Yuan Luo
MedGCN: Graph Convolutional Networks for Multiple Medical Tasks
Mar 31, 2019
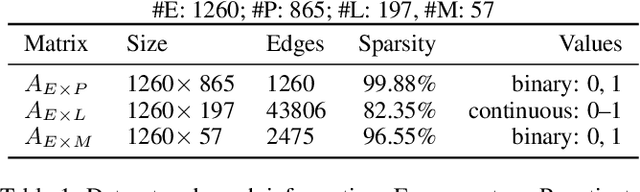
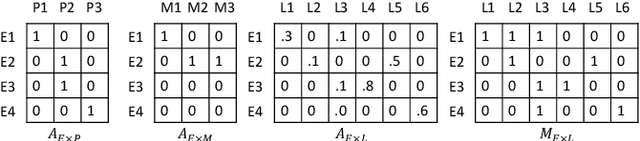
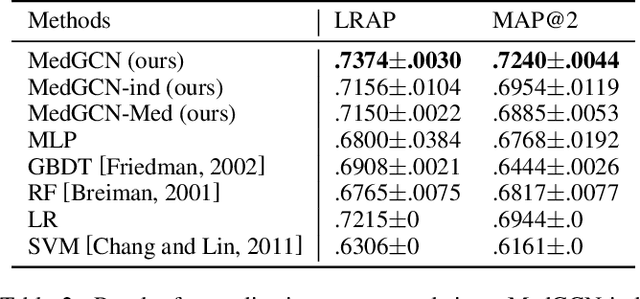
Abstract:Laboratory testing and medication prescription are two of the most important routines in daily clinical practice. Developing an artificial intelligence system that can automatically make lab test imputations and medication recommendations can save cost on potentially redundant lab tests and inform physicians in more effective prescription. We present an intelligent model that can automatically recommend the patients' medications based on their incomplete lab tests, and can even accurately estimate the lab values that have not been taken. We model the complex relations between multiple types of medical entities with their inherent features in a heterogeneous graph. Then we learn a distributed representation for each entity in the graph based on graph convolutional networks to make the representations integrate information from multiple types of entities. Since the entity representations incorporate multiple types of medical information, they can be used for multiple medical tasks. In our experiments, we construct a graph to associate patients, encounters, lab tests and medications, and conduct the two tasks: medication recommendation and lab test imputation. The experimental results demonstrate that our model can outperform the state-of-the-art models in both tasks.
ImageGCN: Multi-Relational Image Graph Convolutional Networks for Disease Identification with Chest X-rays
Mar 31, 2019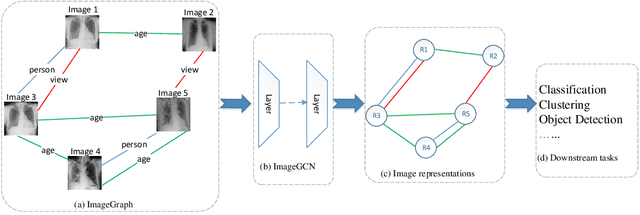
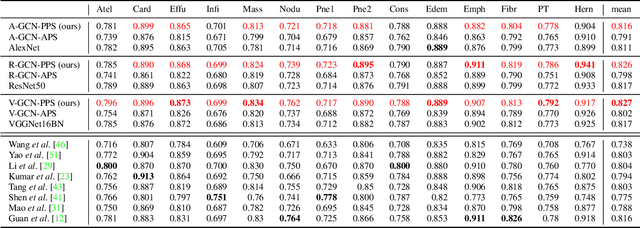
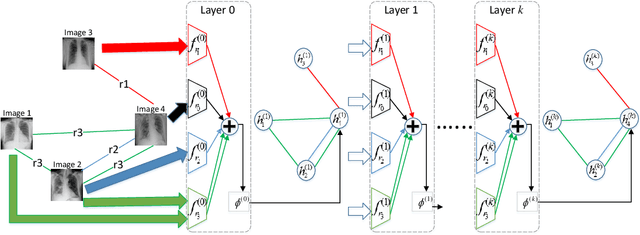
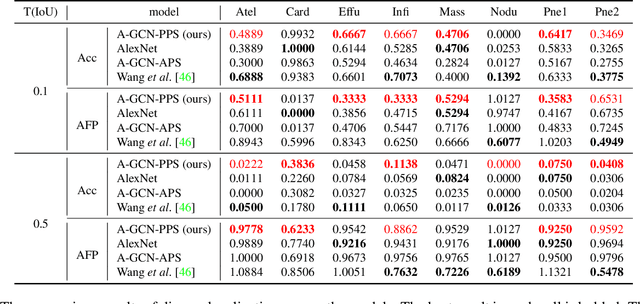
Abstract:Image representation is a fundamental task in computer vision. However, most of the existing approaches for image representation ignore the relations between images and consider each input image independently. Intuitively, relations between images can help to understand the images and maintain model consistency over related images. In this paper, we consider modeling the image-level relations to generate more informative image representations, and propose ImageGCN, an end-to-end graph convolutional network framework for multi-relational image modeling. We also apply ImageGCN to chest X-ray (CXR) images where rich relational information is available for disease identification. Unlike previous image representation models, ImageGCN learns the representation of an image using both its original pixel features and the features of related images. Besides learning informative representations for images, ImageGCN can also be used for object detection in a weakly supervised manner. The Experimental results on ChestX-ray14 dataset demonstrate that ImageGCN can outperform respective baselines in both disease identification and localization tasks and can achieve comparable and often better results than the state-of-the-art methods.
Characterizing Design Patterns of EHR-Driven Phenotype Extraction Algorithms
Nov 15, 2018
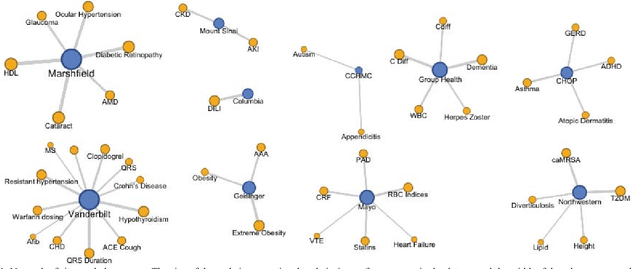
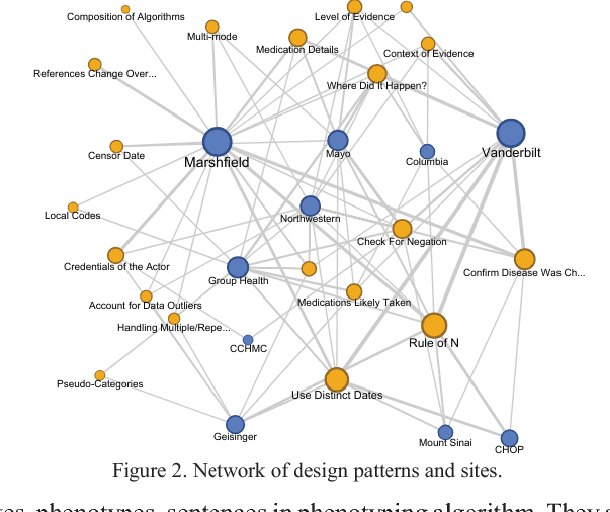
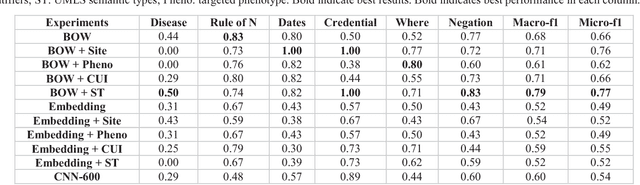
Abstract:The automatic development of phenotype algorithms from Electronic Health Record data with machine learning (ML) techniques is of great interest given the current practice is very time-consuming and resource intensive. The extraction of design patterns from phenotype algorithms is essential to understand their rationale and standard, with great potential to automate the development process. In this pilot study, we perform network visualization on the design patterns and their associations with phenotypes and sites. We classify design patterns using the fragments from previously annotated phenotype algorithms as the ground truth. The classification performance is used as a proxy for coherence at the attribution level. The bag-of-words representation with knowledge-based features generated a good performance in the classification task (0.79 macro-f1 scores). Good classification accuracy with simple features demonstrated the attribution coherence and the feasibility of automatic identification of design patterns. Our results point to both the feasibility and challenges of automatic identification of phenotyping design patterns, which would power the automatic development of phenotype algorithms.
Implementing a Portable Clinical NLP System with a Common Data Model - a Lisp Perspective
Nov 15, 2018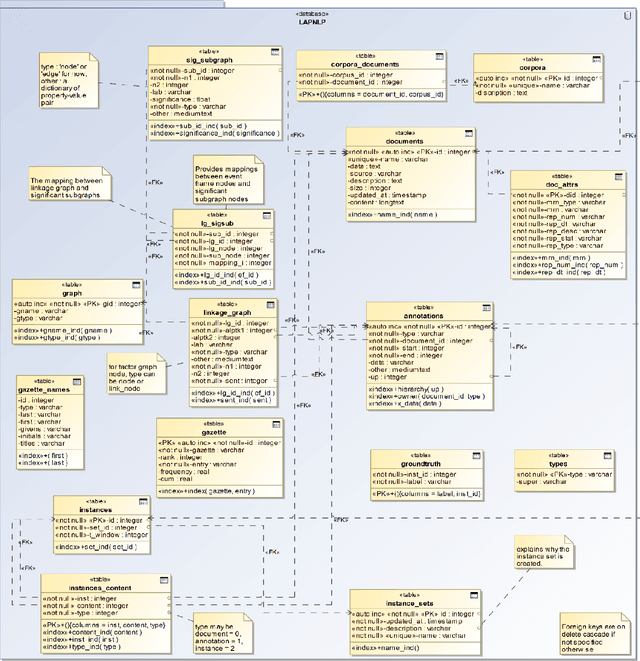
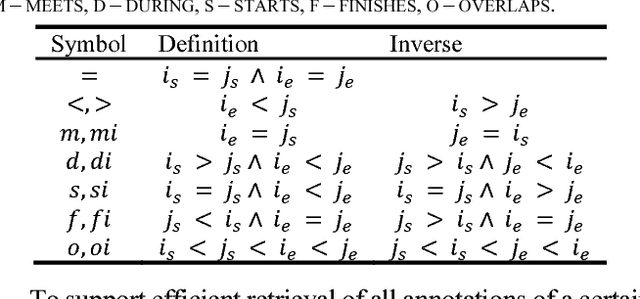

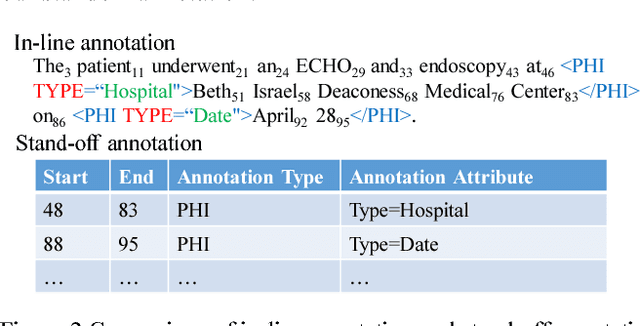
Abstract:This paper presents a Lisp architecture for a portable NLP system, termed LAPNLP, for processing clinical notes. LAPNLP integrates multiple standard, customized and in-house developed NLP tools. Our system facilitates portability across different institutions and data systems by incorporating an enriched Common Data Model (CDM) to standardize necessary data elements. It utilizes UMLS to perform domain adaptation when integrating generic domain NLP tools. It also features stand-off annotations that are specified by positional reference to the original document. We built an interval tree based search engine to efficiently query and retrieve the stand-off annotations by specifying positional requirements. We also developed a utility to convert an inline annotation format to stand-off annotations to enable the reuse of clinical text datasets with inline annotations. We experimented with our system on several NLP facilitated tasks including computational phenotyping for lymphoma patients and semantic relation extraction for clinical notes. These experiments showcased the broader applicability and utility of LAPNLP.
Early Prediction of Acute Kidney Injury in Critical Care Setting Using Clinical Notes
Nov 09, 2018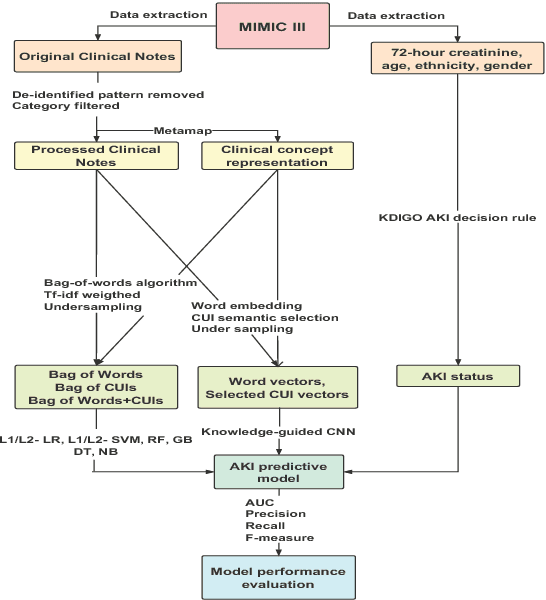
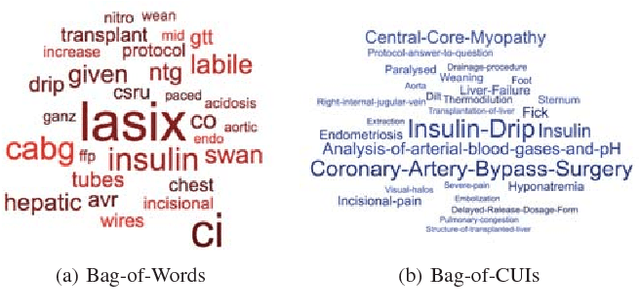

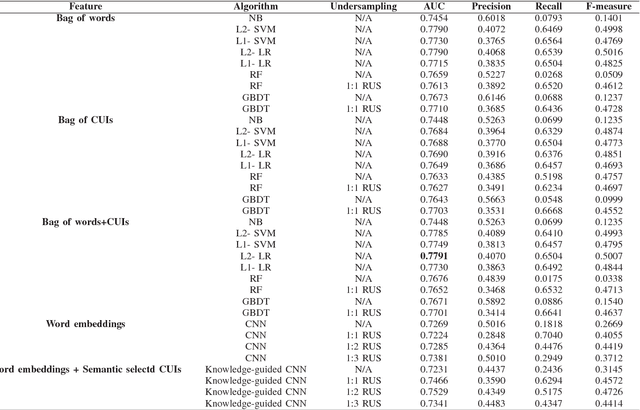
Abstract:Acute kidney injury (AKI) in critically ill patients is associated with significant morbidity and mortality. Development of novel methods to identify patients with AKI earlier will allow for testing of novel strategies to prevent or reduce the complications of AKI. We developed data-driven prediction models to estimate the risk of new AKI onset. We generated models from clinical notes within the first 24 hours following intensive care unit (ICU) admission extracted from Medical Information Mart for Intensive Care III (MIMIC-III). From the clinical notes, we generated clinically meaningful word and concept representations and embeddings, respectively. Five supervised learning classifiers and knowledge-guided deep learning architecture were used to construct prediction models. The best configuration yielded a competitive AUC of 0.779. Our work suggests that natural language processing of clinical notes can be applied to assist clinicians in identifying the risk of incident AKI onset in critically ill patients upon admission to the ICU.
Graph Convolutional Networks for Text Classification
Oct 17, 2018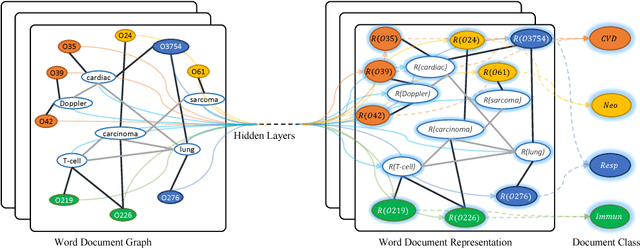
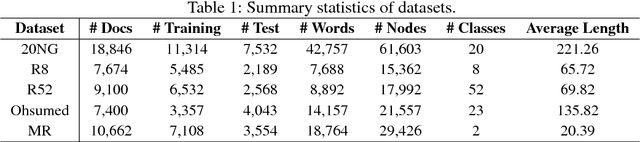
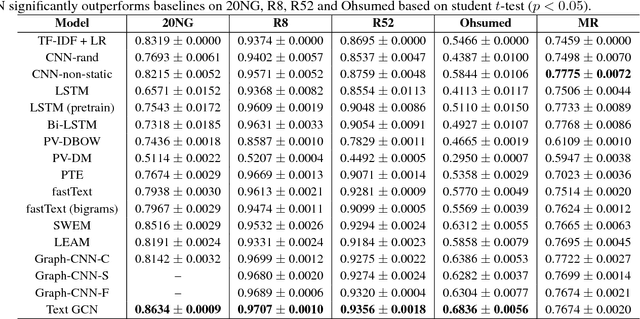
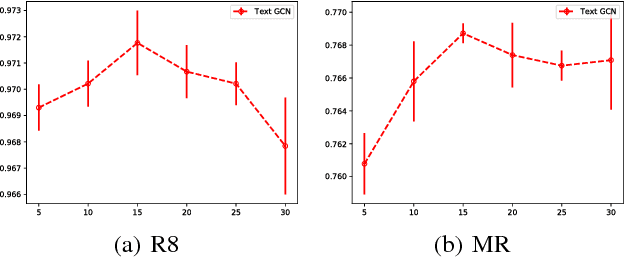
Abstract:Text Classification is an important and classical problem in natural language processing. There have been a number of studies that applied convolutional neural networks (convolution on regular grid, e.g., sequence) to classification. However, only a limited number of studies have explored the more flexible graph convolutional neural networks (convolution on non-grid, e.g., arbitrary graph) for the task. In this work, we propose to use graph convolutional networks for text classification. We build a single text graph for a corpus based on word co-occurrence and document word relations, then learn a Text Graph Convolutional Network (Text GCN) for the corpus. Our Text GCN is initialized with one-hot representation for word and document, it then jointly learns the embeddings for both words and documents, as supervised by the known class labels for documents. Our experimental results on multiple benchmark datasets demonstrate that a vanilla Text GCN without any external word embeddings or knowledge outperforms state-of-the-art methods for text classification. On the other hand, Text GCN also learns predictive word and document embeddings. In addition, experimental results show that the improvement of Text GCN over state-of-the-art comparison methods become more prominent as we lower the percentage of training data, suggesting the robustness of Text GCN to less training data in text classification.
Supervised Nonnegative Matrix Factorization to Predict ICU Mortality Risk
Oct 09, 2018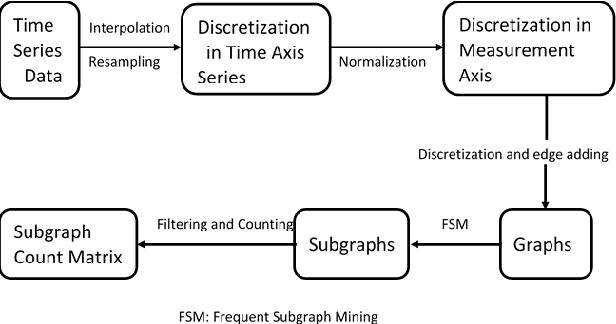
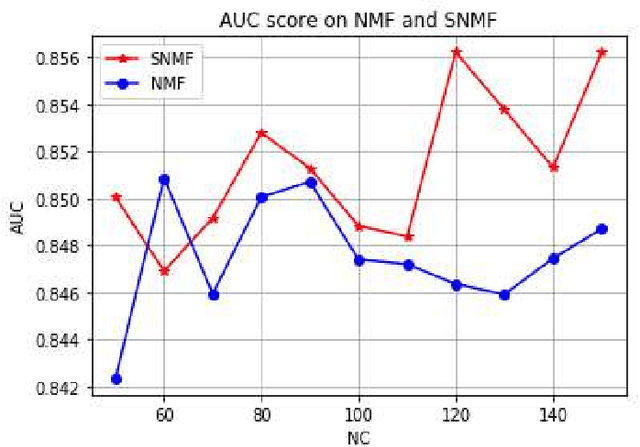
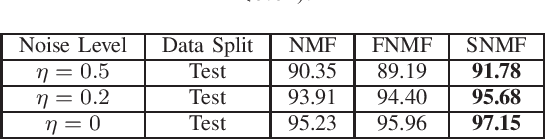
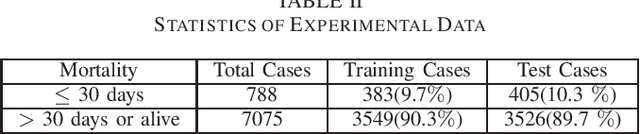
Abstract:ICU mortality risk prediction is a tough yet important task. On one hand, due to the complex temporal data collected, it is difficult to identify the effective features and interpret them easily; on the other hand, good prediction can help clinicians take timely actions to prevent the mortality. These correspond to the interpretability and accuracy problems. Most existing methods lack of the interpretability, but recently Subgraph Augmented Nonnegative Matrix Factorization (SANMF) has been successfully applied to time series data to provide a path to interpret the features well. Therefore, we adopted this approach as the backbone to analyze the patient data. One limitation of the raw SANMF method is its poor prediction ability due to its unsupervised nature. To deal with this problem, we proposed a supervised SANMF algorithm by integrating the logistic regression loss function into the NMF framework and solved it with an alternating optimization procedure. We used the simulation data to verify the effectiveness of this method, and then we applied it to ICU mortality risk prediction and demonstrated its superiority over other conventional supervised NMF methods.
Deep Generative Classifiers for Thoracic Disease Diagnosis with Chest X-ray Images
Sep 20, 2018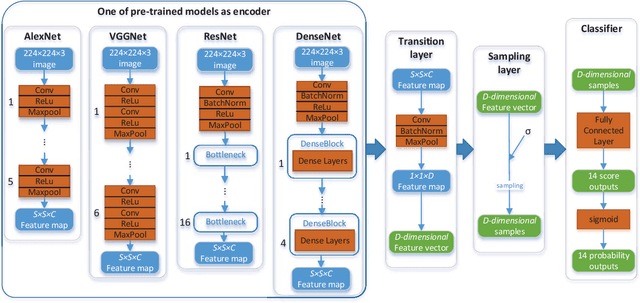
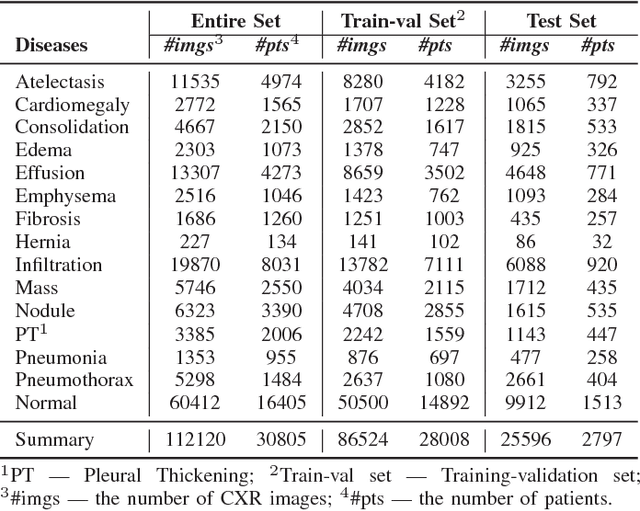
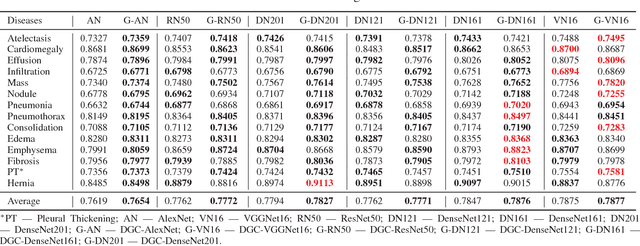
Abstract:Thoracic diseases are very serious health problems that plague a large number of people. Chest X-ray is currently one of the most popular methods to diagnose thoracic diseases, playing an important role in the healthcare workflow. However, reading the chest X-ray images and giving an accurate diagnosis remain challenging tasks for expert radiologists. With the success of deep learning in computer vision, a growing number of deep neural network architectures were applied to chest X-ray image classification. However, most of the previous deep neural network classifiers were based on deterministic architectures which are usually very noise-sensitive and are likely to aggravate the overfitting issue. In this paper, to make a deep architecture more robust to noise and to reduce overfitting, we propose using deep generative classifiers to automatically diagnose thorax diseases from the chest X-ray images. Unlike the traditional deterministic classifier, a deep generative classifier has a distribution middle layer in the deep neural network. A sampling layer then draws a random sample from the distribution layer and input it to the following layer for classification. The classifier is generative because the class label is generated from samples of a related distribution. Through training the model with a certain amount of randomness, the deep generative classifiers are expected to be robust to noise and can reduce overfitting and then achieve good performances. We implemented our deep generative classifiers based on a number of well-known deterministic neural network architectures, and tested our models on the chest X-ray14 dataset. The results demonstrated the superiority of deep generative classifiers compared with the corresponding deep deterministic classifiers.
Distribution Networks for Open Set Learning
Sep 20, 2018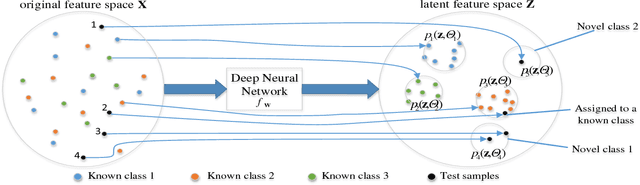
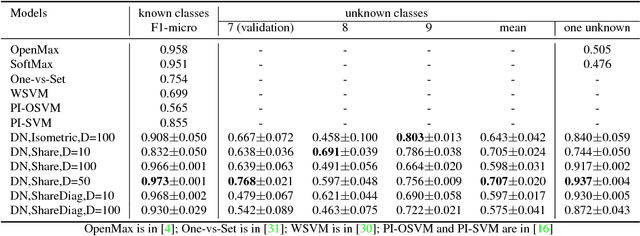
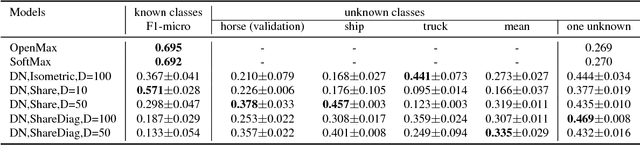
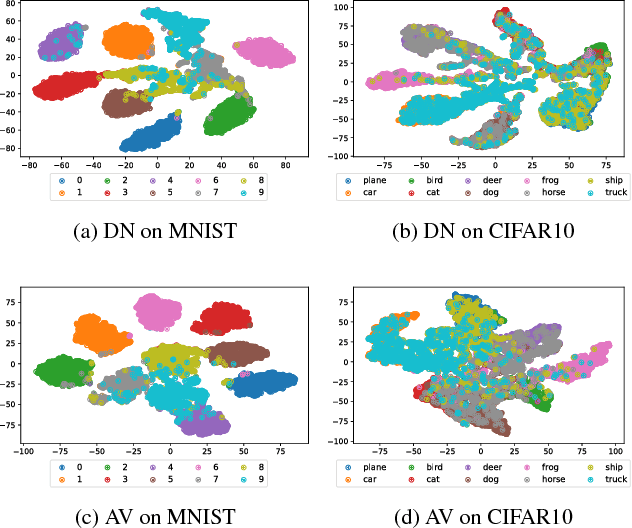
Abstract:In open set learning, a model must be able to generalize to novel classes when it encounters a sample that does not belong to any of the classes it has seen before. Open set learning poses a realistic learning scenario that is receiving growing attention. Existing studies on open set learning mainly focused on detecting novel classes, but few studies tried to model them for differentiating novel classes. We recognize that novel classes should be different from each other, and propose distribution networks for open set learning that can learn and model different novel classes. We hypothesize that, through a certain mapping, samples from different classes with the same classification criterion should follow different probability distributions from the same distribution family. We estimate the probability distribution for each known class and a novel class is detected when a sample is not likely to belong to any of the known distributions. Due to the large feature dimension in the original feature space, the probability distributions in the original feature space are difficult to estimate. Distribution networks map the samples in the original feature space to a latent space where the distributions of known classes can be jointly learned with the network. In the latent space, we also propose a distribution parameter transfer strategy for novel class detection and modeling. By novel class modeling, the detected novel classes can serve as known classes to the subsequent classification. Our experimental results on image datasets MNIST and CIFAR10 and text dataset Ohsumed show that the distribution networks can detect novel classes accurately and model them well for the subsequent classification tasks.
Clinical Text Classification with Rule-based Features and Knowledge-guided Convolutional Neural Networks
Jul 20, 2018



Abstract:Clinical text classification is an important problem in medical natural language processing. Existing studies have conventionally focused on rules or knowledge sources-based feature engineering, but only a few have exploited effective feature learning capability of deep learning methods. In this study, we propose a novel approach which combines rule-based features and knowledge-guided deep learning techniques for effective disease classification. Critical Steps of our method include identifying trigger phrases, predicting classes with very few examples using trigger phrases and training a convolutional neural network with word embeddings and Unified Medical Language System (UMLS) entity embeddings. We evaluated our method on the 2008 Integrating Informatics with Biology and the Bedside (i2b2) obesity challenge. The results show that our method outperforms the state of the art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge