Zoubin Ghahramani
Dima
Bayesian Structured Prediction Using Gaussian Processes
Jul 15, 2013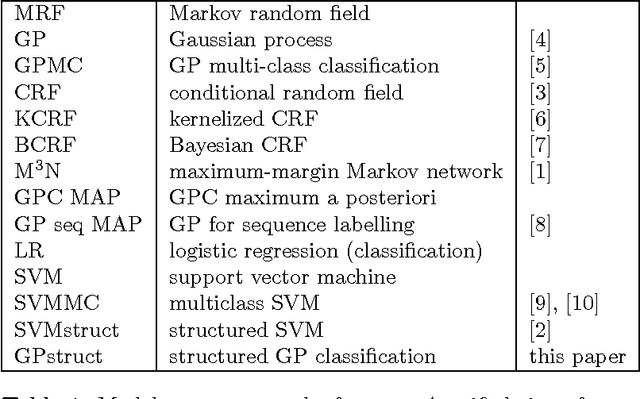
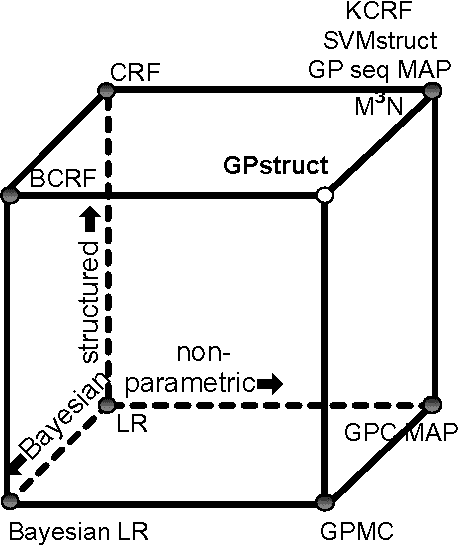
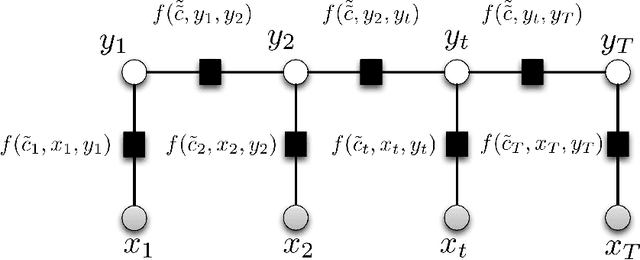
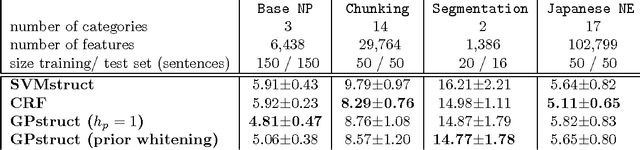
Abstract:We introduce a conceptually novel structured prediction model, GPstruct, which is kernelized, non-parametric and Bayesian, by design. We motivate the model with respect to existing approaches, among others, conditional random fields (CRFs), maximum margin Markov networks (M3N), and structured support vector machines (SVMstruct), which embody only a subset of its properties. We present an inference procedure based on Markov Chain Monte Carlo. The framework can be instantiated for a wide range of structured objects such as linear chains, trees, grids, and other general graphs. As a proof of concept, the model is benchmarked on several natural language processing tasks and a video gesture segmentation task involving a linear chain structure. We show prediction accuracies for GPstruct which are comparable to or exceeding those of CRFs and SVMstruct.
Dynamic Covariance Models for Multivariate Financial Time Series
Jun 02, 2013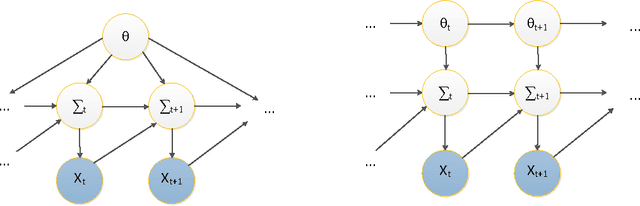
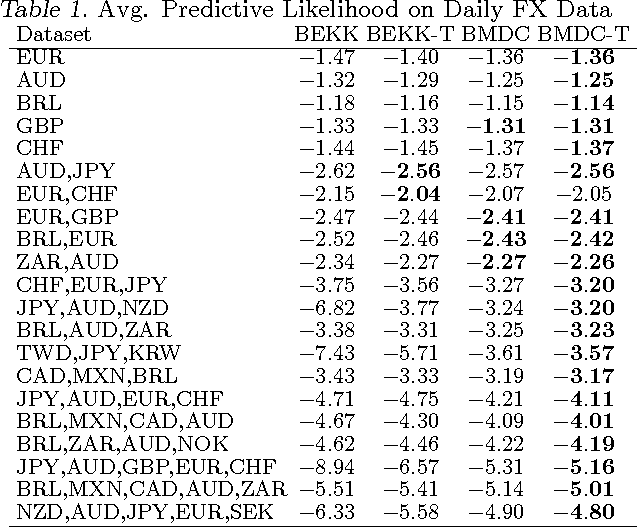
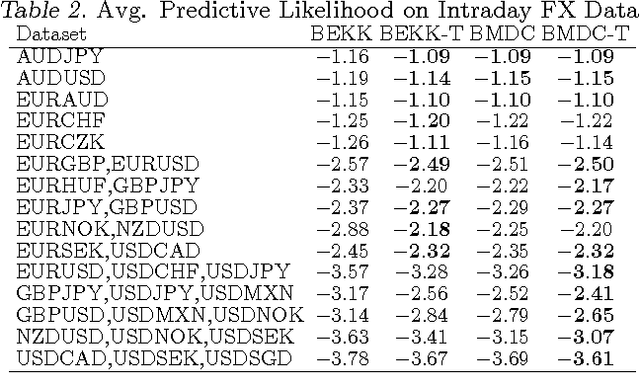
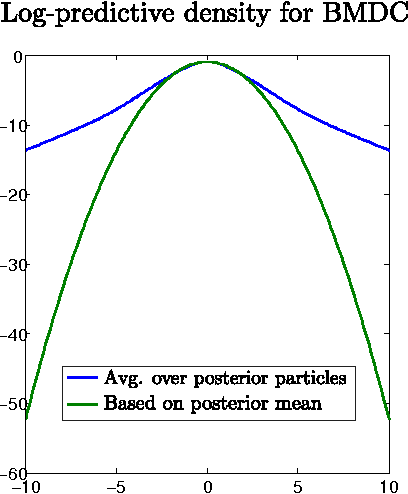
Abstract:The accurate prediction of time-changing covariances is an important problem in the modeling of multivariate financial data. However, some of the most popular models suffer from a) overfitting problems and multiple local optima, b) failure to capture shifts in market conditions and c) large computational costs. To address these problems we introduce a novel dynamic model for time-changing covariances. Over-fitting and local optima are avoided by following a Bayesian approach instead of computing point estimates. Changes in market conditions are captured by assuming a diffusion process in parameter values, and finally computationally efficient and scalable inference is performed using particle filters. Experiments with financial data show excellent performance of the proposed method with respect to current standard models.
Structure Discovery in Nonparametric Regression through Compositional Kernel Search
May 13, 2013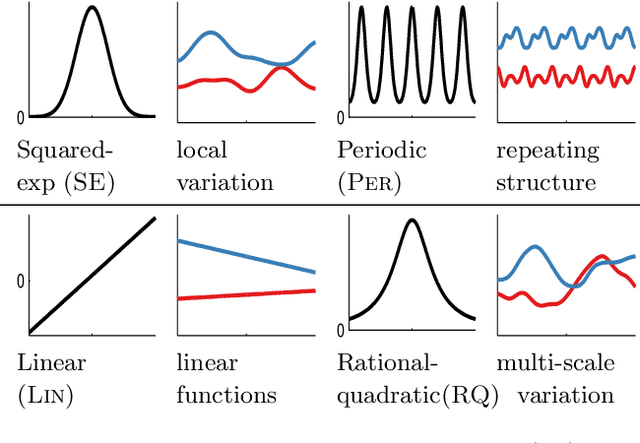
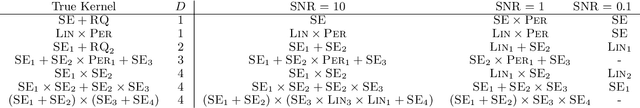

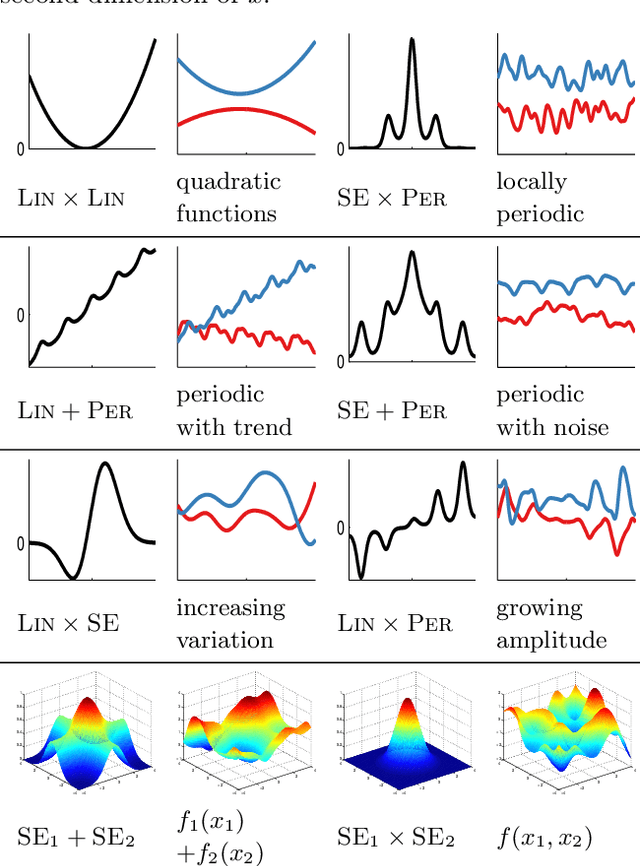
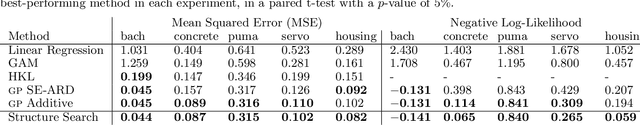
Abstract:Despite its importance, choosing the structural form of the kernel in nonparametric regression remains a black art. We define a space of kernel structures which are built compositionally by adding and multiplying a small number of base kernels. We present a method for searching over this space of structures which mirrors the scientific discovery process. The learned structures can often decompose functions into interpretable components and enable long-range extrapolation on time-series datasets. Our structure search method outperforms many widely used kernels and kernel combination methods on a variety of prediction tasks.
Identifying cancer subtypes in glioblastoma by combining genomic, transcriptomic and epigenomic data
Apr 15, 2013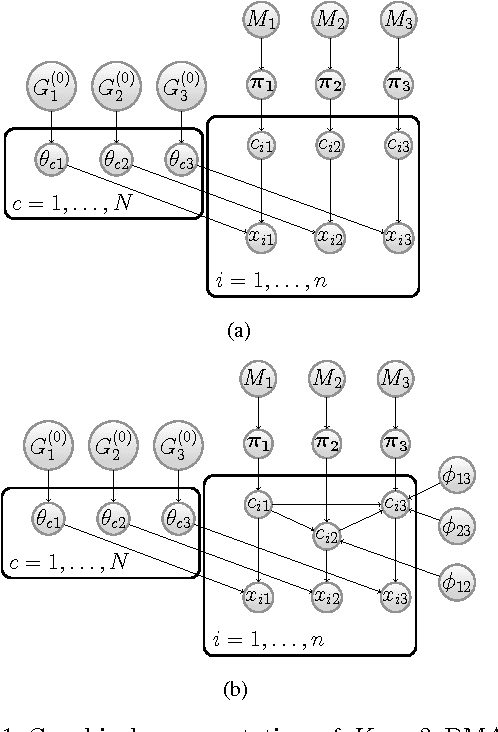
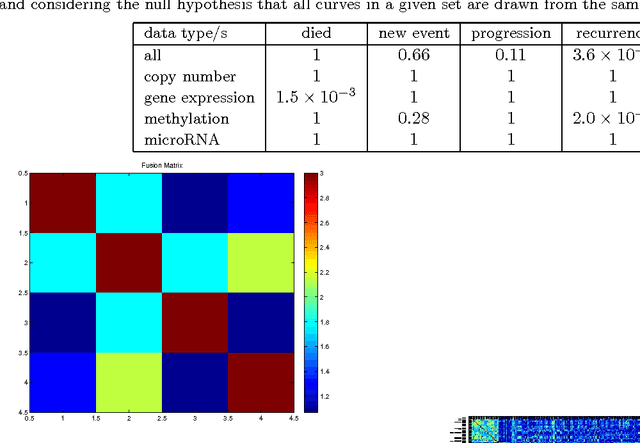
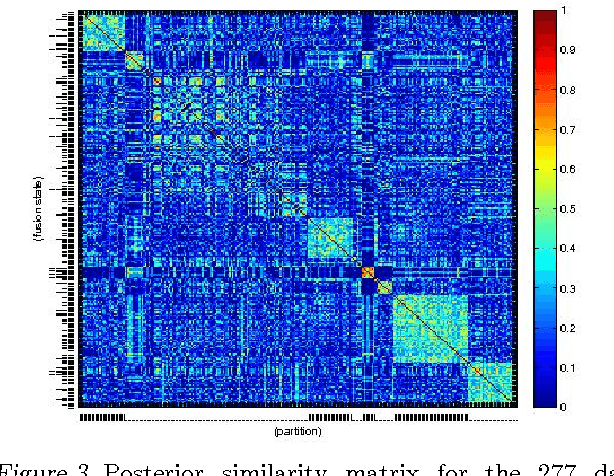
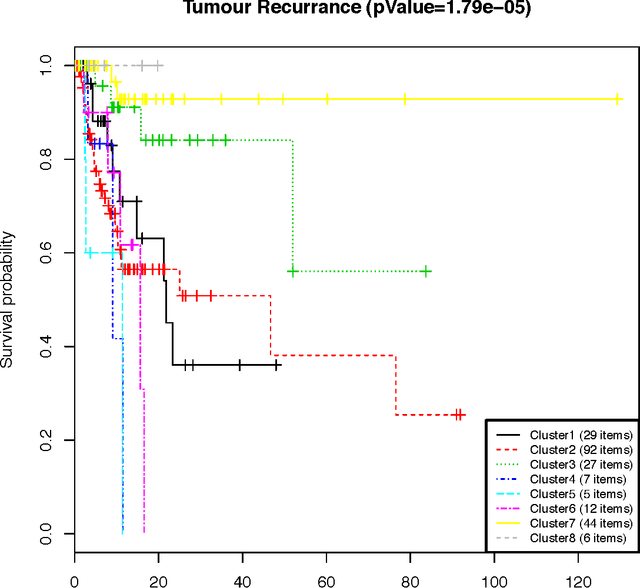
Abstract:We present a nonparametric Bayesian method for disease subtype discovery in multi-dimensional cancer data. Our method can simultaneously analyse a wide range of data types, allowing for both agreement and disagreement between their underlying clustering structure. It includes feature selection and infers the most likely number of disease subtypes, given the data. We apply the method to 277 glioblastoma samples from The Cancer Genome Atlas, for which there are gene expression, copy number variation, methylation and microRNA data. We identify 8 distinct consensus subtypes and study their prognostic value for death, new tumour events, progression and recurrence. The consensus subtypes are prognostic of tumour recurrence (log-rank p-value of $3.6 \times 10^{-4}$ after correction for multiple hypothesis tests). This is driven principally by the methylation data (log-rank p-value of $2.0 \times 10^{-3}$) but the effect is strengthened by the other 3 data types, demonstrating the value of integrating multiple data types. Of particular note is a subtype of 47 patients characterised by very low levels of methylation. This subtype has very low rates of tumour recurrence and no new events in 10 years of follow up. We also identify a small gene expression subtype of 6 patients that shows particularly poor survival outcomes. Additionally, we note a consensus subtype that showly a highly distinctive data signature and suggest that it is therefore a biologically distinct subtype of glioblastoma. The code is available from https://sites.google.com/site/multipledatafusion/
Gaussian Process Vine Copulas for Multivariate Dependence
Feb 16, 2013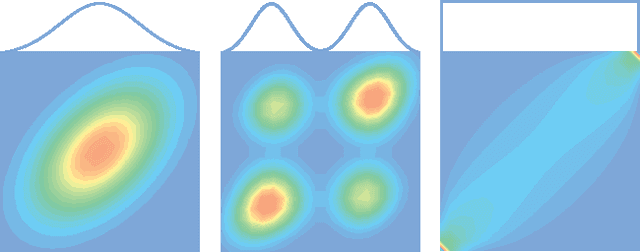
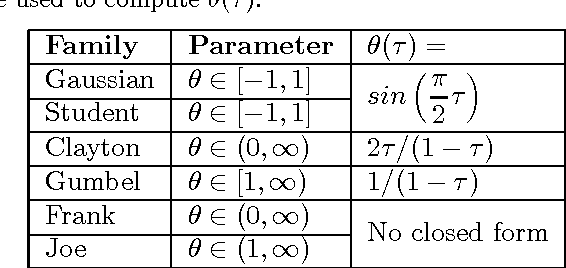
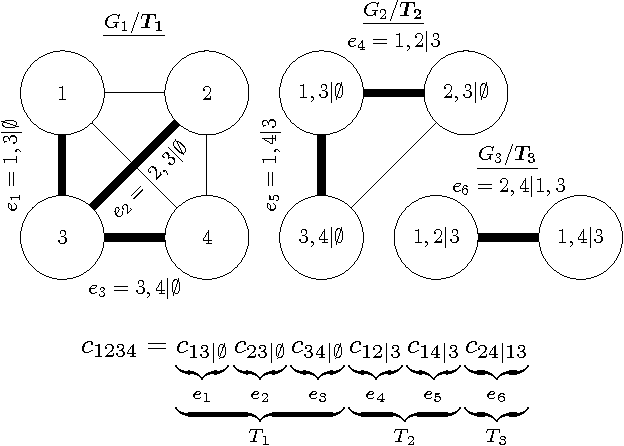
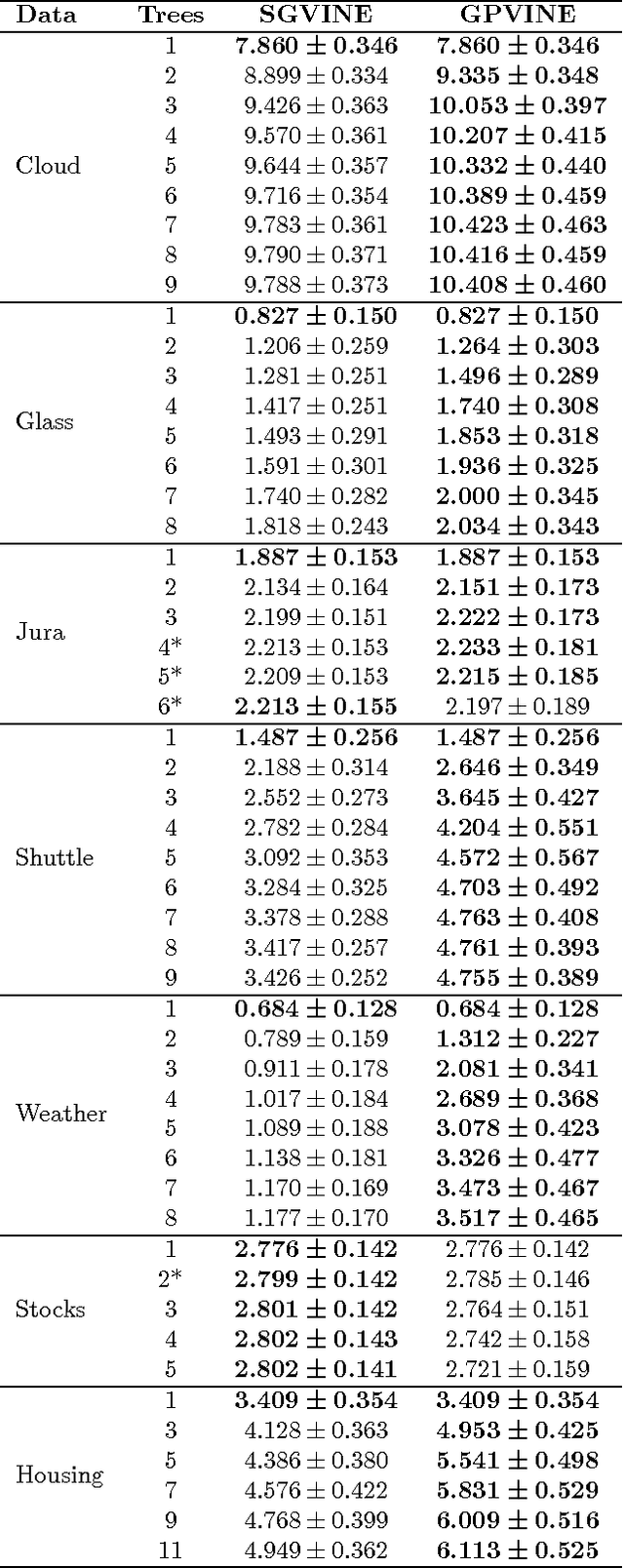
Abstract:Copulas allow to learn marginal distributions separately from the multivariate dependence structure (copula) that links them together into a density function. Vine factorizations ease the learning of high-dimensional copulas by constructing a hierarchy of conditional bivariate copulas. However, to simplify inference, it is common to assume that each of these conditional bivariate copulas is independent from its conditioning variables. In this paper, we relax this assumption by discovering the latent functions that specify the shape of a conditional copula given its conditioning variables We learn these functions by following a Bayesian approach based on sparse Gaussian processes with expectation propagation for scalable, approximate inference. Experiments on real-world datasets show that, when modeling all conditional dependencies, we obtain better estimates of the underlying copula of the data.
On the Convergence of Bound Optimization Algorithms
Oct 19, 2012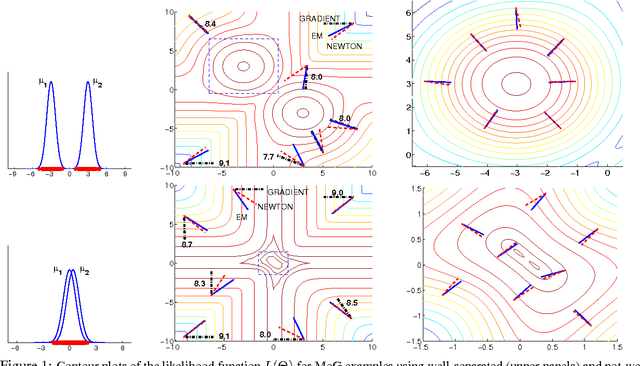
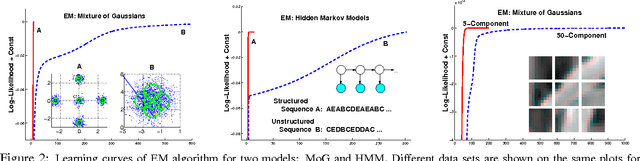
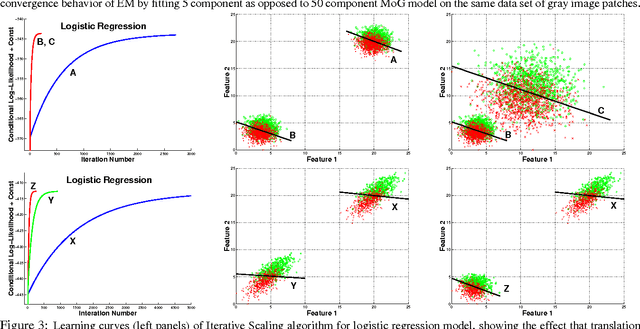
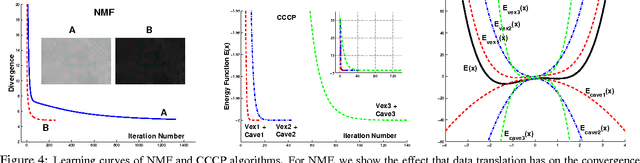
Abstract:Many practitioners who use the EM algorithm complain that it is sometimes slow. When does this happen, and what can be done about it? In this paper, we study the general class of bound optimization algorithms - including Expectation-Maximization, Iterative Scaling and CCCP - and their relationship to direct optimization algorithms such as gradient-based methods for parameter learning. We derive a general relationship between the updates performed by bound optimization methods and those of gradient and second-order methods and identify analytic conditions under which bound optimization algorithms exhibit quasi-Newton behavior, and conditions under which they possess poor, first-order convergence. Based on this analysis, we consider several specific algorithms, interpret and analyze their convergence properties and provide some recipes for preprocessing input to these algorithms to yield faster convergence behavior. We report empirical results supporting our analysis and showing that simple data preprocessing can result in dramatically improved performance of bound optimizers in practice.
Restricting exchangeable nonparametric distributions
Sep 05, 2012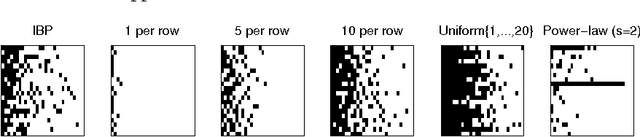


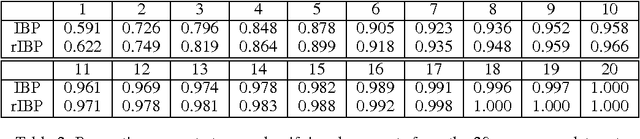
Abstract:Distributions over exchangeable matrices with infinitely many columns, such as the Indian buffet process, are useful in constructing nonparametric latent variable models. However, the distribution implied by such models over the number of features exhibited by each data point may be poorly- suited for many modeling tasks. In this paper, we propose a class of exchangeable nonparametric priors obtained by restricting the domain of existing models. Such models allow us to specify the distribution over the number of features per data point, and can achieve better performance on data sets where the number of features is not well-modeled by the original distribution.
Bayesian and L1 Approaches to Sparse Unsupervised Learning
Aug 17, 2012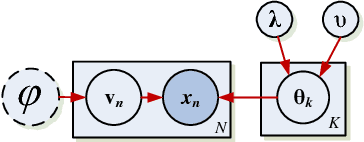
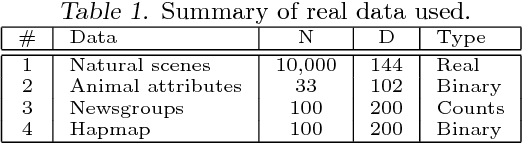
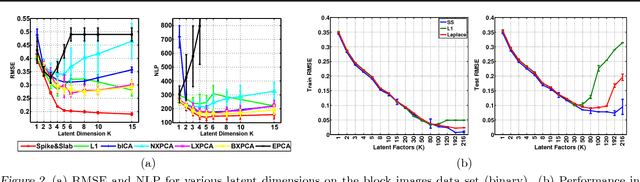
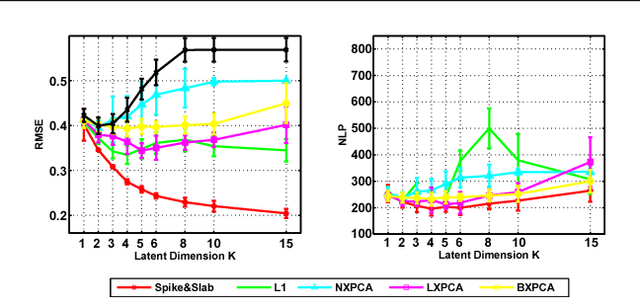
Abstract:The use of L1 regularisation for sparse learning has generated immense research interest, with successful application in such diverse areas as signal acquisition, image coding, genomics and collaborative filtering. While existing work highlights the many advantages of L1 methods, in this paper we find that L1 regularisation often dramatically underperforms in terms of predictive performance when compared with other methods for inferring sparsity. We focus on unsupervised latent variable models, and develop L1 minimising factor models, Bayesian variants of "L1", and Bayesian models with a stronger L0-like sparsity induced through spike-and-slab distributions. These spike-and-slab Bayesian factor models encourage sparsity while accounting for uncertainty in a principled manner and avoiding unnecessary shrinkage of non-zero values. We demonstrate on a number of data sets that in practice spike-and-slab Bayesian methods outperform L1 minimisation, even on a computational budget. We thus highlight the need to re-assess the wide use of L1 methods in sparsity-reliant applications, particularly when we care about generalising to previously unseen data, and provide an alternative that, over many varying conditions, provides improved generalisation performance.
SiGMa: Simple Greedy Matching for Aligning Large Knowledge Bases
Jul 19, 2012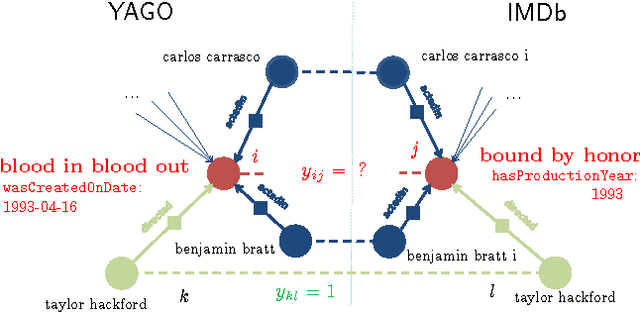
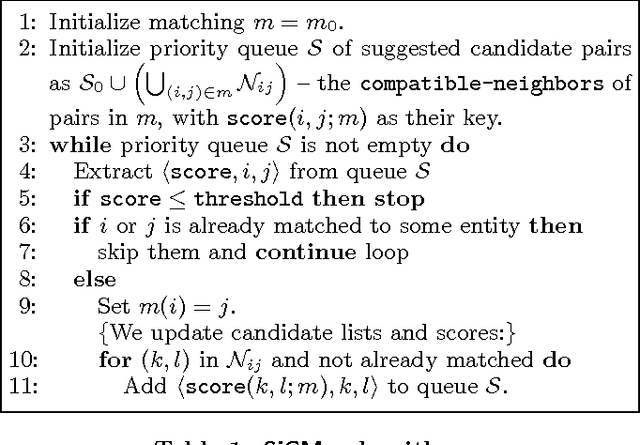

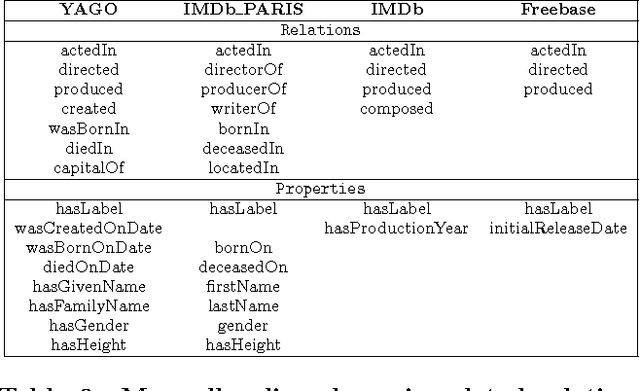
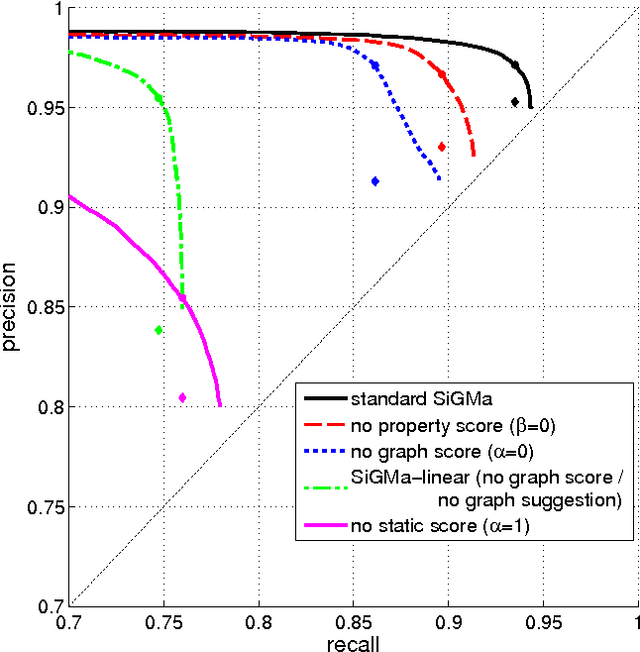
Abstract:The Internet has enabled the creation of a growing number of large-scale knowledge bases in a variety of domains containing complementary information. Tools for automatically aligning these knowledge bases would make it possible to unify many sources of structured knowledge and answer complex queries. However, the efficient alignment of large-scale knowledge bases still poses a considerable challenge. Here, we present Simple Greedy Matching (SiGMa), a simple algorithm for aligning knowledge bases with millions of entities and facts. SiGMa is an iterative propagation algorithm which leverages both the structural information from the relationship graph as well as flexible similarity measures between entity properties in a greedy local search, thus making it scalable. Despite its greedy nature, our experiments indicate that SiGMa can efficiently match some of the world's largest knowledge bases with high precision. We provide additional experiments on benchmark datasets which demonstrate that SiGMa can outperform state-of-the-art approaches both in accuracy and efficiency.
Bayesian Learning in Undirected Graphical Models: Approximate MCMC algorithms
Jul 11, 2012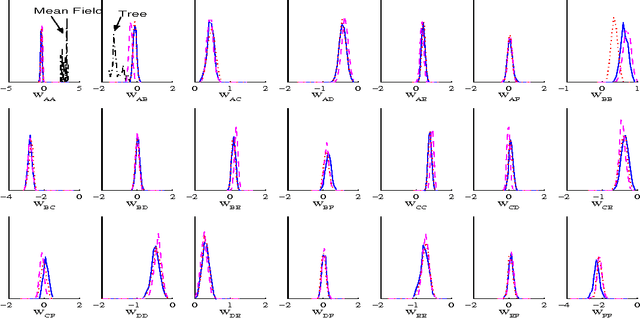
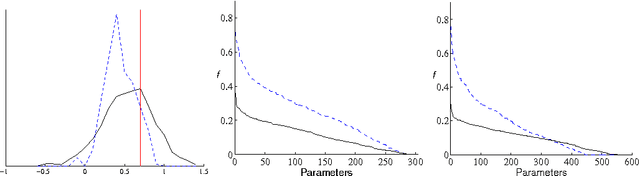
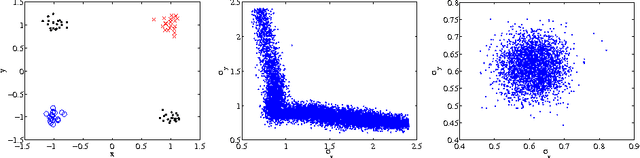
Abstract:Bayesian learning in undirected graphical models|computing posterior distributions over parameters and predictive quantities is exceptionally difficult. We conjecture that for general undirected models, there are no tractable MCMC (Markov Chain Monte Carlo) schemes giving the correct equilibrium distribution over parameters. While this intractability, due to the partition function, is familiar to those performing parameter optimisation, Bayesian learning of posterior distributions over undirected model parameters has been unexplored and poses novel challenges. we propose several approximate MCMC schemes and test on fully observed binary models (Boltzmann machines) for a small coronary heart disease data set and larger artificial systems. While approximations must perform well on the model, their interaction with the sampling scheme is also important. Samplers based on variational mean- field approximations generally performed poorly, more advanced methods using loopy propagation, brief sampling and stochastic dynamics lead to acceptable parameter posteriors. Finally, we demonstrate these techniques on a Markov random field with hidden variables.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge