Weifu Lv
Bridged Semantic Alignment for Zero-shot 3D Medical Image Diagnosis
Jan 07, 2025Abstract:3D medical images such as Computed tomography (CT) are widely used in clinical practice, offering a great potential for automatic diagnosis. Supervised learning-based approaches have achieved significant progress but rely heavily on extensive manual annotations, limited by the availability of training data and the diversity of abnormality types. Vision-language alignment (VLA) offers a promising alternative by enabling zero-shot learning without additional annotations. However, we empirically discover that the visual and textural embeddings after alignment endeavors from existing VLA methods form two well-separated clusters, presenting a wide gap to be bridged. To bridge this gap, we propose a Bridged Semantic Alignment (BrgSA) framework. First, we utilize a large language model to perform semantic summarization of reports, extracting high-level semantic information. Second, we design a Cross-Modal Knowledge Interaction (CMKI) module that leverages a cross-modal knowledge bank as a semantic bridge, facilitating interaction between the two modalities, narrowing the gap, and improving their alignment. To comprehensively evaluate our method, we construct a benchmark dataset that includes 15 underrepresented abnormalities as well as utilize two existing benchmark datasets. Experimental results demonstrate that BrgSA achieves state-of-the-art performances on both public benchmark datasets and our custom-labeled dataset, with significant improvements in zero-shot diagnosis of underrepresented abnormalities.
E3D-GPT: Enhanced 3D Visual Foundation for Medical Vision-Language Model
Oct 18, 2024



Abstract:The development of 3D medical vision-language models holds significant potential for disease diagnosis and patient treatment. However, compared to 2D medical images, 3D medical images, such as CT scans, face challenges related to limited training data and high dimension, which severely restrict the progress of 3D medical vision-language models. To address these issues, we collect a large amount of unlabeled 3D CT data and utilize self-supervised learning to construct a 3D visual foundation model for extracting 3D visual features. Then, we apply 3D spatial convolutions to aggregate and project high-level image features, reducing computational complexity while preserving spatial information. We also construct two instruction-tuning datasets based on BIMCV-R and CT-RATE to fine-tune the 3D vision-language model. Our model demonstrates superior performance compared to existing methods in report generation, visual question answering, and disease diagnosis. Code and data will be made publicly available soon.
Automated assessment of disease severity of COVID-19 using artificial intelligence with synthetic chest CT
Dec 11, 2021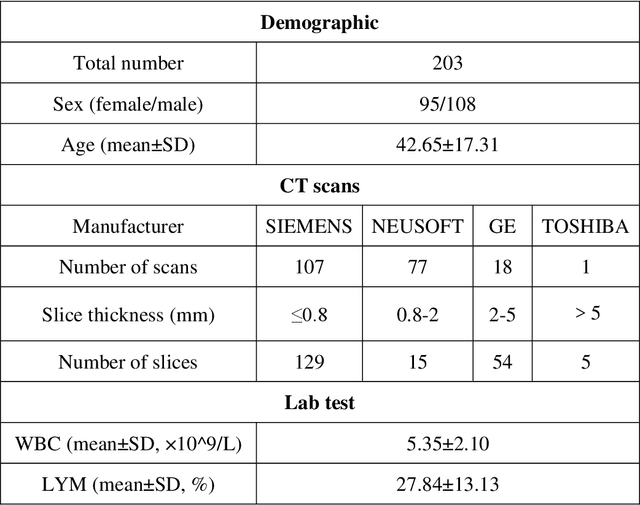
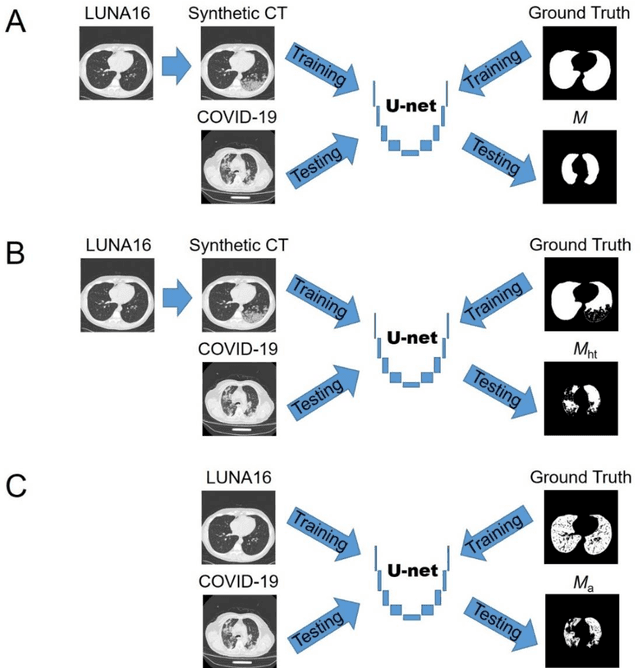
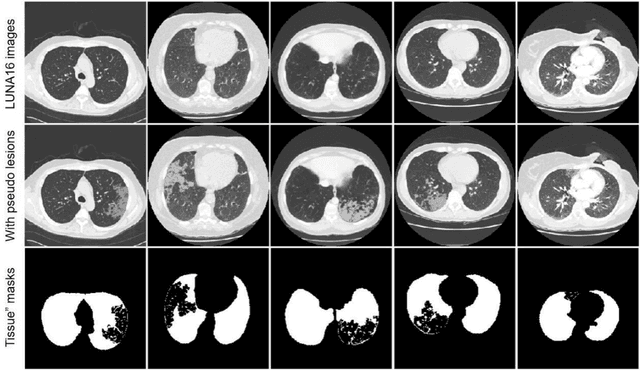
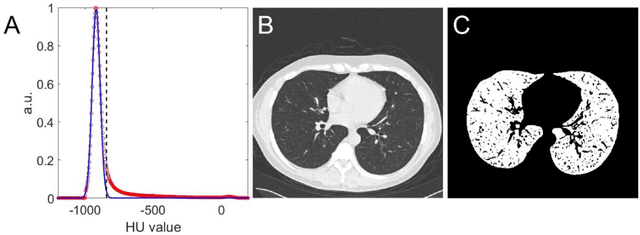
Abstract:Background: Triage of patients is important to control the pandemic of coronavirus disease 2019 (COVID-19), especially during the peak of the pandemic when clinical resources become extremely limited. Purpose: To develop a method that automatically segments and quantifies lung and pneumonia lesions with synthetic chest CT and assess disease severity in COVID-19 patients. Materials and Methods: In this study, we incorporated data augmentation to generate synthetic chest CT images using public available datasets (285 datasets from "Lung Nodule Analysis 2016"). The synthetic images and masks were used to train a 2D U-net neural network and tested on 203 COVID-19 datasets to generate lung and lesion segmentations. Disease severity scores (DL: damage load; DS: damage score) were calculated based on the segmentations. Correlations between DL/DS and clinical lab tests were evaluated using Pearson's method. A p-value < 0.05 was considered as statistical significant. Results: Automatic lung and lesion segmentations were compared with manual annotations. For lung segmentation, the median values of dice similarity coefficient, Jaccard index and average surface distance, were 98.56%, 97.15% and 0.49 mm, respectively. The same metrics for lesion segmentation were 76.95%, 62.54% and 2.36 mm, respectively. Significant (p << 0.05) correlations were found between DL/DS and percentage lymphocytes tests, with r-values of -0.561 and -0.501, respectively. Conclusion: An AI system that based on thoracic radiographic and data augmentation was proposed to segment lung and lesions in COVID-19 patients. Correlations between imaging findings and clinical lab tests suggested the value of this system as a potential tool to assess disease severity of COVID-19.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge