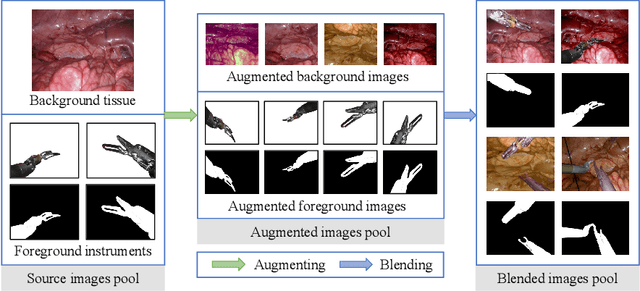
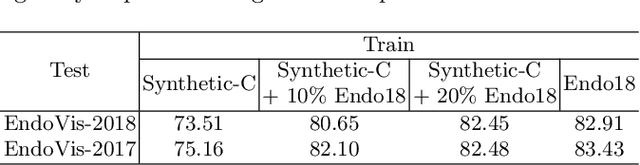
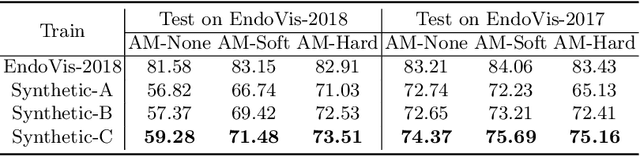


Data diversity and volume are crucial to the success of training deep learning models, while in the medical imaging field, the difficulty and cost of data collection and annotation are especially huge. Specifically in robotic surgery, data scarcity and imbalance have heavily affected the model accuracy and limited the design and deployment of deep learning-based surgical applications such as surgical instrument segmentation. Considering this, we rethink the surgical instrument segmentation task and propose a one-to-many data generation solution that gets rid of the complicated and expensive process of data collection and annotation from robotic surgery. In our method, we only utilize a single surgical background tissue image and a few open-source instrument images as the seed images and apply multiple augmentations and blending techniques to synthesize amounts of image variations. In addition, we also introduce the chained augmentation mixing during training to further enhance the data diversities. The proposed approach is evaluated on the real datasets of the EndoVis-2018 and EndoVis-2017 surgical scene segmentation. Our empirical analysis suggests that without the high cost of data collection and annotation, we can achieve decent surgical instrument segmentation performance. Moreover, we also observe that our method can deal with novel instrument prediction in the deployment domain. We hope our inspiring results will encourage researchers to emphasize data-centric methods to overcome demanding deep learning limitations besides data shortage, such as class imbalance, domain adaptation, and incremental learning. Our code is available at https://github.com/lofrienger/Single_SurgicalScene_For_Segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge