Yukun Tian
GAC: Stabilizing Asynchronous RL Training for LLMs via Gradient Alignment Control
Mar 02, 2026Abstract:Asynchronous execution is essential for scaling reinforcement learning (RL) to modern large model workloads, including large language models and AI agents, but it can fundamentally alter RL optimization behavior. While prior work on asynchronous RL focuses on training throughput and distributional correction, we show that naively applying asynchrony to policy-gradient updates can induce qualitatively different training dynamics and lead to severe training instability. Through systematic empirical and theoretical analysis, we identify a key signature of this instability: asynchronous training exhibits persistently high cosine similarity between consecutive policy gradients, in contrast to the near-orthogonal updates observed under synchronized training. This stale-aligned gradient effect amplifies correlated updates and increases the risk of overshooting and divergence. Motivated by this observation, we propose GRADIENT ALIGNMENT CONTROL(GAC), a simple dynamics-aware stabilization method that regulates asynchronous RL progress along stale-aligned directions via gradient projection. We establish convergence guarantees under bounded staleness and demonstrate empirically that GAC recovers stable, on-policy training dynamics and matches synchronized baselines even at high staleness.
EventAug: Multifaceted Spatio-Temporal Data Augmentation Methods for Event-based Learning
Sep 18, 2024



Abstract:The event camera has demonstrated significant success across a wide range of areas due to its low time latency and high dynamic range. However, the community faces challenges such as data deficiency and limited diversity, often resulting in over-fitting and inadequate feature learning. Notably, the exploration of data augmentation techniques in the event community remains scarce. This work aims to address this gap by introducing a systematic augmentation scheme named EventAug to enrich spatial-temporal diversity. In particular, we first propose Multi-scale Temporal Integration (MSTI) to diversify the motion speed of objects, then introduce Spatial-salient Event Mask (SSEM) and Temporal-salient Event Mask (TSEM) to enrich object variants. Our EventAug can facilitate models learning with richer motion patterns, object variants and local spatio-temporal relations, thus improving model robustness to varied moving speeds, occlusions, and action disruptions. Experiment results show that our augmentation method consistently yields significant improvements across different tasks and backbones (e.g., a 4.87% accuracy gain on DVS128 Gesture). Our code will be publicly available for this community.
PaDNet: Pan-Density Crowd Counting
Nov 07, 2018
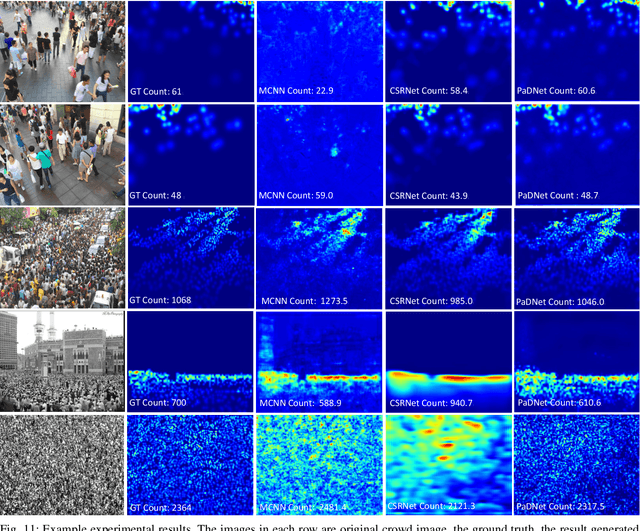
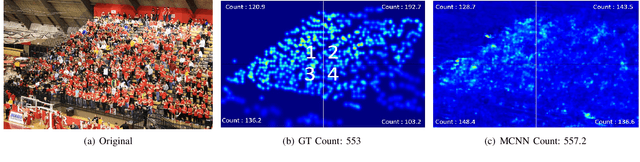
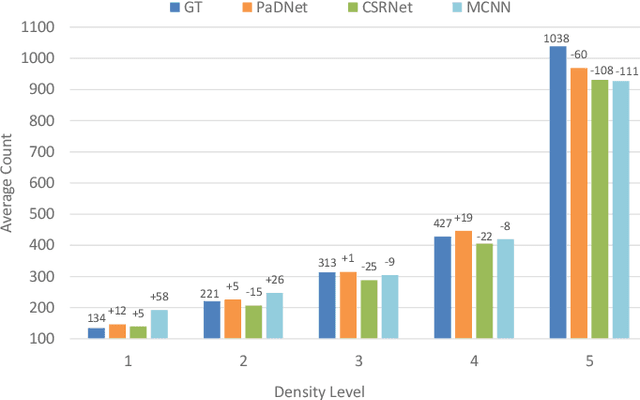


Abstract:Crowd counting in varying density scenes is a challenging problem in artificial intelligence (AI) and pattern recognition. Recently, deep convolutional neural networks (CNNs) are used to tackle this problem. However, the single-column CNN cannot achieve high accuracy and robustness in diverse density scenes. Meanwhile, multi-column CNNs lack effective way to accurately learn the features of different scales for estimating crowd density. To address these issues, we propose a novel pan-density level deep learning model, named as Pan-Density Network (PaDNet). Specifically, the PaDNet learns multi-scale features by three steps. First, several sub-networks are pre-trained on crowd images with different density-levels. Then, a Scale Reinforcement Net (SRN) is utilized to reinforce the scale features. Finally, a Fusion Net fuses all of the scale features to generate the final density map. Experiments on four crowd counting benchmark datasets, the ShanghaiTech, the UCF\_CC\_50, the UCSD, and the UCF-QRNF, indicate that the PaDNet achieves the best performance and has high robustness in pan-density crowd counting compared with other state-of-the-art algorithms.
Soft Activation Mapping of Lung Nodules in Low-Dose CT images
Oct 30, 2018
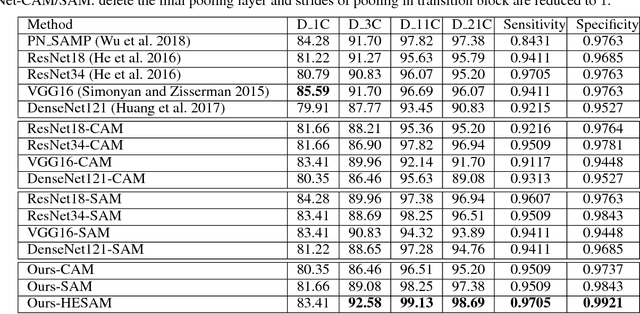
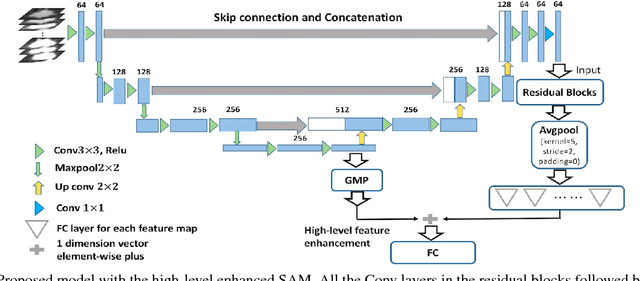
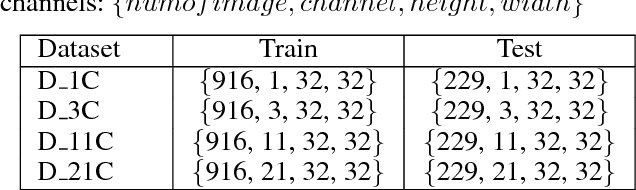
Abstract:As a popular deep learning model, the convolutional neural network (CNN) has produced promising results in analyzing lung nodules and tumors in low-dose CT images. However, this approach still suffers from the lack of labeled data, which is a major challenge for further improvement in the screening and diagnostic performance of CNN. Accurate localization and characterization of nodules provides crucial pathological clues, especially relevant size, attenuation, shape, margins, and growth or stability of lesions, with which the sensitivity and specificity of detection and classification can be increased. To address this challenge, in this paper we develop a soft activation mapping (SAM) to enable fine-grained lesion analysis with a CNN so that it can access rich radiomics features. By combining high-level convolutional features with SAM, we further propose a high-level feature enhancement scheme to localize lesions precisely from multiple CT slices, which helps alleviate overfitting without any additional data augmentation. Experiments on the LIDC-IDRI benchmark dataset indicate that our proposed approach achieves a state-of-the-art predictive performance, reducing the false positive rate. Moreover, the SAM method focuses on irregular margins which are often linked to malignancy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge