Yoshinobu Kawahara
Adaptive action supervision in reinforcement learning from real-world multi-agent demonstrations
May 22, 2023



Abstract:Modeling of real-world biological multi-agents is a fundamental problem in various scientific and engineering fields. Reinforcement learning (RL) is a powerful framework to generate flexible and diverse behaviors in cyberspace; however, when modeling real-world biological multi-agents, there is a domain gap between behaviors in the source (i.e., real-world data) and the target (i.e., cyberspace for RL), and the source environment parameters are usually unknown. In this paper, we propose a method for adaptive action supervision in RL from real-world demonstrations in multi-agent scenarios. We adopt an approach that combines RL and supervised learning by selecting actions of demonstrations in RL based on the minimum distance of dynamic time warping for utilizing the information of the unknown source dynamics. This approach can be easily applied to many existing neural network architectures and provide us with an RL model balanced between reproducibility as imitation and generalization ability to obtain rewards in cyberspace. In the experiments, using chase-and-escape and football tasks with the different dynamics between the unknown source and target environments, we show that our approach achieved a balance between the reproducibility and the generalization ability compared with the baselines. In particular, we used the tracking data of professional football players as expert demonstrations in football and show successful performances despite the larger gap between behaviors in the source and target environments than the chase-and-escape task.
Modeling Nonlinear Dynamics in Continuous Time with Inductive Biases on Decay Rates and/or Frequencies
Dec 26, 2022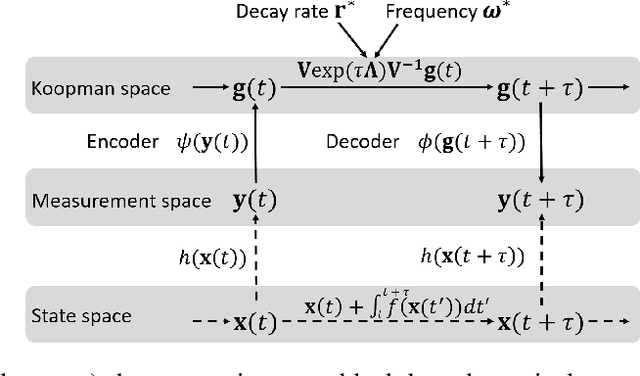
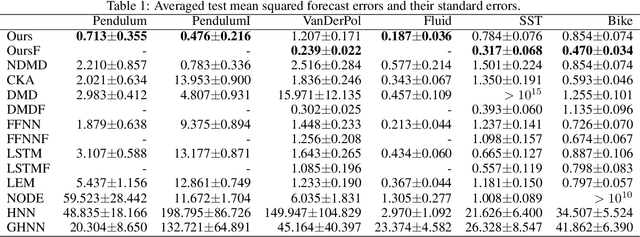
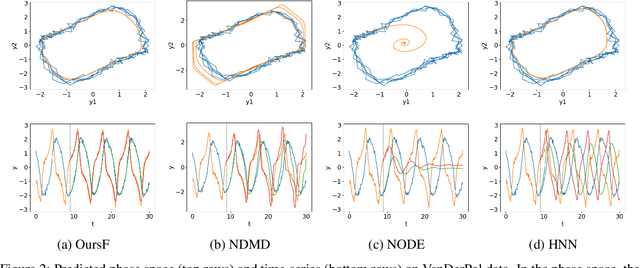
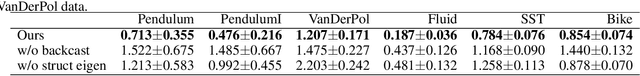
Abstract:We propose a neural network-based model for nonlinear dynamics in continuous time that can impose inductive biases on decay rates and/or frequencies. Inductive biases are helpful for training neural networks especially when training data are small. The proposed model is based on the Koopman operator theory, where the decay rate and frequency information is used by restricting the eigenvalues of the Koopman operator that describe linear evolution in a Koopman space. We use neural networks to find an appropriate Koopman space, which are trained by minimizing multi-step forecasting and backcasting errors using irregularly sampled time-series data. Experiments on various time-series datasets demonstrate that the proposed method achieves higher forecasting performance given a single short training sequence than the existing methods.
Data-driven End-to-end Learning of Pole Placement Control for Nonlinear Dynamics via Koopman Invariant Subspaces
Aug 16, 2022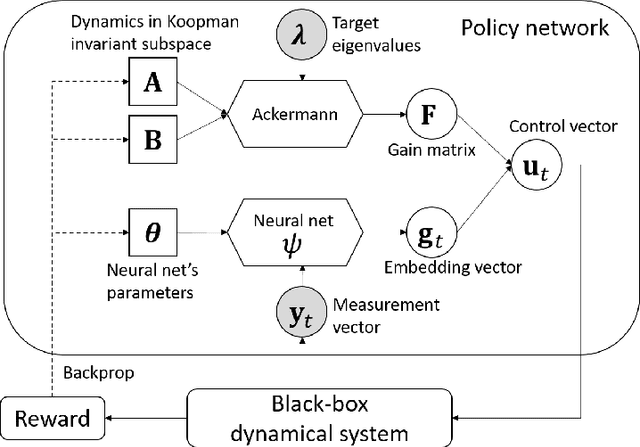
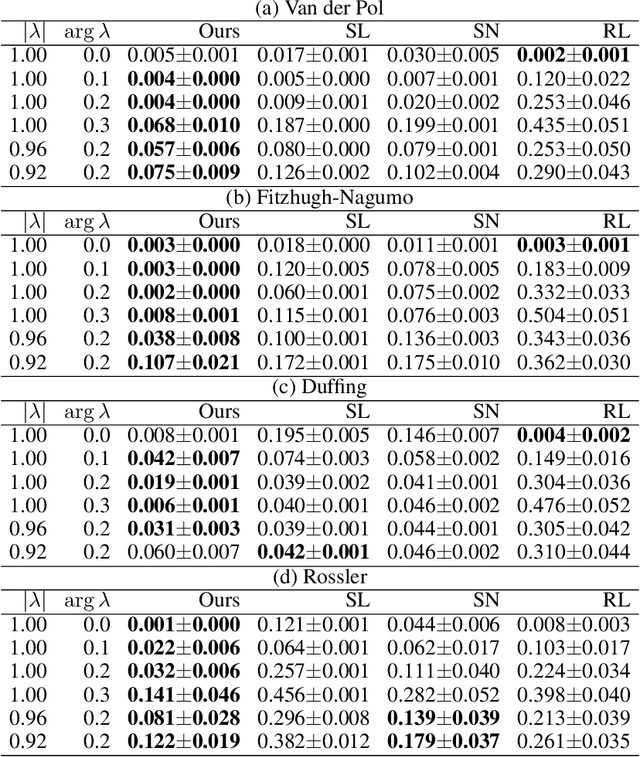

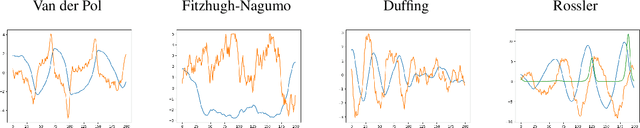
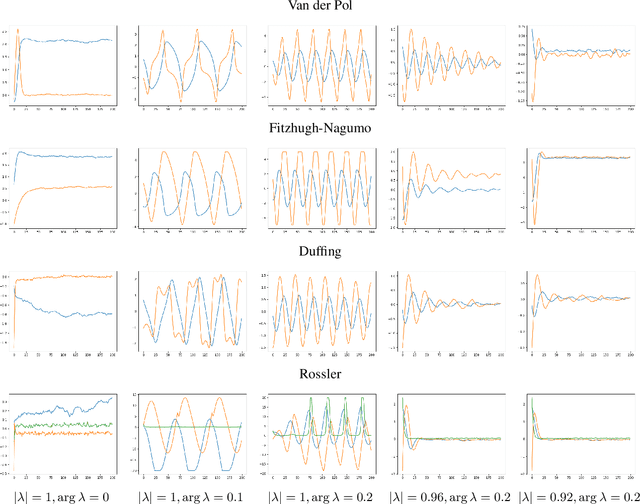
Abstract:We propose a data-driven method for controlling the frequency and convergence rate of black-box nonlinear dynamical systems based on the Koopman operator theory. With the proposed method, a policy network is trained such that the eigenvalues of a Koopman operator of controlled dynamics are close to the target eigenvalues. The policy network consists of a neural network to find a Koopman invariant subspace, and a pole placement module to adjust the eigenvalues of the Koopman operator. Since the policy network is differentiable, we can train it in an end-to-end fashion using reinforcement learning. We demonstrate that the proposed method achieves better performance than model-free reinforcement learning and model-based control with system identification.
Stable Invariant Models via Koopman Spectra
Jul 15, 2022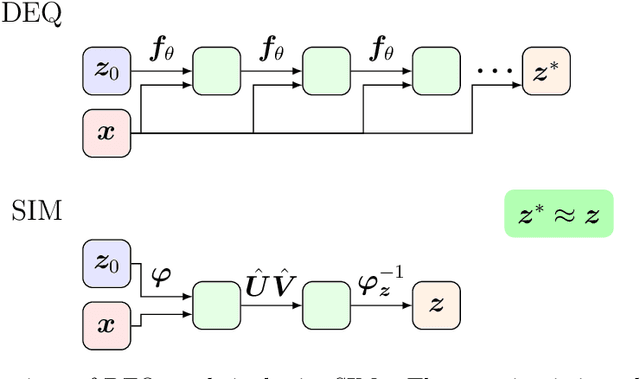

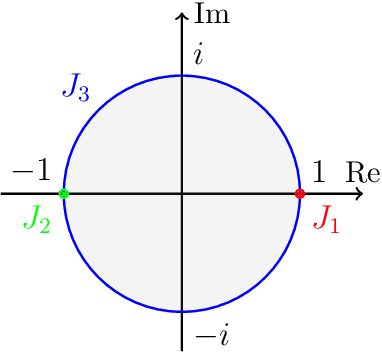
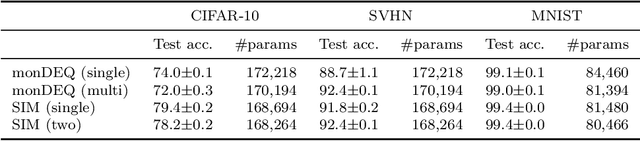
Abstract:Weight-tied models have attracted attention in the modern development of neural networks. The deep equilibrium model (DEQ) represents infinitely deep neural networks with weight-tying, and recent studies have shown the potential of this type of approach. DEQs are needed to iteratively solve root-finding problems in training and are built on the assumption that the underlying dynamics determined by the models converge to a fixed point. In this paper, we present the stable invariant model (SIM), a new class of deep models that in principle approximates DEQs under stability and extends the dynamics to more general ones converging to an invariant set (not restricted in a fixed point). The key ingredient in deriving SIMs is a representation of the dynamics with the spectra of the Koopman and Perron--Frobenius operators. This perspective approximately reveals stable dynamics with DEQs and then derives two variants of SIMs. We also propose an implementation of SIMs that can be learned in the same way as feedforward models. We illustrate the empirical performance of SIMs with experiments and demonstrate that SIMs achieve comparative or superior performance against DEQs in several learning tasks.
Estimating counterfactual treatment outcomes over time in complex multi-agent scenarios
Jun 04, 2022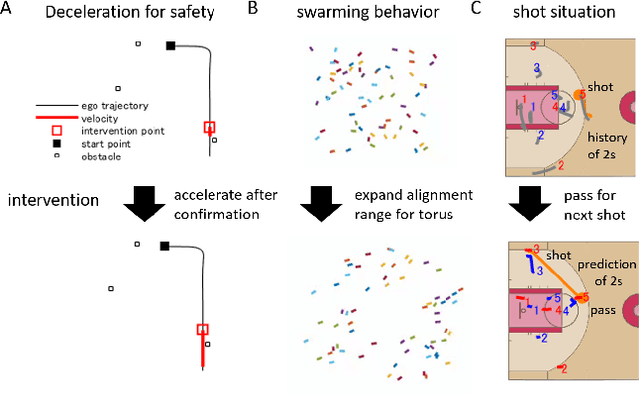
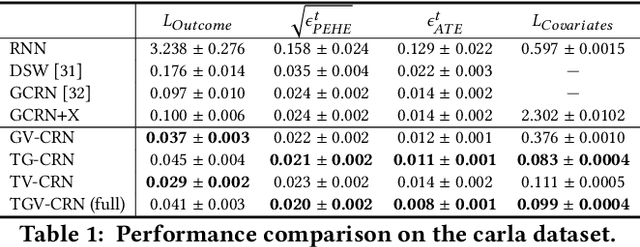
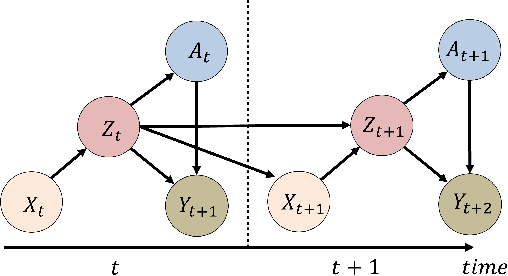
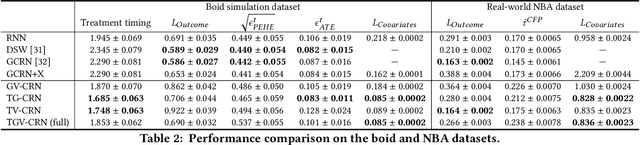
Abstract:Evaluation of intervention in a multi-agent system, e.g., when humans should intervene in autonomous driving systems and when a player should pass to teammates for a good shot, is challenging in various engineering and scientific fields. Estimating the individual treatment effect (ITE) using counterfactual long-term prediction is practical to evaluate such interventions. However, most of the conventional frameworks did not consider the time-varying complex structure of multi-agent relationships and covariate counterfactual prediction. This may sometimes lead to erroneous assessments of ITE and interpretation problems. Here we propose an interpretable, counterfactual recurrent network in multi-agent systems to estimate the effect of the intervention. Our model leverages graph variational recurrent neural networks and theory-based computation with domain knowledge for the ITE estimation framework based on long-term prediction of multi-agent covariates and outcomes, which can confirm under the circumstances under which the intervention is effective. On simulated models of an automated vehicle and biological agents with time-varying confounders, we show that our methods achieved lower estimation errors in counterfactual covariates and the most effective treatment timing than the baselines. Furthermore, using real basketball data, our methods performed realistic counterfactual predictions and evaluated the counterfactual passes in shot scenarios.
Koopman Q-learning: Offline Reinforcement Learning via Symmetries of Dynamics
Nov 02, 2021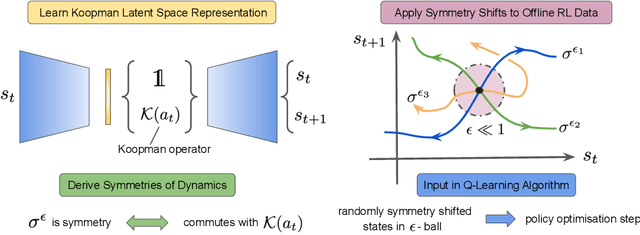
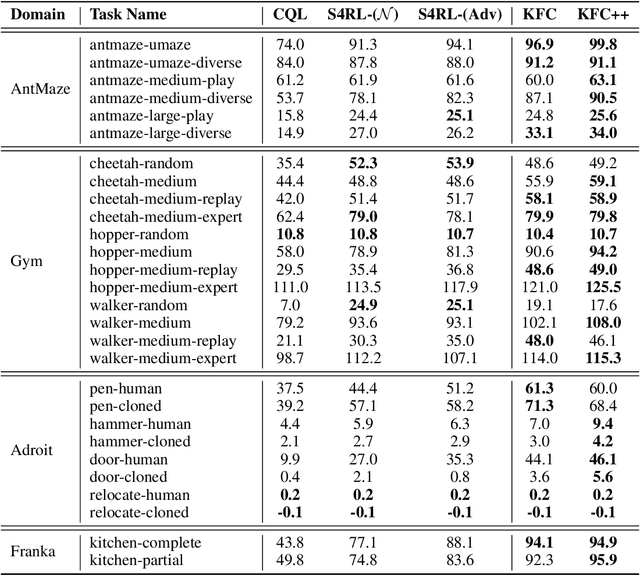
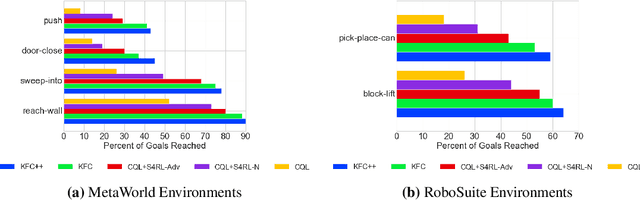
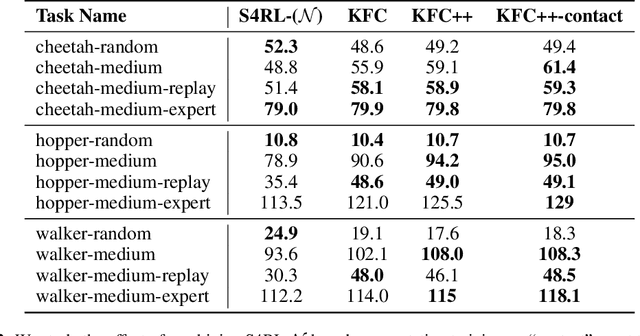
Abstract:Offline reinforcement learning leverages large datasets to train policies without interactions with the environment. The learned policies may then be deployed in real-world settings where interactions are costly or dangerous. Current algorithms over-fit to the training dataset and as a consequence perform poorly when deployed to out-of-distribution generalizations of the environment. We aim to address these limitations by learning a Koopman latent representation which allows us to infer symmetries of the system's underlying dynamic. The latter is then utilized to extend the otherwise static offline dataset during training; this constitutes a novel data augmentation framework which reflects the system's dynamic and is thus to be interpreted as an exploration of the environments phase space. To obtain the symmetries we employ Koopman theory in which nonlinear dynamics are represented in terms of a linear operator acting on the space of measurement functions of the system and thus symmetries of the dynamics may be inferred directly. We provide novel theoretical results on the existence and nature of symmetries relevant for control systems such as reinforcement learning settings. Moreover, we empirically evaluate our method on several benchmark offline reinforcement learning tasks and datasets including D4RL, Metaworld and Robosuite and find that by using our framework we consistently improve the state-of-the-art for Q-learning methods.
Learning interaction rules from multi-animal trajectories via augmented behavioral models
Jul 14, 2021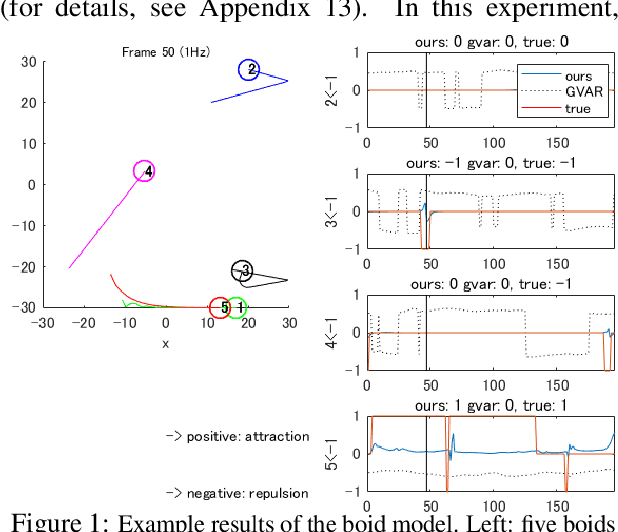
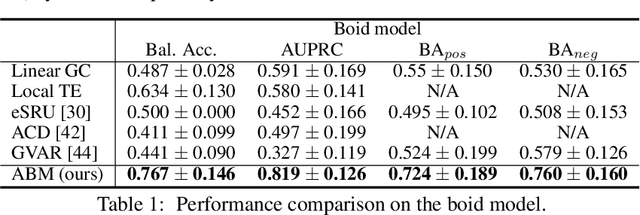
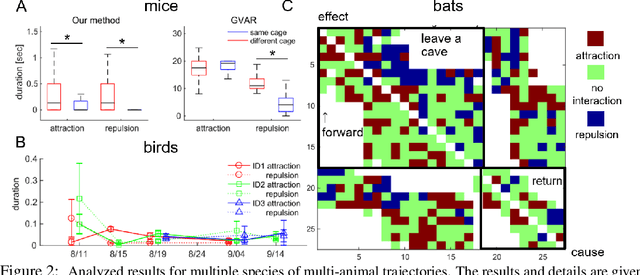
Abstract:Extracting the interaction rules of biological agents from moving sequences pose challenges in various domains. Granger causality is a practical framework for analyzing the interactions from observed time-series data; however, this framework ignores the structures of the generative process in animal behaviors, which may lead to interpretational problems and sometimes erroneous assessments of causality. In this paper, we propose a new framework for learning Granger causality from multi-animal trajectories via augmented theory-based behavioral models with interpretable data-driven models. We adopt an approach for augmenting incomplete multi-agent behavioral models described by time-varying dynamical systems with neural networks. For efficient and interpretable learning, our model leverages theory-based architectures separating navigation and motion processes, and the theory-guided regularization for reliable behavioral modeling. This can provide interpretable signs of Granger-causal effects over time, i.e., when specific others cause the approach or separation. In experiments using synthetic datasets, our method achieved better performance than various baselines. We then analyzed multi-animal datasets of mice, flies, birds, and bats, which verified our method and obtained novel biological insights.
Koopman Spectrum Nonlinear Regulator and Provably Efficient Online Learning
Jun 30, 2021
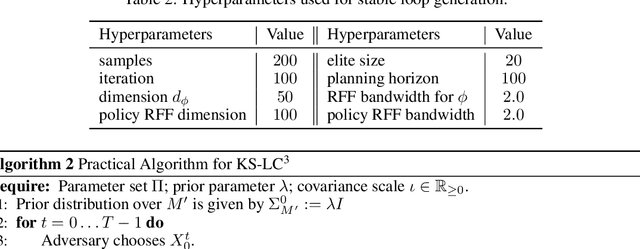

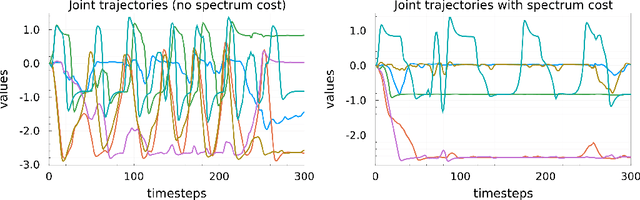

Abstract:Most modern reinforcement learning algorithms optimize a cumulative single-step cost along a trajectory. The optimized motions are often 'unnatural', representing, for example, behaviors with sudden accelerations that waste energy and lack predictability. In this work, we present a novel paradigm of controlling nonlinear systems via the minimization of the Koopman spectrum cost: a cost over the Koopman operator of the controlled dynamics. This induces a broader class of dynamical behaviors that evolve over stable manifolds such as nonlinear oscillators, closed loops, and smooth movements. We demonstrate that some dynamics realizations that are not possible with a cumulative cost are feasible in this paradigm. Moreover, we present a provably efficient online learning algorithm for our problem that enjoys a sub-linear regret bound under some structural assumptions.
A Quadratic Actor Network for Model-Free Reinforcement Learning
Mar 11, 2021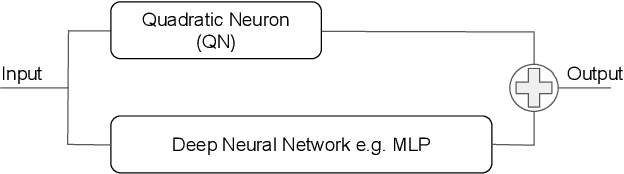
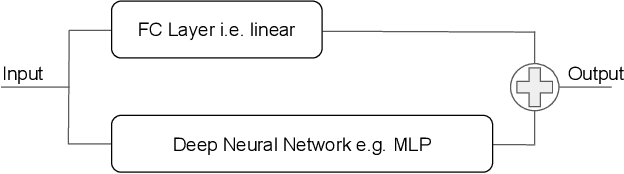
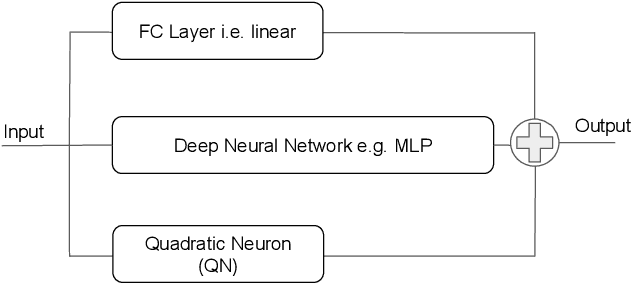
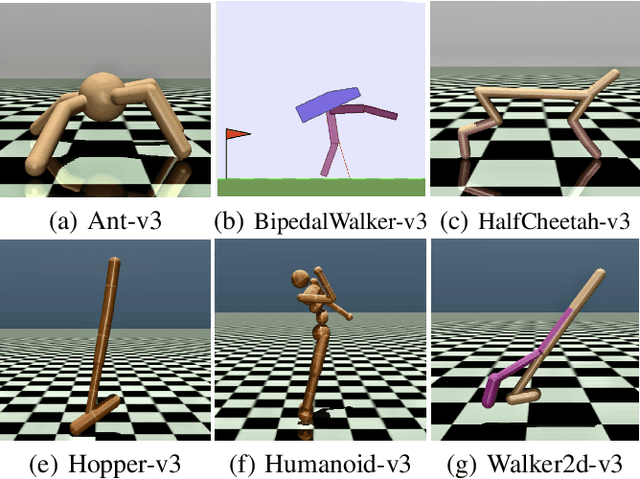
Abstract:In this work we discuss the incorporation of quadratic neurons into policy networks in the context of model-free actor-critic reinforcement learning. Quadratic neurons admit an explicit quadratic function approximation in contrast to conventional approaches where the the non-linearity is induced by the activation functions. We perform empiric experiments on several MuJoCo continuous control tasks and find that when quadratic neurons are added to MLP policy networks those outperform the baseline MLP whilst admitting a smaller number of parameters. The top returned reward is in average increased by $5.8\%$ while being about $21\%$ more sample efficient. Moreover, it can maintain its advantage against added action and observation noise.
Discriminant Dynamic Mode Decomposition for Labeled Spatio-Temporal Data Collections
Feb 19, 2021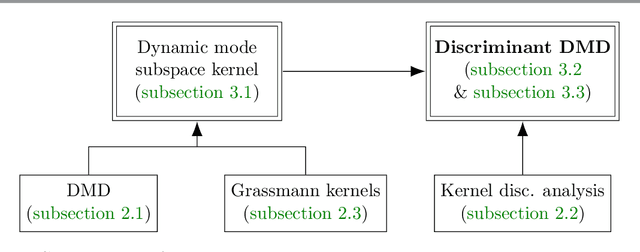
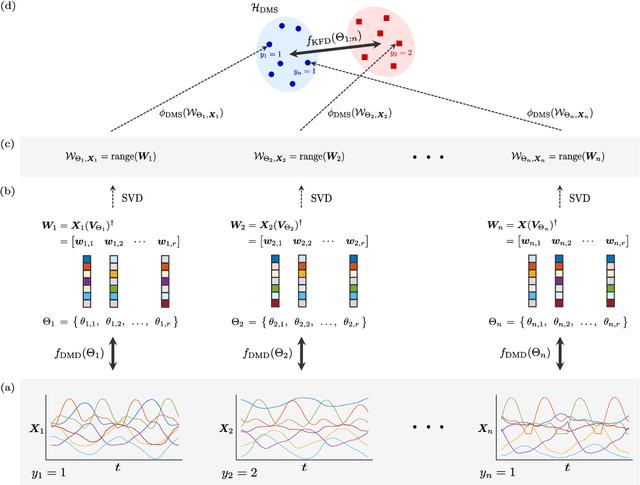
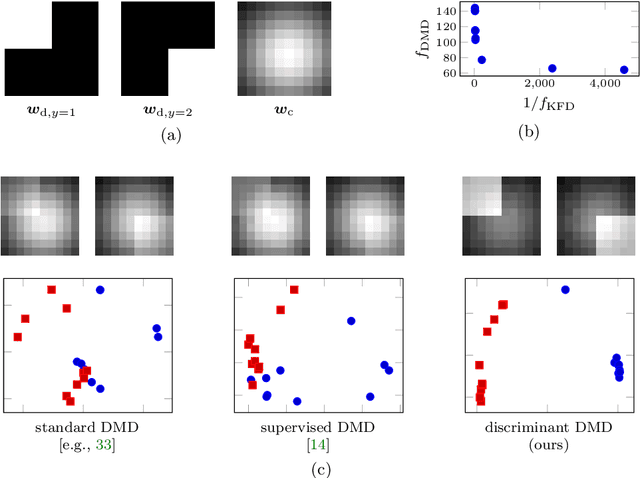
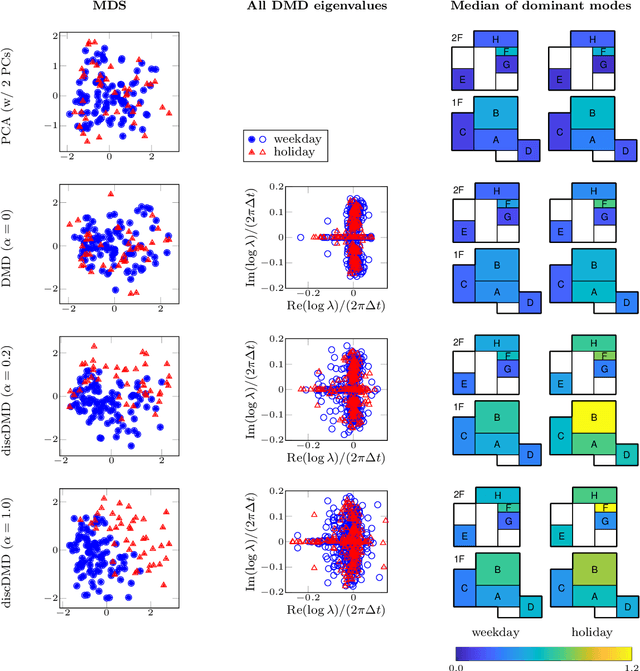
Abstract:Extracting coherent patterns is one of the standard approaches towards understanding spatio-temporal data. Dynamic mode decomposition (DMD) is a powerful tool for extracting coherent patterns, but the original DMD and most of its variants do not consider label information, which is often available as side information of spatio-temporal data. In this work, we propose a new method for extracting distinctive coherent patterns from labeled spatio-temporal data collections, such that they contribute to major differences in a labeled set of dynamics. We achieve such pattern extraction by incorporating discriminant analysis into DMD. To this end, we define a kernel function on subspaces spanned by sets of dynamic modes and develop an objective to take both reconstruction goodness as DMD and class-separation goodness as discriminant analysis into account. We illustrate our method using a synthetic dataset and several real-world datasets. The proposed method can be a useful tool for exploratory data analysis for understanding spatio-temporal data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge