Zudi Lin
AxonEM Dataset: 3D Axon Instance Segmentation of Brain Cortical Regions
Jul 12, 2021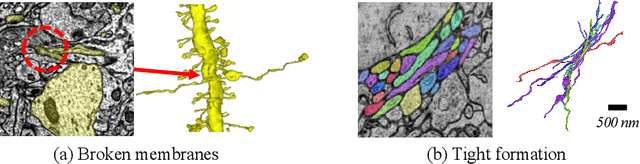
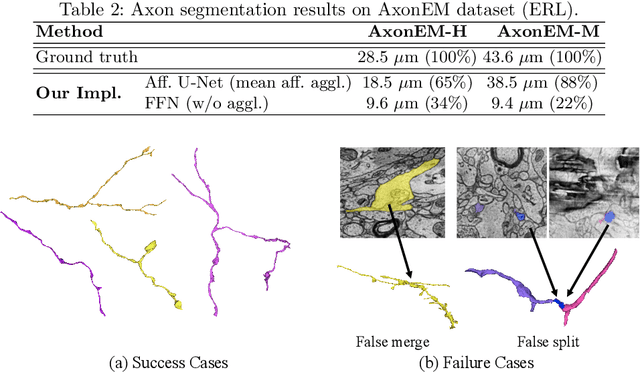

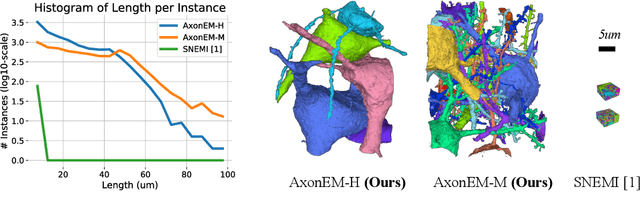

Abstract:Electron microscopy (EM) enables the reconstruction of neural circuits at the level of individual synapses, which has been transformative for scientific discoveries. However, due to the complex morphology, an accurate reconstruction of cortical axons has become a major challenge. Worse still, there is no publicly available large-scale EM dataset from the cortex that provides dense ground truth segmentation for axons, making it difficult to develop and evaluate large-scale axon reconstruction methods. To address this, we introduce the AxonEM dataset, which consists of two 30x30x30 um^3 EM image volumes from the human and mouse cortex, respectively. We thoroughly proofread over 18,000 axon instances to provide dense 3D axon instance segmentation, enabling large-scale evaluation of axon reconstruction methods. In addition, we densely annotate nine ground truth subvolumes for training, per each data volume. With this, we reproduce two published state-of-the-art methods and provide their evaluation results as a baseline. We publicly release our code and data at https://connectomics-bazaar.github.io/proj/AxonEM/index.html to foster the development of advanced methods.
A Topological Nomenclature for 3D Shape Analysis in Connectomics
Sep 27, 2019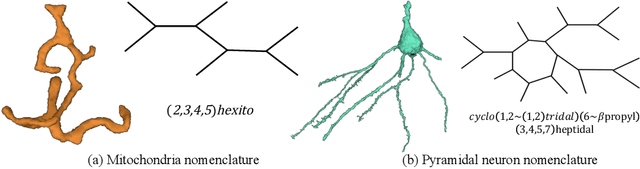
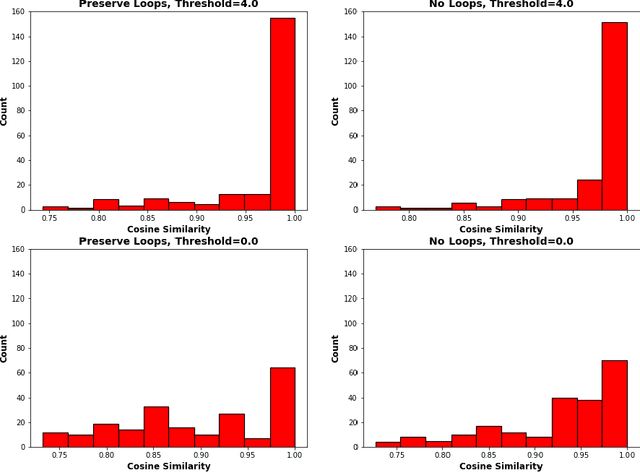
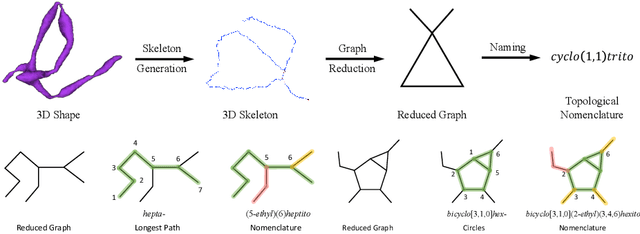
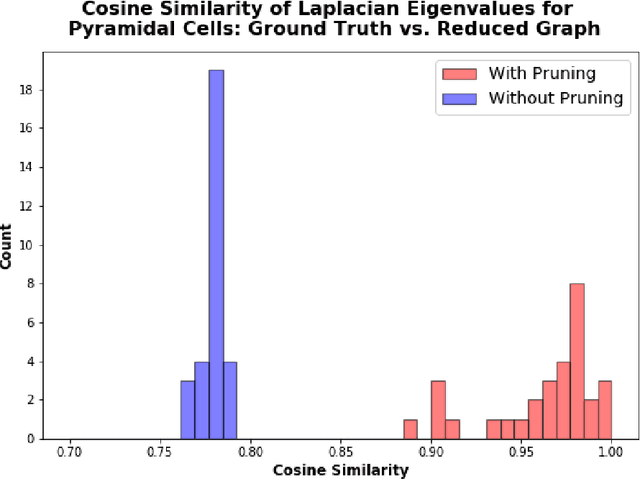
Abstract:An essential task in nano-scale connectomics is the morphology analysis of neurons and organelles like mitochondria to shed light on their biological properties. However, these biological objects often have tangled parts or complex branching patterns, which makes it hard to abstract, categorize, and manipulate their morphology. Here we propose a topological nomenclature to name these objects like chemical compounds for neuroscience analysis. To this end, we convert the volumetric representation into the topology-preserving reduced graph, develop nomenclature rules for pyramidal neurons and mitochondria from the reduced graph, and learn the feature embedding for shape manipulation. In ablation studies, we show that the proposed reduced graph extraction method yield graphs better in accord with the perception of experts. On 3D shape retrieval and decomposition tasks, we show that the encoded topological nomenclature features achieve better results than state-of-the-art shape descriptors. To advance neuroscience, we will release a 3D mesh dataset of mitochondria and pyramidal neurons reconstructed from a 100{\mu}m cube electron microscopy (EM) volume. Code is publicly available at https://github.com/donglaiw/ibexHelper.
White-Box Adversarial Defense via Self-Supervised Data Estimation
Sep 13, 2019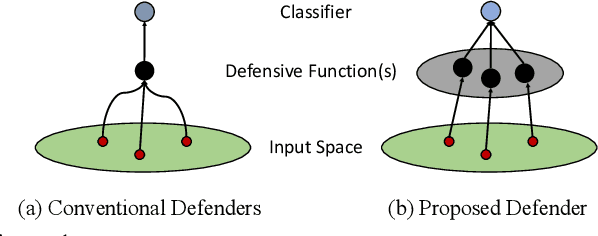
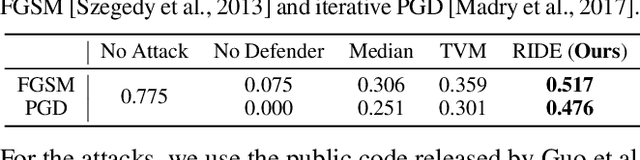
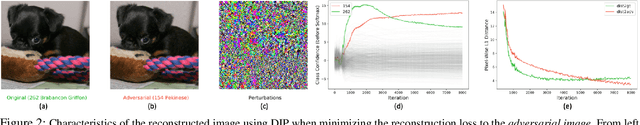
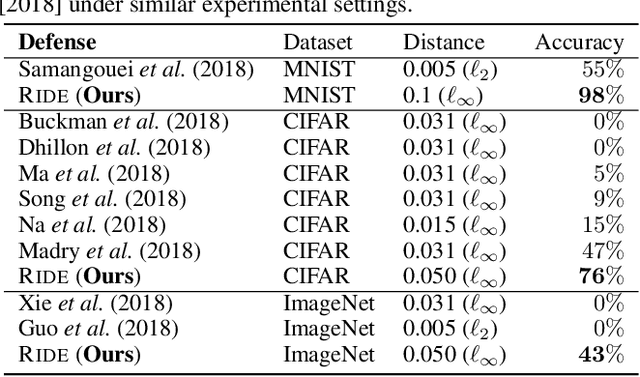
Abstract:In this paper, we study the problem of how to defend classifiers against adversarial attacks that fool the classifiers using subtly modified input data. In contrast to previous works, here we focus on the white-box adversarial defense where the attackers are granted full access to not only the classifiers but also defenders to produce as strong attacks as possible. In such a context we propose viewing a defender as a functional, a higher-order function that takes functions as its argument to represent a function space, rather than fixed functions conventionally. From this perspective, a defender should be realized and optimized individually for each adversarial input. To this end, we propose RIDE, an efficient and provably convergent self-supervised learning algorithm for individual data estimation to protect the predictions from adversarial attacks. We demonstrate the significant improvement of adversarial defense performance on image recognition, eg, 98%, 76%, 43% test accuracy on MNIST, CIFAR-10, and ImageNet datasets respectively under the state-of-the-art BPDA attacker.
FDive: Learning Relevance Models using Pattern-based Similarity Measures
Jul 30, 2019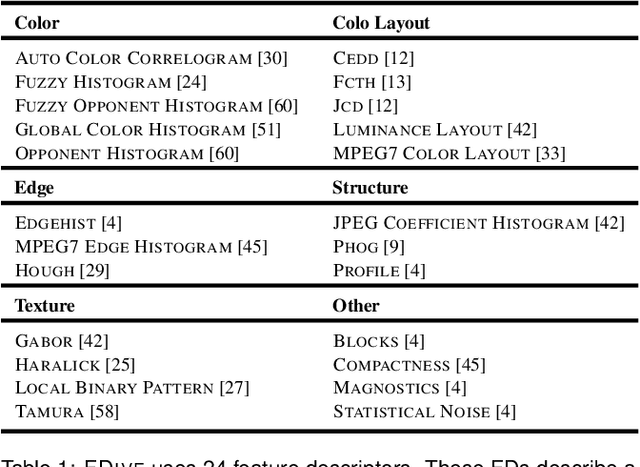
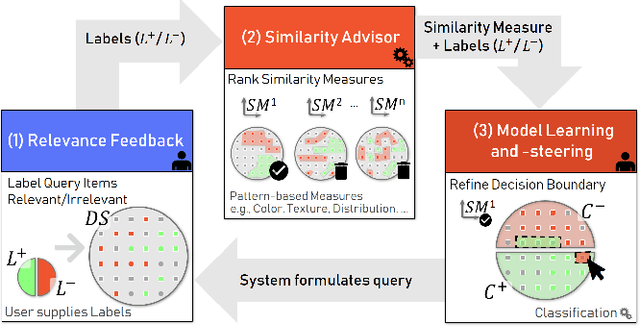
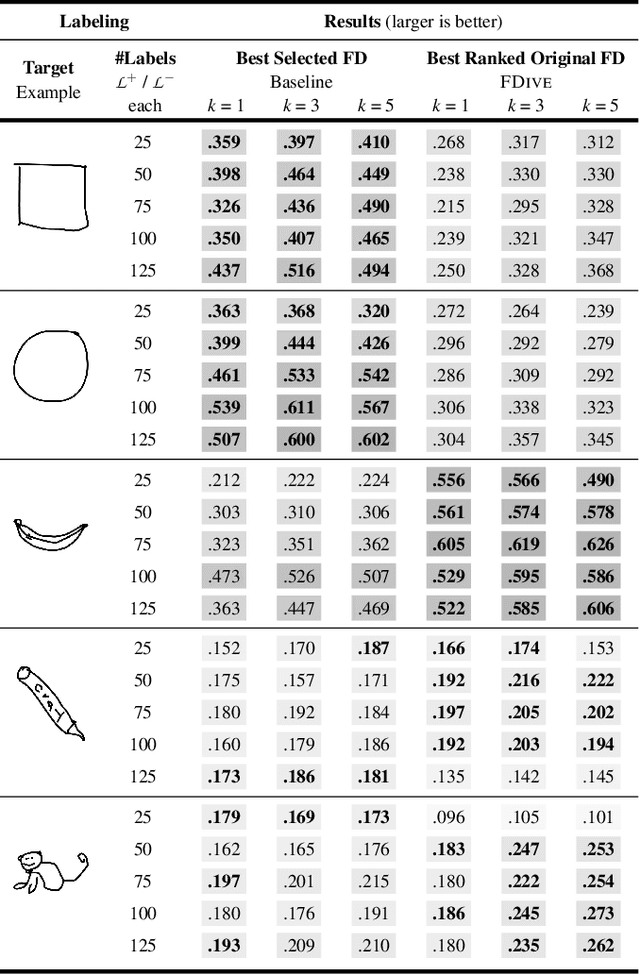
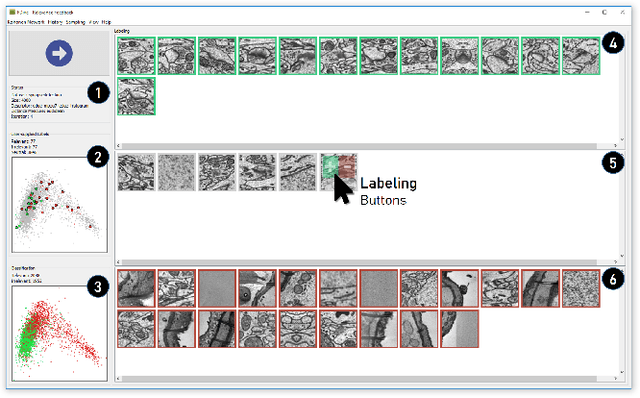
Abstract:The detection of interesting patterns in large high-dimensional datasets is difficult because of their dimensionality and pattern complexity. Therefore, analysts require automated support for the extraction of relevant patterns. In this paper, we present FDive, a visual active learning system that helps to create visually explorable relevance models, assisted by learning a pattern-based similarity. We use a small set of user-provided labels to rank similarity measures, consisting of feature descriptor and distance function combinations, by their ability to distinguish relevant from irrelevant data. Based on the best-ranked similarity measure, the system calculates an interactive Self-Organizing Map-based relevance model, which classifies data according to the cluster affiliation. It also automatically prompts further relevance feedback to improve its accuracy. Uncertain areas, especially near the decision boundaries, are highlighted and can be refined by the user. We evaluate our approach by comparison to state-of-the-art feature selection techniques and demonstrate the usefulness of our approach by a case study classifying electron microscopy images of brain cells. The results show that FDive enhances both the quality and understanding of relevance models and can thus lead to new insights for brain research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge