Xiuzhen Huang
Lung cancer screening with low-dose CT scans using a deep learning approach
Jun 01, 2019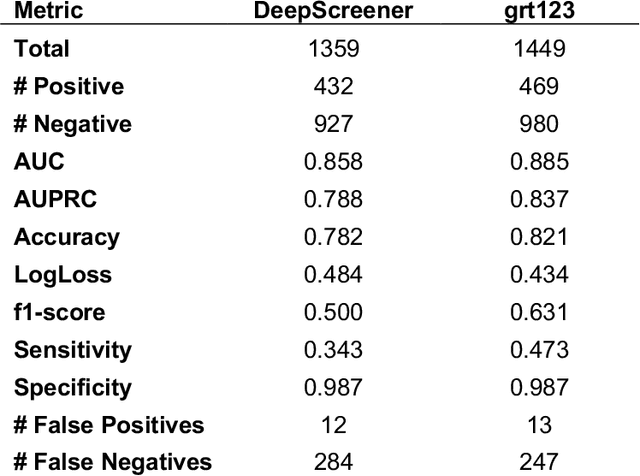
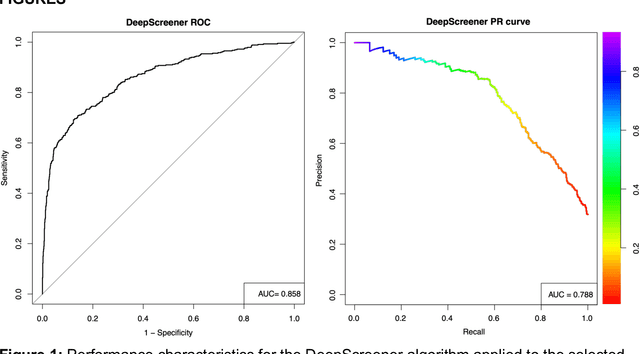
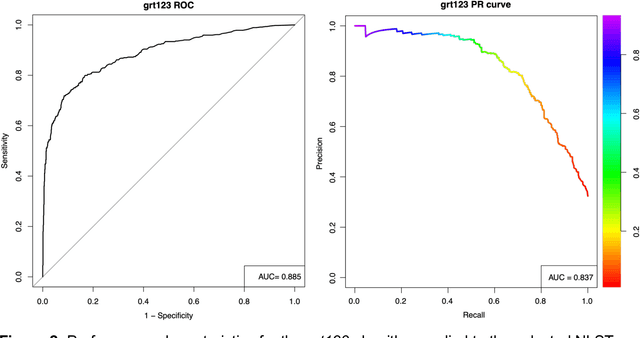
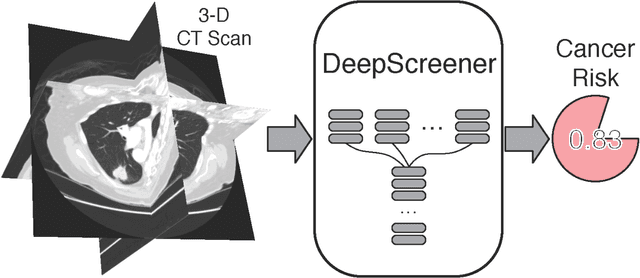
Abstract:Lung cancer is the leading cause of cancer deaths. Early detection through low-dose computed tomography (CT) screening has been shown to significantly reduce mortality but suffers from a high false positive rate that leads to unnecessary diagnostic procedures. Quantitative image analysis coupled to deep learning techniques has the potential to reduce this false positive rate. We conducted a computational analysis of 1449 low-dose CT studies drawn from the National Lung Screening Trial (NLST) cohort. We applied to this cohort our newly developed algorithm, DeepScreener, which is based on a novel deep learning approach. The algorithm, after the training process using about 3000 CT studies, does not require lung nodule annotations to conduct cancer prediction. The algorithm uses consecutive slices and multi-task features to determine whether a nodule is likely to be cancer, and a spatial pyramid to detect nodules at different scales. We find that the algorithm can predict a patient's cancer status from a volumetric lung CT image with high accuracy (78.2%, with area under the Receiver Operating Characteristic curve (AUC) of 0.858). Our preliminary framework ranked 16th of 1972 teams (top 1%) in the Data Science Bowl 2017 (DSB2017) competition, based on the challenge datasets. We report here the application of DeepScreener on an independent NLST test set. This study indicates that the deep learning approach has the potential to significantly reduce the false positive rate in lung cancer screening with low-dose CT scans.
EBIC: an evolutionary-based parallel biclustering algorithm for pattern discover
Jul 26, 2018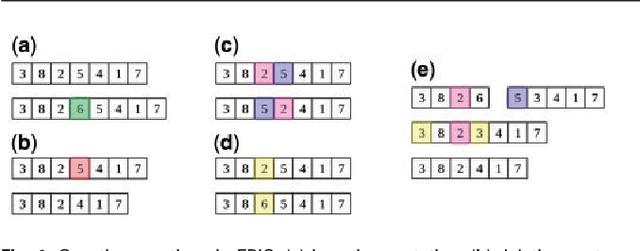
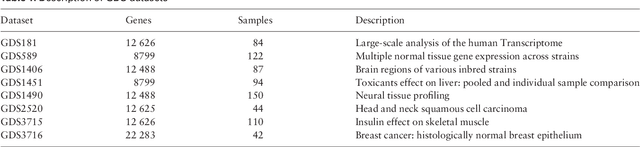
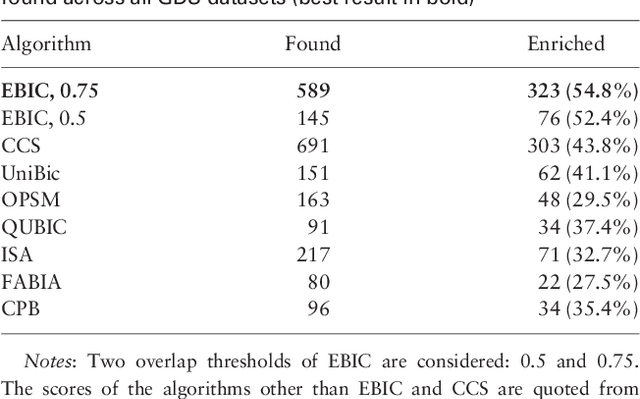
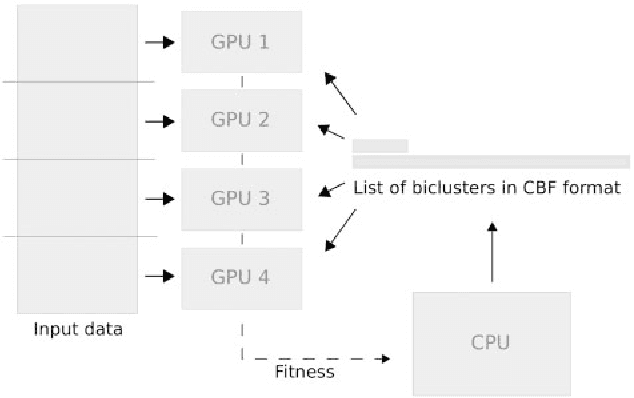
Abstract:In this paper a novel biclustering algorithm based on artificial intelligence (AI) is introduced. The method called EBIC aims to detect biologically meaningful, order-preserving patterns in complex data. The proposed algorithm is probably the first one capable of discovering with accuracy exceeding 50% multiple complex patterns in real gene expression datasets. It is also one of the very few biclustering methods designed for parallel environments with multiple graphics processing units (GPUs). We demonstrate that EBIC outperforms state-of-the-art biclustering methods, in terms of recovery and relevance, on both synthetic and genetic datasets. EBIC also yields results over 12 times faster than the most accurate reference algorithms. The proposed algorithm is anticipated to be added to the repertoire of unsupervised machine learning algorithms for the analysis of datasets, including those from large-scale genomic studies.
Highly accurate model for prediction of lung nodule malignancy with CT scans
Feb 06, 2018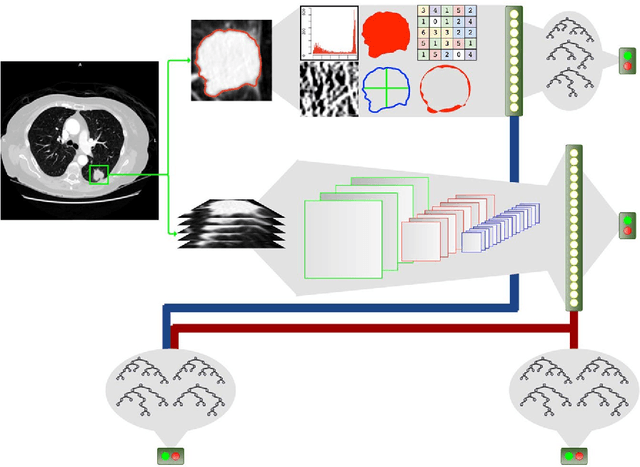
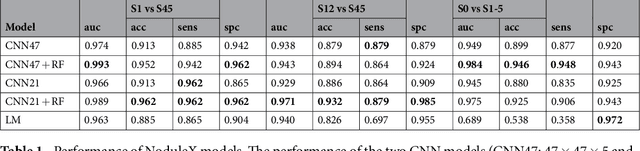
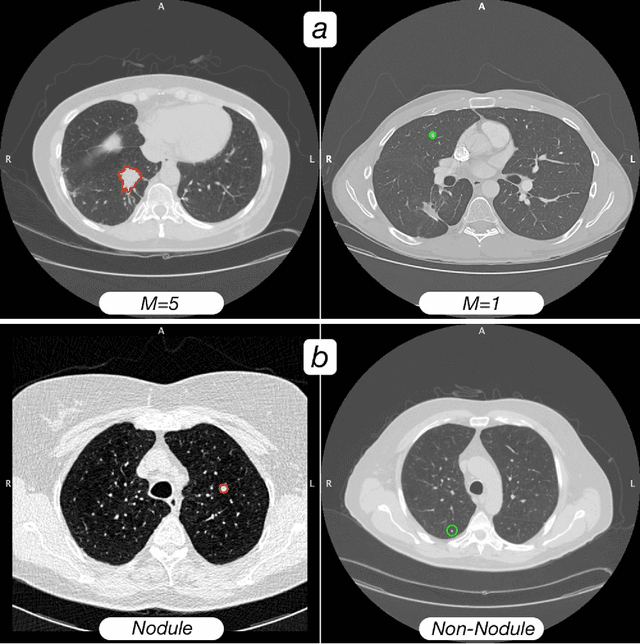
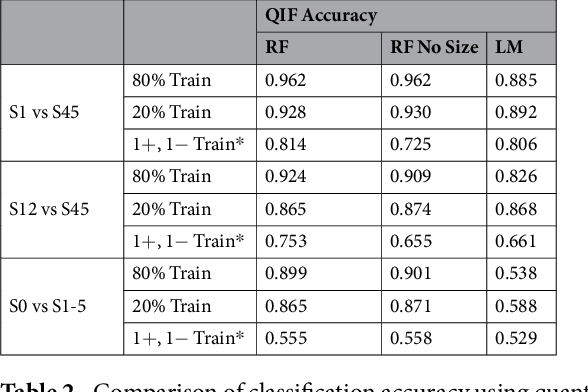
Abstract:Computed tomography (CT) examinations are commonly used to predict lung nodule malignancy in patients, which are shown to improve noninvasive early diagnosis of lung cancer. It remains challenging for computational approaches to achieve performance comparable to experienced radiologists. Here we present NoduleX, a systematic approach to predict lung nodule malignancy from CT data, based on deep learning convolutional neural networks (CNN). For training and validation, we analyze >1000 lung nodules in images from the LIDC/IDRI cohort. All nodules were identified and classified by four experienced thoracic radiologists who participated in the LIDC project. NoduleX achieves high accuracy for nodule malignancy classification, with an AUC of ~0.99. This is commensurate with the analysis of the dataset by experienced radiologists. Our approach, NoduleX, provides an effective framework for highly accurate nodule malignancy prediction with the model trained on a large patient population. Our results are replicable with software available at http://bioinformatics.astate.edu/NoduleX.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge